Abstract
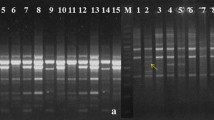
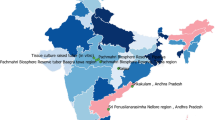
Commiphora wightii (Arn.) Bhandari is a commercially, medicinally and traditionally important tropical shrub widely used to treat various ailments and disorders. Demand of this plant is increasing in the pharmaceutical and perfumery industries due to the presence of guggulsterone E and Z, two important isomers conferring lipid- and cholesterol-lowering, and anti-cancerous properties. Ruthless and u nscientific harvesting of oleo-gum resin by local populations from the wild, with negligible conservation efforts has made this species endangered and led to its inclusion in the Red Data Book of IUCN. It is imperative to have broad information regarding the extent of genetic variability available in the species to accelerate the breeding and conservation programs. Therefore, the present study was undertaken to analyze the extent of genetic variability existing among the C. wightii germplasm collected from Rajasthan and Haryana, the diversity rich Indian states, using ISSR and RAPD markers. A total of 100 (50 each) RAPD and ISSR markers were screened of which 37 RAPD and 43 ISSR primers were able to amplify DNA fragments. RAPD markers were more efficient, detecting 74.16 % polymorphism, compared to ISSR which detected 62.52 % polymorphism. Also, the values of average number of polymorphic bands per assay, polymorphism information content (PIC), diversity index (DI) and marker index (MI) were more for RAPD (7.76, 0.19, 0.38 and 2.53, respectively) than for ISSR (7.02, 0.13, 0.32 and 1.88) markers. The UPGMA dendrogram constructed using individual as well as combined data of the two marker systems separated the collected accessions into two major clusters containing 47 and 4 accessions, respectively, while one accession from Bikaner was not included in any cluster. Genetic similarity values obtained from Jaccard’s coefficient using combined data of both the marker systems were between 0.50 and 0.97. These results indicated the existence of wide genetic variability within this species and can be used for further research in the area of germplasm conservation, population genetics and plant breeding.





Similar content being viewed by others
References
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Ghislain MD, Fazardo ZD, Huaman Z, Hismans RH (1999) Marker assisted sampling of the cultivated andean potato (Solanum fureja) collection using RAPD markers. Genet Reosour Crop Evol 46:547–555
Goulao L, Valdiviesso T, Santana C, Oliveira CM (2001) Comparison between phenotypic characterization using RAPD and ISSR markers and phenotypic data of cultivated chestnut (Castanea sativa Mill.). Genet Resourc Crop Evol 48:329–338
Gupta M, Chyi YS, Romero-Severson J, Owen JL (1994) Amplification of DNA markers from evolutionary diverse genomes using single primers of simple-sequence repeats. Theor Appl Genet 89:998–1006
Gupta S, Srivastava M, Mishra G, Naik P, Chauhan R, Tiwari S (2008) Analogy of ISSR and RAPD markers for comparative analysis of genetic diversity among different Jatropha curcus genotypes. Afr J Biotechnol 7:4230–4243
Haque I, Bandopadhyay R, Mukhopadhyay K (2009) Intraspecific variation in Commiphora wightii populations based on internal transcribed spacer (ITS1-5.8S-ITS2) sequences of rDNA. Diversity 1:89–101
IUCN (2011) IUCN red list of threatened species. Version 2011.1. www.iucnredlist.org
Kant T, Prajapati S, Parmar A (2010) Efficient micropropagation from cotyledonary node cultures of Commiphora wightii (Arn.) Bhandari, an endangered medicinally important desert plant. J Plant Dev 17:37–48
Kulhari A, Sheorayan A, Kalia S, Chaudhury A, Kalia RK (2012) Problems, progress and future prospects of improvement of Commiphora wightii (Arn.) Bhandari, an endangered herbal magic, through modern biotechnological tools: a review. Trees Struct Funct 59:1223–1254
Lal H, Kasera KP (2010) Status and distribution range of Guggul: a critically endangered medicinal plant from the Indian Thar Desert. Sci Cult 76:531–533
Mattioni C, Casasoli M, Gonzalez M, Ipinza R, Villani F (2002) Comparison of ISSR and RAPD markers to characterize three Chilean Nothofagus species. Theor Appl Genet 104:1064–1070
Powell WM, Morgante M, Andre C, Hanafey M, Vogel J, Tingey S, Rafalski A (1996) The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol Breed 2:225–238
Prevost A, Wilkinson MJ (1999) A new system of comparing PCR primers applied to ISSR fingerprinting of potato cultivars. Theor Appl Genet 98:107–112
Rohlf FJ (2000) NTSYS-pc: numerical taxonomy and multivariate analysis system. Version 2.0 Exeter Software, Setauket, New York, USA
Roldan-Ruiz I, Calsyn E, Gilliland TJ, Coll R, Van Eijk MJT, De Loose M (2000) Estimating genetic conformity between related ryegrass (Lolium) varieties. 2. AFLP characterization. Mol Breed 6:593–602
Sankar AA, Moore GA (2001) Evaluation of inter-simple sequence repeat analysis for mapping in Citrus and extension of genetic linkage map. Theor Appl Genet 102:206–214
Satyavati GV, Dwarakanath C, Tripath SN (1969) Experimental studies on the hypocholesterolemic effect of Commiphora mukul Engl. (Guggul). Indian J Med Res 57:1950–1962
Soni V (2010) Conservation of Commiphora wightii, an endangered medicinal shrub, through propagation and planting, and education awareness programs in the Aravali Hills of Rajasthan, India. Conserv Evid 7:27–31
Suthar S, Thul S, Kukreja AK, Ramawat KG (2008) RAPD markers reveal polymorphism in Commiphora wightii, an endangered medicinal tree. J Cell Tiss Res 8:1477–1480
Tsumara Y, Ohba K, Strauss SH (1996) Diversity and inheritance of inter-simple sequence repeat polymorphisms in Douglas-fir (Pseudotsuga menziesii) and sugi (Cryptomeria japonica). Theor Appl Genet 92:40–45
Wang GR, Mahalingan R, Knap HT (1998) (C-A) and (G-A) anchored simple sequence repeats (ASSRs) generated polymorphism in soybean, Glycine max (L.) Merr. Theor Appl Genet 96:1086–1096
Weir BS (1996) Genetic data analysis II: Methods for discrete population genetic data. Sinauer Publisher, Sunderland
Zietkiewicz E, Rafalski A, Labuda D (1994) Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 20:176–183
Acknowledgments
AK thankfully acknowledges the financial assistance provided by DBT, Government of India, New Delhi, under the project sanctioned vide order no- BT/PR 10526/NDB/51/164/2008.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Chandra.
Rights and permissions
About this article
Cite this article
Kulhari, A., Singh, R., Chaudhury, A. et al. Assessment of genetic variability through ISSR and RAPD markers in Commiphora wightii (Arn.) Bhandari. Acta Physiol Plant 37, 113 (2015). https://doi.org/10.1007/s11738-015-1859-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-015-1859-y