Abstract
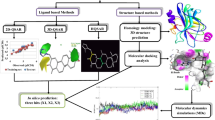
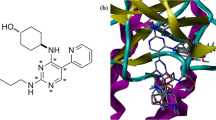
The pivotal role of MRP3 protein in acute leukaemia and the efficacy of natural compounds in cancer treatment have necessitated the current study to identify novel MRP3 inhibitors from natural source. The MRP3 protein was modelled and validated using well-accepted metrics, after which a validated multiple-ligand pharmacophore model (AHHHR_4) was built to screen natural compounds (n = 47,964). The combined pharmacophore screening with molecular docking was conducted to identify the hits drug-like compounds using ADMET profiling. The electronic behaviour of this set of compounds in gas phase was examined using density functional theory. Among the compounds (n = 7) with clean ADMET profile, NPC5486, which possessed the highest binding affinity with MRP3, was further subjected to 50 ns molecular dynamics (MD) simulation to understand its dynamics of binding. Analysis from the resulting MD simulation trajectories of NPC5486 in complex with the model protein alongside that of standard inhibitor (vincristine) showed not only the flexibility and interaction potential of the residues of the MRP3 with NPC5486 as indicated by the RMSF but also stability of the complex as indicated by the RMSD, RoG and number of hydrogen bonds of the ligand–protein complexes. Cluster analysis of the MD simulation trajectory files and dynamics-based MMGBSA computations further revealed that the observed interactions with important residues as well as the free energy contribution per residue were preserved in the dynamic environment. Overall, the current study has revealed drug-like compounds which can serve as potential inhibitors of MRP3 in the treatment of acute leukaemia.















Similar content being viewed by others
References
Akioka K, Masuda K, Harada S, Nakamura T, Okugawa K, Nakano K, Osaka Y, Tsuchiya K, Sako H (2013) Acute renal failure caused by hyperuremic acidemia in ABO-incompatible kidney transplant maintained with cyclosporine and high-dose mizoribine: A case report. Transplant Proc 45(7):2815–2818. https://doi.org/10.1016/j.transproceed.2013.03.052
Ali I, Welch MA, Lu Y, Swaan PW, Brouwer KLR (2017) Identification of novel MRP3 inhibitors based on computational models and validation using an in vitro membrane vesicle assay. Eur J Pharm Sci 103:52–59. https://doi.org/10.1016/j.ejps.2017.02.011
Alyar S, Şen T, Özmen ÜÖ, Alyar H, Adem Ş, Şen C (2019) Synthesis, spectroscopic characterizations, enzyme inhibition, molecular docking study and DFT calculations of new Schiff bases of sulfa drugs. J Mol Struct 1185:416–424
Banerjee P, Eckert AO, Schrey AK, Preissner R (2018) ProTox-II: a webserver for the prediction of toxicity of chemicals. Nucleic Acids Res 46(W1):W257–W263. https://doi.org/10.1093/nar/gky318
Beck TC, Beck KR, Morningstar J, Benjamin MM, Norris RA (2021) Descriptors of cytochrome inhibitors and useful machine learning based methods for the design of safer drugs. Pharmaceuticals 14(5):472
Becke AD (1988) Density-functional exchange-energy approximation with correct asymptotic behavior. Phys Rev A Gen Phys 38(6):3098–3100. https://doi.org/10.1103/physreva.38.3098
Benkert P, Tosatto SCE, Schomburg D (2008) QMEAN: A comprehensive scoring function for model quality assessment. Proteins 71(1):261–277. https://doi.org/10.1002/prot.21715
Berjanskii M, Zhou J, Liang Y, Lin G, Wishart DS (2012) Resolution-by-proxy: a simple measure for assessing and comparing the overall quality of NMR protein structures. J Biomol NMR 53(3):167–180
Bobrowska-Hägerstrand M, Lillås M, Mrówczyñska L, Wróbel A, Shirataki Y, Motohashi N, Hägerstrand H (2006) Resveratrol oligomers are potent MRP1 transport inhibitors. Anticancer Res 26(3A):2081–2084
Bochevarov AD, Harder E, Hughes TF, Greenwood JR, Braden DA, Philipp DM, Rinaldo D, Halls MD, Zhang J, Friesner RA (2013) Jaguar: A high-performance quantum chemistry software program with strengths in life and materials sciences. Int J Quantum Chem 113(18):2110–2142
Borst P, Evers R, Kool M, Wijnholds J (2000) A family of drug transporters: The multidrug resistance-associated proteins. J Natl Cancer Inst 92(16):1295–1302. https://doi.org/10.1093/jnci/92.16.1295
Brooks BR, Brooks CL III, Mackerell AD Jr, Nilsson L, Petrella RJ, Roux B, Won Y, Archontis G, Bartels C, Boresch S (2009) CHARMM: the biomolecular simulation program. J Comput Chem 30(10):1545–1614
Case DA, Berryman J, Betz RM, Cerutti DS, Cheatham Iii TE, Darden TA, Duke RE, Giese TJ, Gohlke H, and Goetz AW (2015) AMBER 2015
Colovos C, Yeates TO (1993) Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci 2(9):1511–1519
Dalton JAR, Jackson RM (2007) An evaluation of automated homology modelling methods at low target–template sequence similarity. Bioinformatics 23(15):1901–1908
DeLano WL (2002) Pymol: An open-source molecular graphics tool. CCP4 Newsl Protein Crystallogr 40(1):82–92
De Vocht T, Buyck C, Deferm N, Qi B, Van Brantegem P, van Vlijmen H, Snoeys J, Hoeben E, Vermeulen A, Annaert P (2021) Identification of novel inhibitors of rat Mrp3. Eur J Pharm Sci 162:105813. https://doi.org/10.1016/j.ejps.2021.105813
Elekofehinti OO, Iwaloye O, Olawale F, Chukwuemeka PO, Folorunso IM (2021) Newly designed compounds from scaffolds of known actives as inhibitors of survivin: computational analysis from the perspective of fragment-based drug design. Silico Pharmacol 9(1):47. https://doi.org/10.1007/s40203-021-00108-8
Fletcher JI, Williams RT, Henderson MJ, Norris MD, Haber M (2016) ABC transporters as mediators of drug resistance and contributors to cancer cell biology. Drug Resist Updat 26:1–9. https://doi.org/10.1016/j.drup.2016.03.001
Fukushima-Uesaka H, Saito Y, Maekawa K, Hasegawa R, Suzuki K, Yanagawa T, Kajio H, Kuzuya N, Noda M, Yasuda K, Tohkin M, Sawada JI (2007) Genetic variations of the ABC transporter gene ABCC3 in a Japanese population. Drug Metab Pharmacokinet 22(2):129–135. https://doi.org/10.2133/dmpk.22.129
Ghose AK, Viswanadhan VN, Wendoloski JJ (1998) Prediction of hydrophobic (lipophilic) properties of small organic molecules using fragmental methods: an analysis of ALOGP and CLOGP methods. J Phys Chem A 102(21):3762–3772
Gilibili RR, Chatterjee S, Bagul P, Mosure KW, Murali BV, Mariappan TT, Mandlekar S, Lai Y (2017) Coproporphyrin-I: A fluorescent, endogenous optimal probe substrate for ABCC2 (MRP2) suitable for vesicle-based MRP2 inhibition assay. Drug Metab Dispos 45(6):604–611. https://doi.org/10.1124/dmd.116.074740
Hagar M, Ahmed HA, Aljohani G, Alhaddad OA (2020) Investigation of some antiviral N-heterocycles as COVID 19 drug: molecular docking and DFT calculations. Int J Mol Sci 21(11):3922
Halgren TA (2009) Identifying and characterizing binding sites and assessing druggability. J Chem Inf Model 49(2):377–389. https://doi.org/10.1021/ci800324m
Hollenstein K, Dawson RJP, Locher KP (2007) Structure and mechanism of ABC transporter proteins. Curr Opin Struct Biol 17(4):412–418
Howlader NNA, Krapcho M, Miller D, Bishop K, Kosary CL, and Yu M (2017) Cronin KA (eds). SEER Cancer Statistics Review, 1975–2014, National Cancer Institute. Bethesda, MD
Huang J, MacKerell ADJ (2013) CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J Comput Chem 34(25):2135–2145. https://doi.org/10.1002/jcc.23354
Iwaloye O, Elekofehinti OO, Momoh AI, Babatomiwa K, Ariyo EO (2020) In silico molecular studies of natural compounds as possible anti-Alzheimer’s agents: ligand-based design. Netw Model Anal Heal Informatics Bioinforma. https://doi.org/10.1007/s13721-020-00262-7
Jiang L, He Y, Luo G, Yang Y, Li G, Zhang Y (2016) Discovery of potential novel microsomal triglyceride transfer protein inhibitors via virtual screening of pharmacophore modelling and molecular docking. Mol Simul 42(15):1223–1232
Jo S, Kim T, Iyer VG, Im W (2008) CHARMM-GUI: a web-based graphical user interface for CHARMM. J Comput Chem 29(11):1859–1865
Khalid M, Ullah MA, Adeel M, Khan MU, Tahir MN, Braga AAC (2019) Synthesis, crystal structure analysis, spectral IR, UV–Vis, NMR assessments, electronic and nonlinear optical properties of potent quinoline based derivatives: interplay of experimental and DFT study. J Saudi Chem Soc 23(5):546–560
Köck K, Ferslew BC, Netterberg I, Yang K, Urban TJ, Swaan PW, Stewart PW, Brouwer KLR (2014) Risk factors for development of cholestatic drug-induced liver injury: inhibition of hepatic basolateral bile acid transporters multidrug resistance-associated proteins 3 and 4. Drug Metab Dispos 42(4):665–674
Lauria A, Ippolito M, Fazzari M, Tutone M, Di Blasi F, Mingoia F, Almerico AM (2010) IKK-β inhibitors: An analysis of drug–receptor interaction by using molecular docking and pharmacophore 3D-QSAR approaches. J Mol Graph Model 29(1):72–81
Le M-T, Hoang V-N, Nguyen D-N, Bui T-H-L, Phan T-V, Huynh PN-H, Tran T-D, Thai K-M (2021) Structure-based discovery of ABCG2 Inhibitors: A homology protein-based pharmacophore modeling and molecular docking approach. Molecules 26(11):3115
Lee C, Yang W, Parr RG (1988) Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Phys Rev B Condens Matter 37(2):785–789. https://doi.org/10.1103/physrevb.37.785
Lee J, Cheng X, Swails JM, Yeom MS, Eastman PK, Lemkul JA, Wei S, Buckner J, Jeong JC, Qi Y, Jo S, Pande VS, Case DA, Brooks CL, MacKerell AD, Klauda JB, Im W (2016) CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J Chem Theory Comput 12(1):405–413. https://doi.org/10.1021/acs.jctc.5b00935
Lyne PD, Lamb ML, Saeh JC (2006) Accurate prediction of the relative potencies of members of a series of kinase inhibitors using molecular docking and MM-GBSA scoring. J Med Chem 49(16):4805–4808
Maynadié M (2015) Acute Leukemia. Rev Francoph Des Lab 2015(471):27–28. https://doi.org/10.1016/s1773-035x(15)30070-8
Miller BR, McGee TD, Swails JM, Homeyer N, Gohlke H, Roitberg AE (2012) MMPBSA.py: An efficient program for end-state free energy calculations. J Chem Theory Comput 8(9):3314–3321. https://doi.org/10.1021/ct300418h
Noroozi-Aghideh A, Safavi A, Ghodousi ES, Rajaeinejad M, Moghaddam PT (2020) Homology modeling and molecular docking of hABCC3/MRP3 with chemotherapeutic agents in acute leukemia. Jundishapur J Nat Pharm Prod. https://doi.org/10.5812/jjnpp.69407
Olawale F, Iwaloye O, Elekofehinti OO (2021a) Virtual screening of natural compounds as selective inhibitors of polo-like kinase-1 at C-terminal polo box and N-terminal catalytic domain. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2021.1991476
Olawale F, Olofinsan K, Iwaloye O, and Ologuntere TE (2021b). Phytochemicals from Nigerian medicinal plants modulate therapeutically-relevant diabetes targets: insight from computational direction. Adv Tradit Med 1–15
Olawale F, Olofinsan K, Iwaloye O, Chukwuemeka PO, Elekofehinti OO (2022) Screening of compounds from Nigerian antidiabetic plants as protein tyrosine phosphatase 1B inhibitor. Comput Toxicol 21:100200. https://doi.org/10.1016/j.comtox.2021.100200
Parthasarathi R, Subramanian V, Roy DR, Chattaraj PK (2004) Electrophilicity index as a possible descriptor of biological activity. Bioorg Med Chem 12(21):5533–5543
Remmert M, Biegert A, Hauser A, Söding J (2011) HHblits: lightning-fast iterative protein sequence searching by HMM-HMM alignment. Nat Methods 9(2):173–175. https://doi.org/10.1038/nmeth.1818
Salentin S, Schreiber S, Haupt VJ, Adasme MF, Schroeder M (2015) PLIP: Fully automated protein-ligand interaction profiler. Nucleic Acids Res 43(W1):W443–W447. https://doi.org/10.1093/nar/gkv315
Saxena S, Abdullah M, Sriram D, Guruprasad L (2018) Discovery of novel inhibitors of Mycobacterium tuberculosis MurG: Homology modelling, structure based pharmacophore, molecular docking, and molecular dynamics simulations. J Biomol Struct Dyn 36(12):3184–3198
Scheffer GL, Kool M, de Haas M, de Vree JML, Pijnenborg ACLM, Bosman DK, Oude Elferink RPJ, Van Der Valk P, Borst P, Scheper RJ (2002) Tissue distribution and induction of human multidrug resistant protein 3. Lab Investig 82(2):193–201. https://doi.org/10.1038/labinvest.3780411
Song XR, Zheng Y, He G, Yang L, Luo YF, He ZY, Li SZ, Li JM, Yu S, Luo X, Hou SX, Wei YQ (2010) Development of PLGA nanoparticles simultaneously loaded with vincristine and verapamil for treatment of hepatocellular carcinoma. J Pharm Sci 99(12):4874–4879. https://doi.org/10.1002/jps.22200
Steinbach D, Legrand O (2007) ABC transporters and drug resistance in leukemia: Was P-gp nothing but the first head of the Hydra? Leukemia 21(6):1172–1176. https://doi.org/10.1038/sj.leu.2404692
Studer G, Tauriello G, Bienert S, Biasini M, Johner N, Schwede T (2021) ProMod3 - A versatile homology modelling toolbox. PLoS Comput Biol 17(1):e1008667. https://doi.org/10.1371/JOURNAL.PCBI.1008667
Tubiana T, Carvaillo J-C, Boulard Y, Bressanelli S (2018) TTClust: a versatile molecular simulation trajectory clustering program with graphical summaries. J Chem Inf Model 58(11):2178–2182
Uchiumi T, Hinoshita E, Haga S, Nakamura T, Tanaka T, Toh S, Furukawa M, Kawabe T, Wada M, Kagotani K, Okumura K, Kohno K, Akiyama SI, Kuwano M (1998) Isolation of a novel human canalicular multispecific organic anion transporter, cMOAT2/MRP3, and its expression in cisplatin-resistant cancer cells with decreased ATP-dependent drug transport. Biochem Biophys Res Commun 252(1):103–110. https://doi.org/10.1006/bbrc.1998.9546
van de Velde ME, Kaspers GL, Abbink FCH, Wilhelm AJ, Ket JCF, van den Berg MH (2017) Vincristine-induced peripheral neuropathy in children with cancer: A systematic review. Crit Rev Oncol Hematol 114:114–130. https://doi.org/10.1016/j.critrevonc.2017.04.004
Van Der Kolk DM, De Vries EGE, Noordhoek L, Van Den Berg E, Van Der Pol MA, Müller M, Vellenga E (2001) Activity and expression of the multidrug resistance proteins P-glycoprotein, MRP1, MRP2, MRP3 and MRP5 in de novo and relapsed acute myeloid leukemia. Leukemia 15(10):1544–1553. https://doi.org/10.1038/sj.leu.2402236
Veber DF, Johnson SR, Cheng H-Y, Smith BR, Ward KW, Kopple KD (2002) Molecular properties that influence the oral bioavailability of drug candidates. J Med Chem 45(12):2615–2623. https://doi.org/10.1021/jm020017n
Waterhouse A, Bertoni M, Bienert S, Studer G, Tauriello G, Gumienny R, Heer FT, de Beer TAP, Rempfer C, Bordoli L, Lepore R, Schwede T (2018) SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res 46(W1):W296–W303. https://doi.org/10.1093/nar/gky427
Willard L, Ranjan A, Zhang H, Monzavi H, Boyko RF, Sykes BD, Wishart DS (2003) VADAR: a web server for quantitative evaluation of protein structure quality. Nucleic Acids Res 31(13):3316–3319
Wu CP, Calcagno AM, Hladky SB, Ambudkar SV, Barrand MA (2005) Modulatory effects of plant phenols on human multidrug-resistance proteins 1, 4 and 5 (ABCC1, 4 and 5). FEBS J 272(18):4725–4740. https://doi.org/10.1111/j.1742-4658.2005.04888.x
Ye ZW, Camus S, Augustijns P, Annaert P (2010) Interaction of eight HIV protease inhibitors with the canalicular efflux transporter ABCC2 (MRP2) in sandwich-cultured rat and human hepatocytes. Biopharm Drug Dispos 31(2–3):178–188. https://doi.org/10.1002/bdd.701
Yu W, He X, Vanommeslaeghe K, MacKerell ADJ (2012) Extension of the CHARMM General Force Field to sulfonyl-containing compounds and its utility in biomolecular simulations. J Comput Chem 33(31):2451–2468. https://doi.org/10.1002/jcc.23067
Zhang YK, Wang YJ, Gupta P, Chen ZS (2015) Multidrug Resistance Proteins (MRPs) and Cancer Therapy. AAPS J 17(4):802–812. https://doi.org/10.1208/s12248-015-9757-1
Acknowledgements
The authors acknowledge the technical support received from Computational Biologists at the Bioinformatics and Molecular Biology Unit, Department of Biochemistry, Federal University of Technology, Akure, Ondo State, Nigeria.
Funding
This research received no external financial or non-financial supports.
Author information
Authors and Affiliations
Contributions
FO helped in project design, generation and analysis of data, interpretation of data, preparation of manuscript draft and approval of manuscript for submission. OI and OO performed data analysis, data interpretation, substantial manuscript revision and approval of manuscript for submission. KO was involved in manuscript draft, substantial manuscript revision and approval of manuscript for submission. II contributed to data acquisition, preparation of manuscript draft and approval of manuscript for submission. GG helped in substantial manuscript revision, project supervision and provision of resources for research and approval of manuscript for submission. All authors have read and approved the manuscript for submission.
Corresponding author
Ethics declarations
Ethical approval
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper. The authors declare the following financial interests/personal relationships which may be considered as potential competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Olawale, F., Iwaloye, O., Olofinsan, K. et al. Homology modelling, vHTS, pharmacophore, molecular docking and molecular dynamics studies for the identification of natural compound-derived inhibitor of MRP3 in acute leukaemia treatment. Chem. Pap. 76, 3729–3757 (2022). https://doi.org/10.1007/s11696-022-02128-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11696-022-02128-w