Abstract
Gamma secretase is a multi-subunit complex with aspartic intramembrane protease activity that is involved in the pathogenesis of Alzheimer’s disease in humans. In Arabidopsis thaliana, γ-secretase subunits are localized to endomembrane system compartments and interact with each other in a similar manner to their human counterparts. Here, we identified the protein partners of two plant γ-secretase subunits, presenilin 2 and PEN-2, by tandem affinity purification and co-immunoprecipitation, respectively. Integral membrane proteins were found to interact with presenilin 2, whereas secreted proteins were found to interact with PEN-2. Microscopy screening revealed that the reticulon family protein, RTNLB1, and two single transmembrane domain proteins, TIR-X and the phytolongin PHYL1.1, interact with presenilins. Finally, we show that RTNLB1 interacts with TIR-X. These results represent a step toward elucidating the functions of γ-secretase subunits in plant cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Intramembrane-cleaving proteases (I-CLiPs) include metalloproteases, serine proteases, and aspartic proteases. Signal peptide proteases (SPP) and presenilins are representative aspartic intramembrane proteases. Presenilins maintain the activity of the γ-secretase complex, which is involved in the pathogenesis of Alzheimer’s disease (Parent and Thinakaran 2010). In mammalian cells, the basic γ-secretase subunits include four integral membrane proteins: presenilin 1 (Ps1) or presenilin 2 (Ps2), APH-1a or APH-1b, PEN-2, and nicastrin (Nct) (Fraering et al. 2004; De Strooper and Annaert 2010; Gertsik et al. 2015; Sun et al. 2015). Presenilin comprises the catalytic core of the γ-secretase complex, and the GXGD and YD motifs in the seventh transmembrane domain (TMD) of presenilin are crucial for its proteolytic activity. The modes of substrate processing and the functions of the presenilin 1 and presenilin 2 complexes are somewhat different (Meckler and Checler 2016; Yonemura et al. 2016; Stanga et al. 2017). There are many known substrates of γ-secretase, including an amyloid precursor protein (APP), cadherins, interleukin receptor 1, Notch receptor, and ligands of Notch receptor (McCarthy et al. 2009; Haapasalo and Kovacs 2011; Wężyk and Żekanowski 2017). In addition to their γ-secretase activity, presenilins maintain cell calcium homeostasis via an independent mechanism (Zhang et al. 2010; Kuo et al. 2015; Lee et al. 2015).
Homologs of mammalian presenilins and other γ-secretase subunits have been investigated in the methanogen Methanoculleus marisnigri, the slime mold Dictyostelium discoideum, the moss Physcomitrella patens, and the model plant Arabidopsis thaliana (Ponting et al. 2002; Khandelwal et al. 2007; McMains et al. 2010; Torres-Arancivia et al. 2010; Smolarkiewicz et al. 2014). In archaea (M. marisnigri), the proteolytic activity of the presenilin homolog PSH is similar to that of human γ-secretase (Dang et al. 2015). γ-Secretase activity has been reported in D. discoideum, and the processing of human APP in this organism has been found to depend on presenilin and nicastrin (McMains et al. 2010). Moreover, in D. discoideum, presenilin is indispensable for effective phagocytosis and development, both of which depend on the non-proteolytic activity of presenilin (McMains et al. 2010; Ludtmann et al. 2014). A P. patens (moss) mutant lacking the only functional presenilin found in this species displays pleiotropic phenotypes caused by cytoskeleton dysfunction, which occurs in a γ-secretase-independent manner (Khandelwal et al. 2007). Introducing P. patens presenilin into MEF PSDKO (Mouse Embryonic Fibroblast Presenilin Double Knock-out) compensated only for PSDKO abnormalities related to γ-secretase-independent functions (Khandelwal et al. 2007). Interestingly, human presenilin 1 was able to complement the presenilin-knockout phenotype in P. patens, even when this human protein was inactivated by mutation (Khandelwal et al. 2007).
In seed plants, homologs of γ-secretase subunits exhibit partial amino acid sequence conservation with human proteins (Ponting et al. 2002; Smolarkiewicz et al. 2014). Two presenilin genes were found in the A. thaliana genome (Ps1: AT1G08700, Ps2: AT2G29900), as well as APH-1 (AT2G31440), PEN-2 (AT5G09310), and nicastrin (AT3G52640). In proteomic studies, Ps1 was identified as a trans-Golgi network (TGN) protein, whereas nicastrin was shown to be a plasma membrane protein (Groen et al. 2014; Jones et al. 2014). Microscopy analysis revealed that A. thaliana γ-secretase is primarily localized to the TGN and prevacuolar compartment (PVC) (Smolarkiewicz et al. 2014). Discrepancies in localization data possibly could be a result of complex assembling that occurs, at least in mammals, in different cellular subcompartments (Dries and Yu 2008). Moreover, direct interactions between homologs of γ-secretase subunits have been identified in A. thaliana: presenilins interact with APH-1, PEN-2, and Nct; APH-1 also interacts with Nct; and PEN-2 also interacts with nicastrin (Smolarkiewicz et al. 2014).
These results strongly suggest that γ-secretase complexes are abundant in A. thaliana. However, γ-secretase activity has not been investigated in plants, and the entire complex has yet to be isolated. Mutated PEN-2N74L exhibits an altered localization and is unable to interact with presenilin 2. Moreover, PEN-2 affects the localization of Ps1 and Nct, and is localized to the TGN/PVC. All of these findings suggest that PEN-2 is involved in protein trafficking (Smolarkiewicz et al. 2014). Furthermore, ps1/ps2 plants exhibit accelerated chlorosis upon dark treatment, likely due to impaired autophagy (Smolarkiewicz et al. 2014).
Here, we investigated the proteins that interact with γ-secretase subunits in A. thaliana. We used tandem affinity purification (TAP) and co-immunoprecipitation analysis to identify the interacting proteins of presenilin 2 and PEN-2, respectively. Some of the interactions were verified by FRET-FLIM. We identified a TIR domain-containing protein, as well as RTNLB1 and a SNARE-like ‘phytolongin’ protein, as γ-secretase subunit partners. Finally, we showed that the subcellular localization of human presenilin 1 in plant protoplasts was the same as that of A. thaliana presenilin 1. Our findings shed light on the functions of presenilins and γ-secretase in plants.
Results
Identification of presenilin-interacting proteins
We identified presenilin 2-interacting proteins by tandem affinity purification (TAP) of Ps2:CGSrhino from PSB-D Arabidopsis root cell cultures. Of the proteins detected by MS, seven were categorized as presenilin-2-interacting proteins (Table 1). Five of these seven proteins are integral membrane proteins: two reticulon family proteins, two Endomembrane protein 70 (EMP70) family proteins, and dihydroflavonol 4-reductase. Reticulons are involved in shaping the ER, establishing plasmodesmata, and FLS2 trafficking (Lee et al. 2011, 2013; Knox et al. 2015; Verena Kriechbaumer et al. 2015). The roles of TMN/EMP70 (TM9-type membrane) proteins in plants have not yet been elucidated. However, the targeting of EMP12 (TMN1) to the Golgi has been extensively studied; both microscopy and proteomic analysis have shown that EMP12 and other EMP70 proteins are Golgi/trans-Golgi proteins, as are presenilins (Nikolovski et al. 2012, 2014; Parsons et al. 2012; Gao et al. 2012; Groen et al. 2014; Smolarkiewicz et al. 2014). Two soluble RNA-binding proteins with RRM/RBP/RNP motifs were also identified as Ps2-interacting proteins.We then subjected most of the identified proteins to colocalization and FRET-FLIM analyses to verify their interactions with presenilins (Figs. 1, 2). Colocalization and FRET-FLIM analysis demonstrated that RTNLB1 colocalizes and interacts with presenilins (Fig. 1; Table 2). No interaction with presenilins was confirmed for the other proteins. However, we detected colocalization (Table 2) of presenilins and TMN8, presenilin 1, as well as presenilins and dihydroflavonol 4-reductase/flavonone protein (Fig. 2a–d). RBP45C and Ps2:GFP were found to be localized to distinct subcellular compartments (Fig. 2f). Moreover, presenilin-interacting RTNLB1 colocalize with TMN8, but also no interaction was shown (Fig. 1d; Table 2).
Expression of RTNLB1 in co-expression with Ps1, Ps2, TMN8, Phyl1.1 in A. thaliana mesophyll protoplasts. Fluorescent signals of particular subunits fused with GFP are shown in green, and those of subunits fused with RFP are shown in red. Colocalization with γ-secretase subunits is shown in yellow (merged). Scale bar, 10 µm
Expression of dihydroflavonol 4-reductase (FP), TMN8:GFP, and RBP45C in A. thaliana mesophyll protoplasts. Fluorescent signals of particular subunits fused with GFP are shown in green, and those of subunits fused with RFP are shown in red. Colocalization with γ-secretase subunits is shown in yellow (merged). Scale bars, 10 µm
Identification of PEN-2-interacting proteins
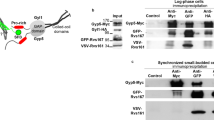
PEN-2:GFP was immunoprecipitated from 35S:PEN-2:GFP A. thaliana cells using an agarose slurry linked to anti-GFP camelid antibodies (GFP-Trap, Chromotek). Five Co-IP replicates were performed using different samples. Identified proteins that were present in the negative control (Co-IP from WT plants) and ribosomal/plastid proteins were subtracted from the result list. Proteins that are co-immunoprecipitated with PEN-2 were different from those identified as interacting with Ps2 via TAP–MS (Tables 1, 3). The presence of BIP1 within the list of PEN-2-interacting proteins might result from the overexpression of this integral membrane protein, which might induce the chaperone activity of BIP1. Half of the identified proteins are secreted proteins (AT1G17860, AT5G48540, AT5G63800, AT5G03350). This result suggests that PEN-2 functions in protein trafficking, as previously suggested (Smolarkiewicz et al. 2014).
Presenilins interact with TIR-X and R-SNARE proteins
Human cells lack substrates with an amino acid motif that is specific for γ-secretase-catalyzed cleavage. However, most animal substrates are type I membrane proteins whose ectodomains must be shed prior to cleavage (Lichtenthaler et al. 2011). To date, no substrate of plant γ-secretase has been identified. Moreover, there are no plant homologs for the most important human γ-secretase substrates, such as APP, cadherins, and Notch receptor. The receptor-like kinase CRINKLY4 (ACR4) was thought to be an Arabidopsis γ-secretase substrate, but this has not been proven experimentally (Walker 2009). We verified several potential plant γ-secretase substrates with single and multiple TMDs, and tested whether they interact with presenilins (data not shown). Proteins that directly interact with presenilin and have a single TMD would represent strong candidates for plant γ-secretase substrates. Two proteins, TIRP and Phyl1.1, were identified by confocal microscopy-based screening as presenilin-interacting proteins.
In mammals, presenilin 1 interacts with interleukin-1 receptor type 1, a Toll-like receptor with a characteristic TIR (Toll-IL-1 receptor) domain (Elzinga et al. 2009). Plant TIR-domain proteins, such as TIR–NBS–LRR, are crucial components of effector-triggered immunity (Burch-Smith et al. 2007; Swiderski et al. 2009; Zhang et al. 2017). TIRP is a small TIR-X family protein (23.5 kDa) with an N-terminal TIR domain and a single TMD (Fig. 3) (Meyers et al. 2002; Nandety et al. 2013). Consensus motifs of the TIR domain were found within the N-terminal fragment of TIRP (Fig. 3). In Arabidopsis protoplasts, TIRP:GFP colocalized with γ-secretase subunits as well as NIP1;1, which was used as an ER marker (Fig. 4). Moreover, FRET-FLIM analysis showed that presenilins, APH-1, PEN-2, and RTNLB1 directly interact with TIRP (Table 2).
TIRP amino acid sequence analysis. a Prediction of a single TMD in the amino acid sequence of TIRP by TMHMM Server, v.2.0; b alignment of the amino acid sequence of TIRP and the conserved TIR-X domain sequence (Meyers et al. 1999) by Clustal Omega
Simultaneous expression of TIRP with γ-secretase subunits, RTNLB1, and ER marker NIP1;1 in A. thaliana mesophyll protoplasts. Fluorescent signals of particular subunits fused with GFP are shown in green, and those of subunits fused with RFP are shown in red. Colocalization of γ-secretase subunits is shown in yellow (merged). Scale bars, 10 µm
In mammalian cells, presenilin 1 interacts with the SNARE proteins Syntaxin 1A and Syntaxin 5 (Smith et al. 2000; Suga et al. 2004). Moreover, Golgi stress and the upregulation of Syntaxin 5 are associated with a reduction in APP processing (Suga et al. 2015). PHYL1.1 belongs to a plant-specific group of R-SNARE proteins named ‘phytolongins’. These proteins lack a canonical SNARE motif, which was replaced by a longer domain known as the ‘Phyl domain’. PHYL1.1 localized to the plasma membrane, Golgi bodies, and post-Golgi compartments when expressed in N. benthamiana epidermal cells (De Marcos Lousa et al. 2016). The similar expression patterns of Phyl2.1, Phyl2.2, and RTNLB11 are thought to be a consequence of their common involvement in ER–PM interactions (De Marcos Lousa et al. 2016). In protoplasts, PHYL1.1:GFP colocalized with presenilins (Fig. 5; Table 2). FRET-FLIM confirmed direct interactions between PHYL1.1, and both Ps1 and Ps2 (FRETEff = 14–16%; Table 2). No interaction was detected between PHYL1.1:GFP and PEN-2:RFP. PHYL1.1 is a potential plant γ-secretase substrate, as it contains a single TMD and can interact with presenilins. When co-expressed in protoplasts, RTNLB1:RFP and PHYL1.1:GFP were observed in different subcellular compartments (Fig. 5d). On the other hand, Phyl1.1 colocalizes with TIRP, but no direct interaction was revealed (Fig. 5e; Table 2).
Both the N- and C-terminal fragments of TIRP interact with presenilins
We investigated whether the presence of the N-terminal fragment of TIRP lacking TMD (TIRPΔTMD) is sufficient for establishing the presenilin–TIRP interaction. TIRPΔTMD:GFP colocalized with Ps1:RFP, but in some cases, these proteins were spatially separated (Fig. 6b). In two-thirds of co-transformed protoplasts, their colocalization was detected (65%, n = 50; Fig. 6a). Such differentiation was not observed for TIRP/Ps1: colocalization was observed in every co-transformed protoplast. FRET-FLIM analysis of the TIRPΔTMD:GFP-Ps1:RFP interaction showed that these molecules directly interact with each other (Table 2). Hence, the transmembrane domain of TIRP is dispensable for establishing the TIRP–presenilin interaction, but the lack of the TMD in TIRP decreases the probability of TIRP and Ps1 colocalization.
However, the truncated TIRPΔTIR:GFP, which lacks the N-terminal fragment but possesses a TMD domain, interacted with Ps2:GFP and APH-1:GFP (Fig. 6c–e; Table 2). Colocalization was observed for TIRPΔTIR and PEN-2, but no interaction was detected between these proteins (Fig. 6d; Table 2). Therefore, the interaction of presenilins with TIRP involves both the intramembrane and TIR-domain-containing fragments, pointing to strong physical interactions between TIRP and presenilins, as supported by the high FRET efficiency values (15–18%).
Overexpression of TIRP in N. benthamiana leads to cell death
The transient overexpression of various TIR–NBS and TIR-X genes in N. benthamiana leads to necrosis, a symptom of the hypersensitive response (Swiderski et al. 2009; Nandety et al. 2013). Similarly, we found that the transient overexpression of TIRP resulted in necrosis at 3 days after inoculation (Fig. 7b). We detected stronger necrotic effects in plants overexpressing TIRPΔTMD:GFP (Fig. 7a) and almost no effect in plants overexpressing Ps1:RFP (Fig. 7c). The differences in phenotypic changes after TIRP and TIRPΔTMD:GFP overexpression were also visible at the cellular level: TIRP:GFP overexpression (Fig. 7d, e) led to slower formation of large cellular protein deposits than TIRPΔTMD:GFP overexpression (Fig. 7f). Two days after transformation, TIRP:GFP was observed in reticular-like compartments (Fig. 7d), and on the third day, large aggregates of this protein were detected (Fig. 7e). Beginning on the second day after transformation, TIRPΔTMD:GFP was present in large aggregates (Fig. 7f). Perhaps these aggregates formed due to processes triggered by oligomerization of the TIR domain, which is thought to be an important component of signaling related to effector-triggered immunity (Swiderski et al. 2009; Nishimura et al. 2017). These phenotypic effects suggest that TIRP is involved in defense responses against pathogens in a similar manner to other TIR-X or TIR proteins (Nandety et al. 2013; Nishimura et al. 2017).
Expression of TIRPΔTMD:GFP (a), TIRP:GFP (b), and Ps1:RFP (c) in tobacco epidermal cells. Confocal images of TIRP:GFP 2 (d) and 3 (e) days after inoculation. Confocal images of TIRPΔTMD:GFP 2 days after inoculation. TIRP:GFP overexpressed in N. benthamiana epidermis was observed in reticular-like compartments. Scale bar, 10 µm
Presenilins do not interact with other TIR-NBS or TIR-X proteins
TIRP belongs to the TIR-X protein family, which is characterized by the presence of a TIR domain and the absence of both LRR and NBS domains (Meyers et al. 2002). Both TNL (TIR–NBS–LRR) and TIR–NBS/TIR-X proteins are involved in immunological processes in plants (Swiderski et al. 2009; Bernoux et al. 2011; Nandety et al. 2013; Zhao et al. 2015; Nishimura et al. 2017). We, therefore, investigated whether other TIR–NBS and TIR-X proteins (TN10:GFP, TN11:GFP, TN21:GFP, TX21:GFP) also interact with presenilins. TIR–NBS and TIR-X were reported to be localized to membranes, the cytosol, or the nucleus (Nandety et al. 2013). When TN10:GFP and TN11:GFP were co-expressed with Ps1:RFP, they did not colocalize (Fig. 8a, b). Also, no colocalization was observed when plasma membrane and cytoplasmic aggregates localized protein, TN21:GFP, was coexpressed with Ps2:RFP (Fig. 8c). TX21:GFP and Ps2:RFP partially colocalized (Fig. 8d). However, the poor condition of protoplasts expressing TX21:GFP precluded analysis of direct protein interactions by FRET-FLIM. The presenilin-TIRP interaction likely differentiates TIRP from other TIR-X proteins.
Presenilins are not responsible for FLS2 trafficking
FLS2 is a flagellin receptor that undergoes ligand-induced endocytosis in an ESCRT-I-dependent manner (Robatzek and Chinchilla 2006; Spallek et al. 2013). In mammals, the γ-secretase complex is involved in regulating intracellular protein trafficking (Smolarkiewicz et al. 2013; Greenough 2016). In A. thaliana, RTNLB1 and RTNLB2 are required for efficient trafficking of FLS2 to the plasma membrane. Moreover, reticulons directly interact with FLS2 (Lee et al. 2011). We examined whether presenilins regulate RTNLB1/RTNLB2-dependent FLS2 trafficking. No difference was observed between the localization of FLS2 in WT vs. ps1/ps2 protoplasts (Fig. 9a). In both genotypes, the trafficking of FLS2:GFP to the plasma membrane was not disturbed, as was reported for FLS2:GFP in rtnlb1/rtnlb2 protoplasts (Lee et al. 2011). To confirm that the trafficking of presenilin 1 depends on RTNLB1, we transformed bti1(rtnlb1) protoplasts with Ps1:GFP. We found that the trafficking of Ps1:GFP was not disturbed in the absence of RTNLB1 (Fig. 10). Expressing FLS2:GFP together with Ps1:RFP resulted in the partial colocalization of these proteins (Fig. 9b). Nonetheless, although FLS2 interacts with RTNLB1, it does not interact with Ps2 (Table 2). These results demonstrate that presenilins are not responsible for RTNLB1/RTNLB2-dependent trafficking of FLS2. Moreover, the trafficking of presenilin 1 does not depend on RTNLB1.
Localization of FLS2:GFP; FLS2 reached the plasma membrane in both WT and ps1/ps2 protoplasts (a). Simultaneous expression of FLS2 with presenilins, PEN-2, and APH-1 (b–d). Fluorescence signals of particular subunits fused with GFP are shown in green, and those of subunits fused with RFP are shown in red. Colocalization of proteins is shown in yellow (merged). Scale bars, 10 µm
A. thaliana presenilins do not form homodimers
Human presenilin 1 was shown to form homodimers in intact cells, but other methods of analysis gave different results (Herl et al. 2006; Midde et al. 2014; Naing et al. 2018). Using FRET-FLIM, we examined whether Arabidopsis presenilins form homodimers in plant cells. FRET-FLIM analysis of the interactions of both Ps1:GFP/Ps1:RFP and Ps2:GFP/Ps2:RFP failed to demonstrate that plant presenilins form homodimers (Table 2). This result suggests that animal and plant γ-secretases differ in terms of their activity or ability to form passive Ca2+ channels in the ER via presenilins (Midde et al. 2014).
Human presenilin 1 is localized to the same subcellular compartments as Arabidopsis γ-secretase subunits in protoplasts
Both human and Arabidopsis presenilins contain conserved amino acid motifs that are crucial for human γ-secretase assembly and activity (Smolarkiewicz et al. 2014). To investigate the functionality of human presenilin 1 (HsPs1) in plant cells, we expressed HsPs1 in protoplasts. HsPs1:GFP colocalized with Arabidopsis Ps1:RFP or PEN-2:RFP (Fig. 11). We also detected colocalization of HsPs1 and both Arabidopsis Ps1:RFP and PEN-2:RFP, but no direct interaction between HsPs1 and PEN-2 was detected by FRET-FLIM (Table 2). Hence, it is likely that human presenilin 1 cannot form an active γ-secretase in plant cells due to its inability to interact with Arabidopsis PEN-2. The phenotype of a presenilin-deficient P. patens mutant was successfully complemented by the expression of human presenilin 1 that had been inactivated by a mutation, suggesting that the non-catalytic function of presenilin likely plays a key role in plant development (Khandelwal et al. 2007). However, it appears that presenilins use evolutionarily conserved targeting mechanisms that enable non-native presenilin 1 to obtain the same subcellular localization as native presenilin 1.
Simultaneous expression of HsPs1 with Arabidopsis presenilins (AtPs1) and AtPEN-2 in A. thaliana mesophyll protoplasts. Fluorescent signals of particular subunits fused with GFP are shown in green, and those of subunits fused with RFP are shown in red. Colocalization of proteins is shown in yellow (merged). Scale bar, 10 µm
Discussion
Plant presenilins of the γ-secretase complex are the least-studied intramembrane proteases in plants (Adam 2013). Preliminary experiments indicated that in A. thaliana the presenilins APH-1, PEN-2, and nicastrin are localized within the endomembrane system and that they interact with each other (Smolarkiewicz et al. 2014; Groen et al. 2014). In mammals, the assembly of the γ-secretase complex is a multistep process involving additional proteins. No γ-secretase subunits (except baits) were detected in TAP–MS and co-immunoprecipitation experiments in plants, perhaps due to the use of single-gene overexpression systems. It is also possible that the assembly of the plant γ-secretase complex is developmentally regulated or induced by stress and that this complex might not occur ubiquitously in all plant tissues.
Here, we provided new data on the interactome of Arabidopsis γ-secretase subunits. The single TMD proteins TIRP and PHYL1.1, and the reticulon family protein, RTNLB1, were identified as presenilin partners. Moreover, TMN8 and dihydroflavonol 4-reductase/flavanone protein occupy the same subcellular compartment as presenilins. The single transmembrane domains of TIRP and PHYL1.1 should be considered to be potential substrates of γ-secretase. Moreover, a TMD is a feature that differentiates TIRP from TIR–NBS–LRR and from most other TIR-X proteins, as most plant TIR-domain proteins lack a TMD (Sanseverino and Ercolano 2012). The closest TIRP homolog, TX14 (AT2G32140), a soluble protein, is thought to be involved in defense responses and salicylic acid signaling (Kato et al. 2014). Arabidopsis plants overexpressing TX14 had a dwarf phenotype and increased expression of defense-related genes (Kato et al. 2014). Other TIR-X- and TIR–NBS-overexpressing lines were characterized by enhanced tolerance to Pst DC3000 and Fusarium oxysporum infection, as well as elevated salicylic acid levels (Nandety et al. 2013). Similarly, the overexpression of various TIR-X and TIR–NBS genes in N. benthamiana triggered necrotic-like processes (Nandety et al. 2013). Some TIR-X and TIR–NBS proteins were found to interact with effector proteins. Preliminary findings pointed to the likely involvement of TIR–NBS and TIR-X proteins in guard complexes that monitor pathogen effector proteins (Nandety et al. 2013). TIRP might integrate effector-triggered immunity with endomembrane proteins, such as presenilin or RTNLB1, which might influence the recognition of pathogen effectors or downstream signaling pathways.
In the current study, both TAP-MS and FRET-FLIM analyses indicated that Ps2 and RTNLB1 interact. We showed that RTNLB1 also interacts with presenilin 1. No interaction was detected between RTNLB1 and PEN-2, the smallest subunit of the γ-secretase complex in animals. Interestingly, mice overexpressing any of the native reticulon-encoding genes had reduced β-amyloid levels, although this was not likely due to a decrease in γ-secretase activity (Shi et al. 2009). In plants, reticulons are responsible for forming the tubular regions of the ER (Sparkes et al. 2010; Tolley et al. 2010; Lee et al. 2013). Additionally, reticulons regulate protein trafficking and immune responses in plants by remodeling the ER structure, forming plasma membrane–ER contact sites and desmotubules (Knox et al. 2015; Levy et al. 2015; Verena Kriechbaumer et al. 2015; Griffing et al. 2016). Reticulons are also involved in other immune-related processes, such as FLS2 trafficking and interactions with the Agrobacterium protein VirB2 (Hwang and Gelvin 2004; Lee et al. 2011). However, we demonstrated that presenilins, which regulate protein trafficking in mammals (Greenough 2016), are not responsible for RTNLB1/RTNLB2-dependent FLS2 trafficking in Arabidopsis. It is possible that the tubular ER structures formed by reticulons are important for the functioning of presenilins and TIRP, as they are mainly localized within the ER subcompartment occupied by RTNLB1. Endomembrane protein 70 family proteins, including TMN12, as well as RTNLB1, were previously identified as RTNLB3- and RTNLB6-interacting proteins (Verena Kriechbaumer et al. 2015). TMN8 colocalizes with presenilins, but no direct interaction was detected. In the current study, we did not demonstrate that TMN8 and RTNLB1 interact. In a proteomic study, EMP70 family proteins were identified together with Ps2 as trans-Golgi residents (Groen et al. 2014). The functions of EMP70 proteins in plants are still elusive, although preliminary results suggest the involvement of A. thaliana EMP70 (TMN7) in regulating cellular Cu2+ levels (Hegelund et al. 2010). It is possible that these proteins are important for the proper functioning of the root elongation zone, which requires large amounts of Cu2+, or for immune responses (Hegelund et al. 2010). The roles of the protein network involving EMP70, reticulons, TIRP, and γ-secretase subunits remain to be deciphered.
Furthermore, little is known about the roles of PHYL1.1 (AT4G10170). PHYL1.1 expression is regulated by a conserved peptide encoded by a type II open reading frame upstream of the main open reading frame sequence (Ebina et al. 2015). Phytolongins are R-SNARE proteins involved in membrane fusion. These proteins occupy different subcellular compartments. PHYL1.1 is localized to the plasma membrane, Golgi, and post-Golgi (De Marcos Lousa et al. 2016). Presenilins, but not PEN-2, interact with PHYL1.1, suggesting that presenilins play specific roles in plant cells. Numerous studies have shown that mammalian presenilins have γ-secretase-independent functions (Smolarkiewicz et al. 2013). PEN-2 is recruited to autophagosomes together with ATG8, suggesting that PEN-2 might be involved in autophagy. Autophagy is also associated with the establishment of disease resistance and the hypersensitive response (Hofius et al. 2011). In an analysis of proteins that co-immunoprecipitated with PEN-2, a few (mostly secreted and ER) proteins were identified. One of these proteins was DGL1, which was also found to interact with RTNLB3 and RTNLB6 (Verena Kriechbaumer et al. 2015). Although we still lack a full understanding of the roles of plant γ-secretase, in the current study, we obtained new molecular data about proteins that interact with two γ-secretase subunits in A. thaliana cells. These results provide preliminary information about the functional roles of plant γ-secretase. Nonetheless, the roles of plant γ-secretase, the assembly of the γ-secretase complex, and its abundance in plants remain to be elucidated.
Materials and methods
Plant materials and cultivation
Arabidopsis thaliana ecotype Columbia (Col-0) was used as the wild type. The following T-DNA insertion lines were used: ps1/ps2 (Smolarkiewicz et al. 2014) and bti1 (SALK 069542C). Plants were grown in a Versatile Environmental Test Chamber MLR-350H (Sanyo) at 22 °C under a 16-h light/8-h dark cycle at 70% relative humidity (RH). Nicotiana benthamiana was grown at 28 °C under a 16-h light/8-h dark cycle at 70% RH.
Vector construction
To amplify TIRP, ABRC clone S63497 was used. TIRDom was obtained by amplification of the sequence encoding the 200 N-terminal amino acids of TIRP, harboring a TIR domain but not a TMD. PHYL1.1 was amplified from ABRC clone U22726. TMN8/EMP1 and RBP45C were amplified from mRNA derived from root and rosette leaves, respectively. The dihydroflavonol 4-reductase gene was amplified from RNA isolated from flowers. The PCR products were cloned into pENTR/D-TOPO using a pENTR/D-TOPO Cloning Kit (Life Technologies) or by NotI/AscI-based cloning (Fast Digest, Thermo Fisher Scientific). Cloning into destination plasmids from the pSITE or pSAT series was performed using an LR Clonase II Kit (Life Technologies). The use of these destination vectors allowed genes to be transiently overexpressed in plant cells together with an N-terminal or C-terminal fluorescent protein tag. The names of the protein constructs with fluorescent protein tags reflect N-to-C peptide order.
Transient transformation
Transient expression assays were performed using protoplasts isolated from Arabidopsis mesophyll cells as previously described (Smolarkiewicz et al. 2014). Agrobacterium tumefaciens GV3101 cells transformed with binary vectors from the pSITE series were cultivated overnight (4 mL, 28 °C, 180 rpm) in LB medium, centrifuged (4 °C, 6 min., 8000g), and resuspended in T buffer (100 mM MES pH 5,7; 100 mM MgCl2; 150 µM acetosyringone) to OD600 0.15–0.3. After 3 h of incubation with shaking (room temperature, 70 rpm), 1 mL of A. tumefaciens cells was injected into the epidermis of 4-week-old N. benthamiana leaves using a 2-mL syringe.
Microscopy
All images were obtained with a Nikon A1R confocal microscope. Protoplast images were acquired using Apo 40×/1.25 WI λS DIC N2, Plan Apo VC 60×/1.4 Oil DIC N2, or Plan Apo VC 100×/1.4 Oil DIC N2 objectives. For N. benthamiana imaging, a Plan Apo VC 20×/0.75 DIC N2 objective was used. EGFP fluorescence was excited with a 488-nm laser and captured using a 500–550-nm emission filter. mRFP and mCherry were excited with a 561-nm laser and captured using a 570–620-nm emission filter. A spectral detector was used to verify the fluorescence spectra of the fluorochromes. Colocalization was quantified as Pearson’s correlation coefficient (PCE) for at least 20 independent protoplasts. Protein–protein interactions were analyzed by FRET-FLIM as described earlier (Smolarkiewicz et al. 2014). Donor fluorescence was excited with a 485-nm Pulsed Diode Laser; PDL 800-D; 50 mHz. Picoquant’s SymPhoTime 64 software was used for data analysis. Complete fluorescence lifetime decays were calculated per pixel for selected fragments and fitted using a double exponential decay model. Histograms were drawn based on data from protoplasts lacking a distinct peak of low lifetime signals (< 1 ns), which was considered to represent nonspecific signals coming from excited chloroplasts. FRET efficiency was calculated as FRETEff = (1 − τDA/τD) × 100%, where τD is the fluorescence lifetime of the donor in the absence of acceptor and τDA is that of the donor in the presence of acceptor. The lifetime of free eGFP in A. thaliana mesophyll protoplasts was 2.49 ns, and the lifetime of eGFP in the presence of mRFP was 2.5 ns, which is equal to 0% FRETEff and cut-off threshold. A shorter photon lifetime of the donor in the presence of acceptor would make the FRETEff > 0%, indicating that the distance between the donor and acceptor molecules was < 10 nm. A distance of 10 nm corresponds to a direct interaction between proteins (Day and Davidson 2013).
SDS-PAGE and immunoblotting
Prior to loading, the centrifuged protoplasts were denaturated at 95 °C for 5 min in SB buffer (0.16 M Tris, pH 6.8, 7% SDS, 33% glycerol, and 2% β-mercaptoethanol). Proteins were separated in a 10% polyacrylamide gel with 0.05% SDS. Semi-dry electrotransfer from gel to polyvinylidene fluoride (PVDF, Immobilon-P) was performed at 0.8 mA/cm2 for 45 min. Anti-GFP antibody (Santa Cruz Biotechnologies, sc-9996) was used at a 1:1000 dilution in PBST (phosphate buffered saline with Tween® 20). Peroxidase-conjugated secondary antibody was used at a 1:2500 dilution in PBST (Santa Cruz Biotechnologies, sc-2005).
PEN-2 immunoprecipitation
PEN-2:GFP was immunoprecipitated from the 35S:PEN-2:GFP overexpression line and from WT col-0 as a negative control. Leaves of 4-week-old A. thaliana plants were ground in liquid nitrogen, suspended in lysis buffer (10 mM Tris/Cl pH 7.5, 150 mM NaCl, 0.5 mM EDTA, 1 mM PMSF, Protease Inhibitor Cocktail [Sigma-Aldrich], and 1% Nonidet P40), and incubated on ice for 30 min. After centrifugation (12,000g; 4 °C; 10 min.), the pellet was resuspended in dilution buffer (10 mM Tris/Cl, pH 7.5, 150 mM NaCl, 0.5 mM EDTA, 1 mM PMSF, and Protease Inhibitor Cocktail [Sigma-Aldrich]). The protein extracts were incubated for 1 h at 4 °C with shaking (200 rpm) with agarose slurry linked to anti-GFP camelid antibodies (GFP-Trap, Chromotek) and washed with dilution buffer. Proteins were identified in a publicly available database by fragment ion analysis and peptide mass fingerprinting using MASCOT at the Mass Spectrometry Laboratory (Institute of Biochemistry and Biophysics, Polish Academy of Sciences, Warsaw) using standard protocols. Data sets were searched against the TAIR10 database using the following settings: tryptic peptides with one missed cleavage site, 20 ppm for peptides, and 0.6 Da for fragment ion mass tolerance. Proteins were reported as being successfully identified if their probability value was below 0.05 (P < 0.05). Only these hits that were present in at least three out of five biological replicates and were not present in any out of two negative controls were taken into account. Ribosomal, mitochondrial, plastids peptides, and hits with emPAI equal or lower than 0.05, were not placed on results list.
Tandem affinity purification
PSB-D cell culture and transformation, protein purification, and MS analysis were performed in Geert de Jaeger’s laboratory as described previously (Van Leene et al. 2007, 2008). Presenilin 2 with CGSrhino tag on its C-terminus was overexpressed in Arabidopsis PSB-D root cell cultures. The cells were solubilized with 1% digitonin, 1% Triton X-100, and 1% C12E8. Proteins purified with Ps2 were identified with an LTQ Orbitrap Velos mass spectrometer (Thermo Fisher Scientific) and searched against the TAIR10 database. The results were compared to known background proteins, annotated as background in previous TAP experiments in Geert de Jaeger’s laboratory. These proteins were subtracted from the result list.
Author contribution statement
Conceptualization, TS; Methodology, TS and GJ; Investigation, TS, RK, SW, KG, ES, GJ, KL; Writing-Original Draft Preparation, TS; Supervision, PW; Funding Acquisition, PW, KL and TS.
Abbreviations
- APH-1:
-
Anterior pharynx defective 1
- APP:
-
Amyloid precursor protein
- Co-IP:
-
Co-immunoprecipitation
- FLS2:
-
Flagellin sensitive 2
- FRET-FLIM:
-
Förster resonance energy transfer by fluorescence lifetime imaging
- GFP:
-
Green fluorescence protein
- MEF:
-
Mouse embryonic fibroblast
- PEN-2:
-
Presenilin enhancer 2
- Ps:
-
Presenilin
- PSDKO:
-
Presenilin double knock-out
- RFP:
-
Red fluorescence protein
- SNARE:
-
Soluble NSF(N-ethylmaleimide-sensitive factor) attachment protein) receptor
- TAP:
-
Tandem affinity purification
- TIR:
-
Toll/interleukin-1 receptor homology domain
- TMD:
-
Transmembrane domain
References
Adam Z (2013) Emerging roles for diverse intramembrane proteases in plant biology. Biochim Biophys Acta 1828:2933–2936. https://doi.org/10.1016/j.bbamem.2013.05.013
Bernoux M, Ve T, Williams S et al (2011) Structural and functional analysis of a plant resistance protein TIR domain reveals interfaces for self-association, signaling, and autoregulation. Cell Host Microbe 9:200–211. https://doi.org/10.1016/j.chom.2011.02.009
Burch-Smith TM, Schiff M, Caplan JL et al (2007) A novel role for the TIR domain in association with pathogen-derived elicitors. PLoS Biol 5:e68. https://doi.org/10.1371/journal.pbio.0050068
Dang S, Wu S, Wang J et al (2015) Cleavage of amyloid precursor protein by an archaeal presenilin homologue PSH. Proc Natl Acad Sci. https://doi.org/10.1073/pnas.1502150112
De Strooper B, Annaert W (2010) Novel research horizons for presenilins and γ-secretases in cell biology and disease. Annu Rev Cell Dev Biol 26:235–260. https://doi.org/10.1146/annurev-cellbio-100109-104117
De Marcos Lousa C, Soubeyrand E, Bolognese P et al (2016) Subcellular localization and trafficking of phytolongins (non-SNARE longins) in the plant secretory pathway. J Exp Bot 67:2627–2639. https://doi.org/10.1093/jxb/erw094
Dries D, Yu G (2008) Assembly, Maturation, And Trafficking Of The γ- secretase complex in Alzheimer’s disease. Curr Alzheimer Res 5:132–146
Ebina I, Takemoto-Tsutsumi M, Watanabe S et al (2015) Identification of novel Arabidopsis thaliana upstream open reading frames that control expression of the main coding sequences in a peptide sequence-dependent manner. Nucleic Acids Res. https://doi.org/10.1093/nar/gkv018
Elzinga BM, Twomey C, Powell JC et al (2009) Interleukin-1 receptor type 1 is a substrate for gamma-secretase-dependent regulated intramembrane proteolysis. J Biol Chem 284:1394–1409. https://doi.org/10.1074/jbc.M803108200
Fraering PC, Ye W, Strub J-M et al (2004) Purification and characterization of the human gamma-secretase complex. Biochemistry 43:9774–9789. https://doi.org/10.1021/bi0494976
Gao C, Yu CKY, Qu S et al (2012) The Golgi-localized Arabidopsis endomembrane protein12 contains both endoplasmic reticulum export and Golgi retention signals at its C terminus. Plant Cell 24:2086–2104. https://doi.org/10.1105/tpc.112.096057
Gertsik N, Chiu D, Li Y-M (2015) Complex regulation of Î3-secretase: from obligatory to modulatory subunits. Front Aging Neurosci 6:1–10. https://doi.org/10.3389/fnagi.2014.00342
Greenough MA (2016) The role of presenilin in protein trafficking and degradation—implications for metal homeostasis. J Mol Neurosci 60:289–297. https://doi.org/10.1007/s12031-016-0826-4
Griffing LR, Lin C, Perico C et al (2016) Plant ER geometry and dynamics: biophysical and cytoskeletal control during growth and biotic response. Protoplasma. https://doi.org/10.1007/s00709-016-0945-3
Groen AJ, Sancho-Andre G, Breckels LM, Gatto L, Aniento F, Lilley KS (2014) Identification of Trans-Golgi Network Proteins in Arabidopsis thaliana Root Tissue. J Proteome Res 13(2):763–776. https://doi.org/10.1021/pr4008464
Haapasalo A, Kovacs DM (2011) The many substrates of presenilin/gamma-secretase. J Alzheimers Dis 25:3–28. https://doi.org/10.3233/JAD-2011-101065
Hegelund JN, Jahn TP, Baekgaard L et al (2010) Transmembrane nine proteins in yeast and Arabidopsis affect cellular metal contents without changing vacuolar morphology. Physiol Plant 140:355–367. https://doi.org/10.1111/j.1399-3054.2010.01404.x
Herl L, Lleo A, Thomas AV et al (2006) Detection of presenilin-1 homodimer formation in intact cells using fluorescent lifetime imaging microscopy. Biochem Biophys Res Commun 340:668–674. https://doi.org/10.1016/j.bbrc.2005.12.063
Hofius D, Munch D, Bressendorff S et al (2011) Role of autophagy in disease resistance and hypersensitive response-associated cell death. Cell Death Differ 18:1257–1262. https://doi.org/10.1038/cdd.2011.43
Hooper CM, Castleden IR, Tanz SK et al (2017) SUBA4: The interactive data analysis centre for Arabidopsis subcellular protein locations. Nucleic Acids Res 45:D1064–D1074. https://doi.org/10.1093/nar/gkw1041
Hwang H-H, Gelvin SB (2004) Plant proteins that interact with VirB2, the Agrobacterium tumefaciens pilin protein, mediate plant transformation. Plant Cell 16:3148–3167. https://doi.org/10.1105/tpc.104.026476
Jones AM, Xuan Y, Xu M et al (2014) Border control—a membrane-linked interactome of Arabidopsis. Sci (New York NY) 344:711–716
Kato H, Saito T, Ito H et al (2014) Overexpression of the TIR-X gene results in a dwarf phenotype and activation of defense-related gene expression in Arabidopsis thaliana. J Plant Physiol 171:382–388. https://doi.org/10.1016/j.jplph.2013.12.002
Khandelwal A, Chandu D, Roe CM et al (2007) Moonlighting activity of presenilin in plants is independent of gamma-secretase and evolutionarily conserved. Proc Natl Acad Sci USA 104:13337–13342. https://doi.org/10.1073/pnas.0702038104
Knox K, Wang P, Kriechbaumer V et al (2015) Putting the squeeze on plasmodesmata: a role for reticulons in primary plasmodesmata formation. Plant Physiol 168:1563–1572. https://doi.org/10.1104/pp.15.00668
Kuo IY, Hu J, Ha Y, Ehrlich BE (2015) Presenilin-like GxGD membrane proteases have dual roles as proteolytic enzymes and ion channels. J Biol Chem 1:jbc.M114.629584. https://doi.org/10.1074/jbc.M114.629584
Lee HY, Bowen CH, Popescu GV et al (2011) Arabidopsis RTNLB1 and RTNLB2 Reticulon-like proteins regulate intracellular trafficking and activity of the FLS2 immune receptor. Plant Cell 23:3374–3391. https://doi.org/10.1105/tpc.111.089656
Lee H, Sparkes I, Gattolin S et al (2013) An Arabidopsis reticulon and the atlastin homologue RHD3-like2 act together in shaping the tubular endoplasmic reticulum. New Phytol 197:481–489. https://doi.org/10.1111/nph.12038
Lee J-H, McBrayer MK, Wolfe DM et al (2015) Presenilin 1 maintains lysosomal Ca2+ homeostasis via TRPML1 by regulating vATPase-mediated lysosome acidification. Cell Rep 12:1430–1444. https://doi.org/10.1016/j.celrep.2015.07.050
Levy A, Zheng JY, Lazarowitz SG (2015) Synaptotagmin SYTA forms ER-plasma membrane junctions that are recruited to plasmodesmata for plant virus movement. Curr Biol 25:2018–2025. https://doi.org/10.1016/j.cub.2015.06.015
Lichtenthaler SF, Haass C, Steiner H (2011) Regulated intramembrane proteolysis–lessons from amyloid precursor protein processing. J Neurochem 117:779–796. https://doi.org/10.1111/j.1471-4159.2011.07248.x
Ludtmann MHR, Otto GP, Schilde C et al (2014) An ancestral non-proteolytic role for presenilin proteins in multicellular development of the social amoeba Dictyostelium discoideum. J Cell Sci. https://doi.org/10.1242/jcs.140939
McCarthy JV, Twomey C, Wujek P (2009) Presenilin-dependent regulated intramembrane proteolysis and gamma-secretase activity. Cell Mol life Sci C 66:1534–1555
McMains VC, Myre M, Kreppel L, Kimmel AR (2010) Dictyostelium possesses highly diverged presenilin/gamma-secretase that regulates growth and cell-fate specification and can accurately process human APP: a system for functional studies of the presenilin/gamma-secretase complex. Dis Model Mech 3:581–594. https://doi.org/10.1242/dmm.004457
Meckler X, Checler F (2016) Presenilin 1 and presenilin 2 target γ-secretase complexes to distinct cellular compartments. J Biol Chem 291:12821–12837. https://doi.org/10.1074/jbc.M115.708297
Meyers BC, Dickerman AW, Michelmore RW et al (1999) Plant disease resistance genes encode members of an ancient and diverse protein family within the nucleotide-binding superfamily. Plant J 20:317–332. https://doi.org/10.1046/j.1365-313X.1999.00606.x
Meyers BC, Morgante M, Michelmore RW (2002) TIR-X and TIR-NBS proteins: two new families related to disease resistance TIR-NBS-LRR proteins encoded in Arabidopsis and other plant genomes. Plant J 32:77–92
Midde K, Rich R, Saxena A et al (2014) Membrane topology of human presenilin-1 in SK-N-SH cells determined by fluorescence correlation spectroscopy and fluorescent energy transfer. Cell Biochem Biophys 70:923–932. https://doi.org/10.1007/s12013-014-9999-z
Naing S-H, Oliver RC, Weiss KL et al (2018) Solution structure of an intramembrane aspartyl protease via small angle neutron scattering. Biophys J 114:602–608. https://doi.org/10.1016/j.bpj.2017.12.017
Nandety RS, Caplan JL, Cavanaugh K et al (2013) The role of TIR-NBS and TIR-X proteins in plant basal defense responses. Plant Physiol 162:1459–1472. https://doi.org/10.1104/pp.113.219162
Nikolovski N, Rubtsov D, Segura MP et al (2012) Putative glycosyltransferases and other plant Golgi apparatus proteins are revealed by lopit proteomics. Plant Physiol 160:1037–1051. https://doi.org/10.1104/pp.112.204263
Nikolovski N, Shliaha PV, Gatto L et al (2014) Label-free protein quantification for plant Golgi protein localization and abundance. Plant Physiol 166:1033–1043. https://doi.org/10.1104/pp.114.245589
Nishimura MT, Anderson RG, Cherkis KA et al (2017) TIR-only protein RBA1 recognizes a pathogen effector to regulate cell death in Arabidopsis. Proc Natl Acad Sci 114:E2053–E2062. https://doi.org/10.1073/pnas.1620973114
Parent AT, Thinakaran G (2010) Modeling presenilin-dependent familial Alzheimer’s disease: emphasis on presenilin substrate-mediated signaling and synaptic function. Int J Alzheimers Dis. https://doi.org/10.4061/2010/825918
Parsons HT, Christiansen K, Knierim B et al (2012) Isolation and proteomic characterization of the Arabidopsis Golgi defines functional and novel components involved in plant cell wall biosynthesis. Plant Physiol 159:12–26. https://doi.org/10.1104/pp.111.193151
Ponting CP, Hutton M, Nyborg A et al (2002) Identification of a novel family of presenilin homologues. Hum Mol Genet 11:1037–1044
Robatzek S, Chinchilla D (2006) Ligand-induced endocytosis of the pattern recognition receptor FLS2 in Arabidopsis. Genes Dev. https://doi.org/10.1101/gad.366506.nized
Sanseverino W, Ercolano MR (2012) In silico approach to predict candidate R proteins and to define their domain architecture. BMC Res Notes 5:678. https://doi.org/10.1186/1756-0500-5-678
Shi Q, Prior M, He W et al (2009) Reduced amyloid deposition in mice overexpressing RTN3 is adversely affected by preformed dystrophic neurites. J Neurosci 29:9163–9173. https://doi.org/10.1523/jneurosci.5741-08.2009
Smith SKF, Anderson H, Yu G et al (2000) Identification of syntaxin 1A as a novel binding protein for presenilin-1. Mol Brain Res 78:100–107. https://doi.org/10.1016/S0169-328X(00)00079-6
Smolarkiewicz M, Skrzypczak T, Wojtaszek P (2013) The very many faces of presenilins and the γ-secretase complex. Protoplasma 250:997–1011. https://doi.org/10.1007/s00709-013-0494-y
Smolarkiewicz M, Skrzypczak T, Michalak M et al (2014) Gamma-secretase subunits associate in intracellular membrane compartments in Arabidopsis thaliana. J Exp Bot. https://doi.org/10.1093/jxb/eru147
Spallek T, Beck M, Ben Khaled S et al (2013) ESCRT-I mediates FLS2 endosomal sorting and plant immunity. PLoS Genet. https://doi.org/10.1371/journal.pgen.1004035
Sparkes I, Tolley N, Aller I et al (2010) Five Arabidopsis reticulon isoforms share endoplasmic reticulum location, topology, and membrane-shaping properties. Plant Cell 22:1333–1343. https://doi.org/10.1105/tpc.110.074385
Stanga S, Vrancx C, Tasiaux B et al (2017) Specificity of presenilin-1- and presenilin-2-dependent γ-secretases towards substrate processing. J Cell Mol Med XX:1–11. https://doi.org/10.1111/jcmm.13364
Suga K, Tomiyama T, Mori H, Akagawa K (2004) Syntaxin 5 interacts with presenilin holoproteins, but not with their N- or C-terminal fragments, and affects beta-amyloid peptide production. Biochem J 381:619–628. https://doi.org/10.1042/BJ20040618
Suga K, Saito A, Mishima T, Akagawa K (2015) ER and Golgi stresses increase ER-Golgi SNARE Syntaxin5: implications for organelle stress and βAPP processing. Neurosci Lett 604:30–35. https://doi.org/10.1016/j.neulet.2015.07.017
Sun L, Zhao L, Yang G et al (2015) Structural basis of human γ-secretase assembly. Proc Natl Acad Sci 112:6003–6008. https://doi.org/10.1073/pnas.1506242112
Swiderski MR, Birker D, Jones JDG et al (2009) The TIR domain of TIR-NB-LRR resistance proteins is a signaling domain involved in cell death induction. Mol Plant Microbe Interact 22:157–165
Tolley N, Sparkes I, Craddock CP et al (2010) Transmembrane domain length is responsible for the ability of a plant reticulon to shape endoplasmic reticulum tubules in vivo. Plant J 64:411–418. https://doi.org/10.1111/j.1365-313X.2010.04337.x
Torres-Arancivia C, Ross CM, Chavez J et al (2010) Identification of an archaeal presenilin-like intramembrane protease. PLoS One. https://doi.org/10.1371/journal.pone.0013072
Van Leene J, Stals H, Eeckhout D et al (2007) A tandem affinity purification-based technology platform to study the cell cycle interactome in Arabidopsis thaliana. Mol Cell Proteomics 6:1226–1238. https://doi.org/10.1074/mcp.M700078-MCP200
Van Leene J, Witters E, Inzé D, De Jaeger G (2008) Boosting tandem affinity purification of plant protein complexes. Trends Plant Sci 13:517–520. https://doi.org/10.1016/j.tplants.2008.08.002
Verena Kriechbaumer V, Botchway SW, Slade SE et al (2015) Reticulomics: Protein-protein interaction studies with two plasmodesmata-localised reticulon family proteins identify binding partners enriched at plasmodesmata, ER and the plasma membrane. Plant Physiol. https://doi.org/10.1104/pp.15.01153
Walker R (2009) Presenilin complexes in arabidopsis: novel plant cell-signalling components? PhD Thesis, University of Edinburgh. http://hdl.handle.net/1842/4090
Wężyk M, Żekanowski C (2017) Presenilins interactome in alzheimer’s disease and pathological ageing. SENESCENCE, InTechOpen, Book. https://doi.org/10.5772/intechopen.68748
Yonemura Y, Futai E, Yagishita S et al (2016) Specific combinations of presenilins and Aph1s affect the substrate specificity and activity of γ-secretase. Biochem Biophys Res Commun 478:1751–1757. https://doi.org/10.1016/j.bbrc.2016.09.018
Zhang H, Sun S, Herreman a et al (2010) Role of presenilins in neuronal calcium homeostasis. J Neurosci 30:8566–8580
Zhang X, Bernoux M, Bentham AR et al (2017) Multiple functional self-association interfaces in plant TIR domains. Proc Natl Acad Sci 114:E2046–E2052. https://doi.org/10.1073/pnas.1621248114
Zhao T, Rui L, Li J et al (2015) A Truncated NLR protein, TIR-NBS2, is required for activated defense responses in the exo70B1 mutant. PLOS Genet 11:e1004945. https://doi.org/10.1371/journal.pgen.1004945
Acknowledgements
RTNLB1 pCAMBIA 1300 was kindly provided by Prof. Chris Hawes (Oxford Brookes University, UK). FLS2 pCAMBIA 2300 was kindly provided by Prof. Silke Robatzek (The Sainsbury Laboratory, UK). TN 10, TN11, TN21, and TX21 cloned in pEarly Gate 103 were kindly provided by Prof. Blake Meyers (Delaware Biotechnology Institute, USA).
Funding
This research was funded by National Science Centre grants (2013/09/N/NZ3/00532, 2012/05/B/NZ3/01026, 2014/15/NZ3/00863) and the “flora robotica” project funded by EU Horizon 2020, FET Grant agreement no. 640959.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by Z. Zhang.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Skrzypczak, T., Krela, R., Wadurkar, S. et al. Characterization of the γ-secretase subunit interactome in Arabidopsis thaliana. Acta Physiol Plant 41, 19 (2019). https://doi.org/10.1007/s11738-019-2811-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-019-2811-3