Abstract
Objective
Gastrointestinal stromal tumors (GISTs) can rapidly proliferate through angiogenesis. Previous studies indicated the potential influence of microRNA on the progression of tumor immature angiogenesis. This study aimed to explore the specific mechanism by which microRNA-409-5p (miR-409-5p) contributes to GIST.
Methods
To identify genes potentially involved in the development and progression of GIST, the differences of miR-409-5p between tumors and adjacent tissues were first analyzed. Following this analysis, target genes were predicted. To further investigate the function of miRNA in GIST cells, two GIST cell lines (GIST-T1 and GIST882) were transfected with lentiviruses that stably expressed miR-409-5p and scrambled miRNA (negative control). Later, the cells were subjected to Western blotting and ELSA to determine any differences in angiogenesis-related genes.
Results
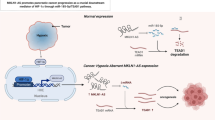
In GISTs, there was a decrease in the expression levels of miR-409-5p compared to the adjacent tissues. It was observed that the upregulation of miR-409-5p in GIST cell lines effectively inhibited the proteins hypoxia-inducible transcription factor 1β (HIF1β) and vascular endothelial growth factor A (VEGF-A). Further investigations revealed that miR-409-5p acted as an inhibitor of angiogenesis by binding to the 3′-UTR of Lysine-specific demethylase 4D (KDM4D) mRNA. Moreover, the combination of miR-409-5p with imatinib enhanced its inhibitory effect on angiogenesis.
Conclusion
This study demonstrated that the miRNA-409-5p/KDM4D/HIF1β/VEGF-A signaling pathway could serve as a novel target for the development of therapeutic strategies for the treatment of imatinib-resistance in GIST patients.
Similar content being viewed by others
References
Corless CL, Barnett CM, Heinrich MC. Gastrointestinal stromal tumours: origin and molecular oncology. Nat Rev Cancer, 2011,11(12):865–878
Miettinen M, Lasota J. Histopathology of gastrointestinal stromal tumor. J Surg Oncol, 2011,104(8):865–873
Joensuu H, Hohenberger P, Corless CL. Gastrointestinal stromal tumour. Lancet, 2013,382(9896):973–983
ESMO/European Sarcoma Network Working Group. Gastrointestinal stromal tumors: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol, 2012,23(Suppl 7):vii49–55
Goettsch WG, Bos SD, Breekveldt-Postma N, et al. Incidence of gastrointestinal stromal tumours is underestimated: results of a nation-wide study. Eur J Cancer, 2005,41(18):2868–2872
Roberts PJ, Eisenberg B. Clinical presentation of gastrointestinal stromal tumors and treatment of operable disease. Eur J Cancer, 2002,38 Suppl 5:S37–S38
Demetri GD, von Mehren M, Blanke CD, et al. Efficacy and safety of imatinib mesylate in advanced gastrointestinal stromal tumors. N Engl J Med, 2002,347(7):472–480
Blanke CD, Demetri GD, von Mehren M, et al. Long-term results from a randomized phase II trial of standard-versus higher-dose imatinib mesylate for patients with unresectable or metastatic gastrointestinal stromal tumors expressing KIT. J Clin Oncol, 2008,26(4):620–625
Nishida T, Shirao K, Sawaki A, et al. Efficacy and safety profile of imatinib mesylate (ST1571) in Japanese patients with advanced gastrointestinal stromal tumors: a phase II study (STI571B1202). Int J Clin Oncol, 2008,13(3):244–251
Joensuu H, Vehtari A, Riihimäki J, et al. Risk of recurrence of gastrointestinal stromal tumour after surgery: an analysis of pooled population-based cohorts. The Lancet Oncology, 2012,13(3):265–274
Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell, 2009,136(2):215–233
Mattick JS. The genetic signatures of noncoding RNAs. PLoS Genet, 2009,5(4):e1000459
Lu J, Getz G, Miska EA, et al. MicroRNA expression profiles classify human cancers. Nature, 2005,435(7043):834–838
Ji J, Shi J, Budhu A, et al. MicroRNA expression, survival, and response to interferon in liver cancer. N Engl J Med, 2009,361(15):1437–1447
Ling H, Fabbri M, Calin GA. MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat Rev Drug Discov, 2013,12(11):847–865
Bader AG. miR-34 - a microRNA replacement therapy is headed to the clinic. Front Genet, 2012,3:120
Bader AG, Brown D, Stoudemire J, Lammers P. Developing therapeutic microRNAs for cancer. Gene Ther, 2011,18(12):1121–1126
Chen L, Zheng J, Zhang Y, et al. Tumor-specific expression of microRNA-26a suppresses human hepatocellular carcinoma growth via cyclin-dependent and -independent pathways. Mol Ther, 2011,19(8):1521–1528
Hartmann P, Zhou Z, Natarelli L, et al. Endothelial Dicer promotes atherosclerosis and vascular inflammation by miRNA-103-mediated suppression of KLF4. Nat Commun, 2016,7:10521
Jeon YJ, Kim T, Park D, et al. miRNA-mediated TUSC3 deficiency enhances UPR and ERAD to promote metastatic potential of NSCLC. Nat Commun, 2018,9(1):5110
Josson S, Gururajan M, Hu P, et al. miR-409-3p/-5p promotes tumorigenesis, epithelial-to-mesenchymal transition, and bone metastasis of human prostate cancer. Clin Cancer Res, 2014,20(17):4636–4646
Prakash R, John AA, Singh D. miR-409-5p negatively regulates Wnt/Beta catenin signaling pathway by targeting Lrp-8. J Cell Physiol, 2019,234(12):23507–23517
Yu H, Xing H, Han W, et al. MicroRNA-409-5p is upregulated in breast cancer and its downregulation inhibits cancer development through downstream target of RSU1. Tumour Biol, 2017,39(5):1010428317701647
de Rinaldis E, Gazinska P, Mera A, et al. Integrated genomic analysis of triple-negative breast cancers reveals novel microRNAs associated with clinical and molecular phenotypes and sheds light on the pathways they control. BMC Genomics, 2013,14:643
Wang G, Yang X, Li C, et al. PIK3R3 induces epithelial-to-mesenchymal transition and promotes metastasis in colorectal cancer. Mol Cancer Ther, 2014,13(7):1837–1847
Lu X, Kang Y. Hypoxia and hypoxia-inducible factors: master regulators of metastasis. Clin Cancer Res, 2010,16(24):5928–5935
Eswarappa SM, Fox PL. Antiangiogenic VEGF-Ax: A New Participant in Tumor Angiogenesis. Cancer Res, 2015,75(14):2765–2769
Goel HL, Mercurio AM. VEGF targets the tumour cell. Nat Rev Cancer, 2013,13(12):871–882
Semenza GL. Defining the role of hypoxia-inducible factor 1 in cancer biology and therapeutics. Oncogene, 2010,29(5):625–634
Kumar MS, Lu J, Mercer KL, et al. Impaired microRNA processing enhances cellular transformation and tumorigenesis. Nat Genet, 2007,39(5):673–677
Garzon R, Marcucci G, Croce CM. Targeting microRNAs in cancer: rationale, strategies and challenges. Nat Rev Drug Discov, 2010,9(10):775–789
Lin S, Gregory RI. MicroRNA biogenesis pathways in cancer. Nat Rev Cancer, 2015,15(6):321–333
Bai R, Weng C, Dong H, et al. MicroRNA-409-3p suppresses colorectal cancer invasion and metastasis partly by targeting GAB1 expression. Int J Cancer, 2015,137(10):2310–2322
Hu F, Li H, Liu L, et al. Histone demethylase KDM4D promotes gastrointestinal stromal tumor progression through HIF1beta/VEGFA signalling. Mol Cancer, 2018,17(1):107
Gebert LFR, MacRae IJ. Regulation of microRNA function in animals. Nat Rev Mol Cell Biol, 2019,20(1):21–37
Issler O, Chen A. Determining the role of microRNAs in psychiatric disorders. Nat Rev Neurosci, 2015,16(4):201–212
Kota J, Chivukula RR, O’Donnell KA, et al. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell, 2009,137(6):1005–1017
Janssen HL, Reesink HW, Lawitz EJ, et al. Treatment of HCV infection by targeting microRNA. N Engl J Med, 2013,368(18):1685–1694
Acknowledgmenets
We are grateful to the members of Gui-hua WANG’s lab and Jun-bo HU’s lab for their critical input and suggestions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
All authors have no conflicts of interest to declare.
Additional information
The study was supported by the National Natural Science Foundation of China (No. 81372323 and No. 81802426).
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Qiu, C., Feng, Yd. & Yang, X. MicroRNA-409-5p Inhibits GIST Tumorigenesis and Improves Imatinib Resistance by Targeting KDM4D Expression. CURR MED SCI 43, 935–946 (2023). https://doi.org/10.1007/s11596-023-2715-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11596-023-2715-8