Abstract
Quinoxaline-based compounds show promising inhibition of protoporphyrinogen oxidase (PPO), a key enzyme in chlorophyll production and a prime target for herbicide development. However, their precise molecular interactions remain largely unexplored. This study investigates 1-propyl-3-phenylquinoxalin-2(1H)-one (Qnz), a novel quinoxaline derivative designed and synthesized as a representative system. We examined its interaction, at the atomic level, with the active site of Nicotiana tabacum PPO (NtPPO) using an integrated molecular docking and DFT-based quantum mechanics cluster approach. The nature and strength of observed noncovalent interactions were theoretically evaluated using quantum theory of atoms-in-molecules (QTAIM), independent gradient model (IGM), and natural bond orbital (NBO) methods. Our analysis revealed a fascinating set of unconventional interactions that contribute to the stability of the Qnz–NtPPO complex, namely weak hydrogen bonding, homopolar dihydrogen interactions, π–stacking, and carbonyl–carbonyl interactions. Interestingly, the study uncovered the unexpected role of several less common amino acids—including Gly178, Ser235, and nonpolar aliphatic leucines—in facilitating molecular recognition. Moreover, the employed computational approaches have proven to be a powerful tool for analyzing interactions within the binding pocket.
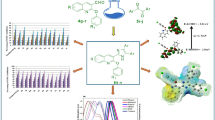
Graphical abstract











Similar content being viewed by others
References
Abad N, Guelmami L, Haouas A, Hajji M, El Hafi M, Sebhaoui J, Guerfel T, Mague JT, Essassi EM, Ramli Y (2023) Synthesis, non-covalent interactions and chemical reactivity of 1-pentyl-3-phenylquinoxalin-2(1H)-one—Structural and computational studies. J Mol Struct 1286:135622. https://doi.org/10.1016/J.MOLSTRUC.2023.135622
Adasme MF, Linnemann KL, Bolz SN, Kaiser F, Salentin S, Haupt VJ, Schroeder M (2021) PLIP 2021: expanding the scope of the protein–ligand interaction profiler to DNA and RNA. Nucleic Acids Res 49:W530–W534. https://doi.org/10.1093/nar/gkab294
Alanazi MA, Arafa WAA, Althobaiti IO, Altaleb HA, Bakr RB, Elkanzi NAA (2022) Green design, synthesis, and molecular docking study of novel quinoxaline derivatives with insecticidal potential against Aphis craccivora. ACS Omega 7:27674–27689. https://doi.org/10.1021/acsomega.2c03332
Avram S, Funar-Timofei S, Borota A, Chennamaneni SR, Manchala AK, Muresan S (2014) Quantitative estimation of pesticide-likeness for agrochemical discovery. J Cheminform 6:42. https://doi.org/10.1186/s13321-014-0042-6
Bauzá A, Frontera A (2022) Noncovalent interactions involving Group 6 in biological systems: the case of molybdopterin and tungstopterin cofactors. Chem Eur J 28:e202201660. https://doi.org/10.1002/chem.202201660
Blomberg MRA, Borowski T, Himo F, Liao R-Z, Siegbahn PEM (2014) Quantum chemical studies of mechanisms for metalloenzymes. Chem Rev 114:3601–3658. https://doi.org/10.1021/cr400388t
Boys SF, Bernardi F (1970) The calculation of small molecular interactions by the differences of separate total energies. some procedures with reduced errors. Mol Phys 19:553–566. https://doi.org/10.1080/00268977000101561
Briceño-Vargas FM, Quesadas-Rojas M, Mirón-López G, Cáceres-Castillo D, Carballo RM, Mena-Rejón GJ, Quijano-Quiñones RF (2023) Molecular orbital and topological electron density study of n → π* interactions: amides and thioamides cases. RSC Adv 13:31321–31329. https://doi.org/10.1039/D3RA06038A
Dayan FE, Duke SO (2002) Herbicides, protoporphyrinogen oxidase inhibitors. encyclopedia of agrochemicals. Wiley, New York. https://doi.org/10.1002/047126363X.agr204
Dennington R, Keith TA, Millam JM (2016) GaussView Version 6. Semichem Inc., Shawnee Mission KS
Eberhardt J, Santos-Martins D, Tillack AF, Forli S (2021) AutoDock Vina 1.2.0: new docking methods, expanded force field, and python bindings. J Chem Inf Model 61:3891–3898. https://doi.org/10.1021/acs.jcim.1c00203
Fernández Riveras JA, Frontera A, Bauzá A (2021) Selenium chalcogen bonds are involved in protein–carbohydrate recognition: a combined PDB and theoretical study. Phys Chem Chem Phys 23:17656–17662. https://doi.org/10.1039/D1CP01929E
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H, Caricato M, Li X, Hratchian HP, Izmaylov AF, Bloino J, Zheng G, Sonnenberg JL, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Montgomery J, Peralta JE, Ogliaro F, Bearpark M, Heyd JJ, Brothers E, Kudin KN, Staroverov VN, Kobayashi R, Normand J, Raghavachari K, Rendell A, Burant JC, Iyengar SS, Tomasi J, Cossi M, Rega N, Millam JM, Klene M, Knox JE, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Martin RL, Morokuma K, Zakrzewski VG, Voth GA, Salvador P, Dannenberg JJ, Dapprich S, Daniels AD, Farkas O, Foresman JB, Ortiz JV, Cioslowski J, Fox DJ (2009) Gaussian 09, Revision D01. Gaussian Inc., Wallingford
García-Gutiérrez P, Zubillaga RA, Ibarra IA, Martínez A, Vargas R, Garza J (2021) Non-conventional interactions of N3 inhibitor with the main protease of SARS-CoV and SARS-CoV-2. Comput Struct Biotechnol J 19:4669–4675. https://doi.org/10.1016/j.csbj.2021.08.015
Grabowski SJ (2019) A−H…σ hydrogen bonds: dihydrogen and cycloalkanes as proton acceptors. ChemPhysChem 20:565–574. https://doi.org/10.1002/cphc.201900045
Grimme S, Antony J, Ehrlich S, Krieg H (2010) A consistent and accurate ab initio parametrization of density functional dispersion correction (DFT-D) for the 94 elements H-Pu. J Chem Phys 132:154104. https://doi.org/10.1063/1.3382344
Hajji M, Abad N, Habib MA, Elmgirhi SMH, Guerfel T (2021a) Computational chemistry methods for modelling non-covalent interactions and chemical reactivity—an overview. J Indian Chem Soc 98:100208. https://doi.org/10.1016/J.JICS.2021.100208
Hajji M, Al-Otaibi JS, Belkhiria M, Dhifaoui S, Habib MA, Elmgirhi SMH, Mtiraoui H, Bel-Hadj-Tahar R, Msaddek M, Guerfel T (2021b) Structural and computational analyses of a 2-propanolammonium-chlorocadmate(II) assembly: pivotal role of hydrogen bonding and H–H interactions. J Mol Struct 1223:128998. https://doi.org/10.1016/j.molstruc.2020.128998
Hajji M, Haouas A, Abad N, Guerfel T (2023) The unconventional noncovalent interactions control: crystallographic and theoretical analyses of the crystalline structure of 1,1′-(1-Chloro-4-methoxyphenyl)dibenzene as a Case Study. ChemistrySelect 8:e202302624. https://doi.org/10.1002/slct.202302624
Hao G-F, Zuo Y, Yang S-G, Yang G-F (2011) Protoporphyrinogen oxidase inhibitor: an ideal target for herbicide discovery. Chimia (Aarau) 65:961. https://doi.org/10.2533/chimia.2011.961
Hao G-F, Tan Y, Yang S-G, Wang Z-F, Zhan C-G, Xi Z, Yang G-F (2013) Computational and experimental insights into the mechanism of substrate recognition and feedback inhibition of protoporphyrinogen oxidase. PLoS ONE 8:e69198. https://doi.org/10.1371/journal.pone.0069198
Hou Y, Bai Y, Lu C, Wang Q, Wang Z, Gao J, Xu H (2023) Applying molecular docking to pesticides. Pest Manag Sci. https://doi.org/10.1002/ps.7700
Humphrey W, Dalke A, Schulten K (1996) VMD: Visual molecular dynamics. J Mol Graph 14:33–38. https://doi.org/10.1016/0263-7855(96)00018-5
Jacobs JM, Jacobs NJ (1984) Protoporphyrinogen oxidation, an enzymatic step in heme and chlorophyll synthesis: partial characterization of the reaction in plant organelles and comparison with mammalian and bacterial systems. Arch Biochem Biophys 229:312–319. https://doi.org/10.1016/0003-9861(84)90157-7
Jena S, Dutta J, Tulsiyan KD, Sahu AK, Choudhury SS, Biswal HS (2022) Noncovalent interactions in proteins and nucleic acids: beyond hydrogen bonding and π-stacking. Chem Soc Rev 51:4261–4286. https://doi.org/10.1039/D2CS00133K
Koch M, Breithaupt C, Kiefersauer R, Freigang J, Huber R, Messerschmidt A (2004) Crystal structure of protoporphyrinogen IX oxidase: a key enzyme in haem and chlorophyll biosynthesis. EMBO J 23:1720–1728. https://doi.org/10.1038/sj.emboj.7600189
Laskowski RA, Swindells MB (2011) LigPlot+: multiple ligand-protein interaction diagrams for drug discovery. J Chem Inf Model 51:2778–2786. https://doi.org/10.1021/ci200227u
Lefebvre C, Khartabil H, Boisson J, Contreras-García J, Piquemal J, Hénon E (2018) The independent gradient model: a new approach for probing strong and weak interactions in molecules from wave function calculations. ChemPhysChem 19:724–735. https://doi.org/10.1002/cphc.201701325
Liu X-H, Yu W, Min L-J, Wedge DE, Tan C-X, Weng J-Q, Wu H-K, Cantrell CL, Bajsa-Hirschel J, Hua X-W, Duke SO (2020) Synthesis and pesticidal activities of new quinoxalines. J Agric Food Chem 68:7324–7332. https://doi.org/10.1021/acs.jafc.0c01042
Liu H-Y, Yu L-K, Qin S-N, Yang H-Z, Wang D-W, Xi Z (2023) Design, synthesis, and metabolism studies of N-1,4-diketophenyltriazinones as protoporphyrinogen IX oxidase inhibitors. J Agric Food Chem 71(7):3225–3238. https://doi.org/10.1021/acs.jafc.2c09082
Losev TV, Gerasimov IS, Panova MV, Lisov AA, Abdyusheva YR, Rusina PV, Zaletskaya E, Stroganov OV, Medvedev MG, Novikov FN (2023) Quantum mechanical-cluster approach to solve the bioisosteric replacement problem in drug design. J Chem Inf Model 63:1239–1248. https://doi.org/10.1021/acs.jcim.2c01212
Lu T, Chen F (2012) Multiwfn: a multifunctional wavefunction analyzer. J Comput Chem 33:580–592. https://doi.org/10.1002/jcc.22885
Mandal N, Pratik SM, Datta A (2017) Exploring ultrashort hydrogen-hydrogen nonbonded contacts in constrained molecular cavities. J Phys Chem B 121:825–834. https://doi.org/10.1021/acs.jpcb.6b12391
Martínez A, García-Gutiérrez P, Zubillaga RA, Garza J, Vargas R (2021) Main interactions of dopamine and risperidone with the dopamine D2 receptor. Phys Chem Chem Phys 23:14224–14230. https://doi.org/10.1039/D1CP01637G
Mei L-C, Chen H-M, Dong A-Y, Huang G-Y, Liu Y-W, Zhang X, Wang W, Hao G-F, Yang G-F (2022) Pesticide informatics platform (PIP): an international platform for pesticide discovery, residue, and risk evaluation. J Agric Food Chem 70:6617–6623. https://doi.org/10.1021/acs.jafc.2c02141
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791. https://doi.org/10.1002/jcc.21256
Muchowska KB, Pascoe DJ, Borsley S, Smolyar IV, Mati IK, Adam C, Ling NGS, Cockroft SL (2020) Reconciling electrostatic and n→π* orbital contributions in carbonyl interactions. Angew Chem 132:14710–14716. https://doi.org/10.1002/ange.202005739
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612. https://doi.org/10.1002/jcc.20084
Qin X, Sun L, Wen X, Yang X, Tan Y, Jin H, Cao Q, Zhou W, Xi Z, Shen Y (2010) Structural insight into unique properties of protoporphyrinogen oxidase from Bacillus subtilis. J Struct Biol 170:76–82. https://doi.org/10.1016/j.jsb.2009.11.012
Rahim A, Saha P, Jha KK, Sukumar N, Sarma BK (2017) Reciprocal carbonyl–carbonyl interactions in small molecules and proteins. Nat Commun 8:78. https://doi.org/10.1038/s41467-017-00081-x
Rocha-Santos A, Chaves EJF, Grillo IB, de Freitas AS, Araújo DAM, Rocha GB (2021) Thermochemical and quantum descriptor calculations for gaining insight into ricin toxin A (RTA) Inhibitors. ACS Omega 6:8764–8777. https://doi.org/10.1021/acsomega.0c02588
Schrödinger LLC (2021) The PyMOL molecular graphics system. Version 2:5
Shi J, Zhao L, Wang J, Cao H, Gao S, Ye F, Fu Y (2023) Identification of 4-hydroxyphenylpyruvate dioxygenase inhibitors by virtual screening, molecular docking, molecular dynamic simulation. J Sci Food Agric 103:5547–5559. https://doi.org/10.1002/jsfa.12629
Steiner T (1996) C-H-O hydrogen bonding in crystals. Crystallogr Rev 6:1–51. https://doi.org/10.1080/08893119608035394
Sukhoverkov KV, Corral MG, Leroux J, Haywood J, Johnen P, Newton T, Stubbs KA, Mylne JS (2021) Improved herbicide discovery using physico-chemical rules refined by antimalarial library screening. RSC Adv 11:8459–8467. https://doi.org/10.1039/D1RA00914A
Sun Z, Wei C, Wu S, Zhang W, Song R, Hu D (2022) Synthesis, Anti-potato virus Y activities, and interaction mechanisms of novel quinoxaline derivatives bearing dithioacetal moiety. J Agric Food Chem 70:7029–7038. https://doi.org/10.1021/acs.jafc.2c01898
Sun J (2023) PyMOL ChatGPT Plugin. GitHub repository. https://github.com/JinyuanSun/ChatMol.git
Tang X, Zhou Q, Zhan W, Hu D, Zhou R, Sun N, Chen S, Wu W, Xue W (2022) Synthesis of novel antibacterial and antifungal quinoxaline derivatives. RSC Adv 12:2399–2407. https://doi.org/10.1039/D1RA07559D
Tian W, Chen C, Lei X, Zhao J, Liang J (2018) CASTp 3.0: computed atlas of surface topography of proteins. Nucleic Acids Res 46:W363–W367. https://doi.org/10.1093/nar/gky473
Wang Z-W, Zhao L-X, Gao S, Leng X-Y, Yu Y, Fu Y, Ye F (2021b) Quinoxaline derivatives as herbicide safeners by improving Zea mays tolerance. Pestic Biochem Physiol 179:104958. https://doi.org/10.1016/j.pestbp.2021.104958
Wang D, Wang B, Xi Z (2021) Development of Protoporphyrinogen IX oxidase inhibitors for sustainable agriculture. pp. 11–41 in: Crop Protection Products for Sustainable Agriculture. https://doi.org/10.1021/bk-2021-1390.ch002
Wojtkowiak K, Michalczyk M, Zierkiewicz W, Jezierska A, Panek JJ (2022) Chalcogen bond as a factor stabilizing ligand conformation in the binding pocket of carbonic anhydrase IX receptor Mimic. Int J Mol Sci 23:13701. https://doi.org/10.3390/ijms232213701
Zagar C, Liebl R, Theodoridis G, Witschel M (2019) Protoporphyrinogen IX Oxidase Inhibitors. In: Modern crop protection compounds. Weinheim, Germany, pp. 173–211. https://doi.org/10.1002/9783527699261.ch3
Zhao H, Tang S, Zhang Q, Du L (2017) Weak hydrogen bonding competition between O-H⋯π and O–H⋯Cl. RSC Adv 7:22485–22491. https://doi.org/10.1039/C7RA00901A
Zhao L, Peng J, Liu F, Zou Y, Gao S, Fu Y, Ye F (2022a) Discovery of novel phenoxypyridine as promising protoporphyrinogen IX oxidase inhibitors. Pestic Biochem Physiol 184:105102. https://doi.org/10.1016/j.pestbp.2022.105102
Zhao L-X, Peng J-F, Hu J-J, Zou Y-L, Yin M-L, Wang Z-X, Gao S, Fu Y, Ye F (2022b) Design, synthesis, herbicidal activity, and the molecular docking study of novel diphenyl ether derivatives as protoporphyrinogen IX oxidase inhibitors. J Mol Struct 1258:132670. https://doi.org/10.1016/j.molstruc.2022.132670
Zhao W, Huang Y, Hao G-F (2022c) Pesticide informatics expands the opportunity for structure-based molecular design and optimization. Adv Agrochem 1:139–147. https://doi.org/10.1016/j.aac.2022.11.006
Zheng B-F, Zuo Y, Huang G-Y, Wang Z-Z, Ma J-Y, Wu Q-Y, Yang G-F (2023a) Synthesis and biological activity evaluation of benzoxazinone-pyrimidinedione hybrids as potent protoporphyrinogen IX oxidase inhibitor. J Agric Food Chem 71(39):14221–14231. https://doi.org/10.1021/acs.jafc.3c03593
Zheng B-F, Wang Z-Z, Dong J, Ma J-Y, Zuo Y, Wu Q-Y, Yang G-F (2023b) Discovery of a subnanomolar inhibitor of protoporphyrinogen IX oxidase via fragment-based virtual screening. J Agric Food Chem 71(23):8746–8756. https://doi.org/10.1021/acs.jafc.3c00168
Acknowledgements
The authors extend their appreciation to Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2024R95), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia. The author, Melek Hajji, acknowledges the invaluable support and resources provided by Assalam International University-Sirte, throughout this research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hajji, M., Abad, N., Dallel, M. et al. Noncovalent interactions between quinoxalines and protoporphyrinogen oxidase (PPO): a computational case study for herbicidal applications. Chem. Pap. (2024). https://doi.org/10.1007/s11696-024-03485-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11696-024-03485-4