Abstract
Breast cancer pathological image segmentation (BCPIS) holds significant value in assisting physicians with quantifying tumor regions and providing treatment guidance. However, achieving fine-grained semantic segmentation remains a major challenge for this technology. The complex and diverse morphologies of breast cancer tissue structures result in high costs for manual annotation, thereby limiting the sample size and annotation quality of the dataset. These practical issues have a significant impact on the segmentation performance. To overcome these challenges, this study proposes a semi-supervised learning model based on classification-guided segmentation. The model first utilizes a multi-scale convolutional network to extract rich semantic information and then employs a multi-expert cross-layer joint learning strategy, integrating a small number of labeled samples to iteratively provide the model with class-generated multi-cue pseudo-labels and real labels. Given the complexity of the breast cancer samples and the limited sample quantity, an innovative approach of augmenting additional unlabeled data was adopted to overcome this limitation. Experimental results demonstrate that, although the proposed model falls slightly behind supervised segmentation models, it still exhibits significant progress and innovation. The semi-supervised model in this study achieves outstanding performance, with an IoU (Intersection over Union) value of 71.53%. Compared to other semi-supervised methods, the model developed in this study demonstrates a performance advantage of approximately 3%. Furthermore, the research findings indicate a significant correlation between the classification and segmentation tasks in breast cancer pathological images, and the guidance of a multi-expert system can significantly enhance the fine-grained effects of semi-supervised semantic segmentation.
Graphical abstract
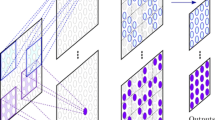
Overall architecture diagram of the model. During the training process, the model is trained by iteratively feeding it labeled and unlabeled data. When presented with unlabeled data, the model leverages the generators 1 and 2 of multiple expert systems to generate images suitable for fine-grained recognition training, along with pseudo mask labels for segmentation training. On the other hand, when presented with labeled images, the model undergoes convergence optimization training.







Similar content being viewed by others
References
Lei S, Zheng R, Zhang S, Wang S, Chen R, Sun K, Zeng H, Zhou J, Wei W (2021) Global patterns of breast cancer incidence and mortality: a population-based cancer registry data analysis from 2000 to 2020. Cancer Commun 41(11):1183–1194
Zuo T-T, Zheng R-S, Zeng H-M, Zhang S-W (2013) Chen W-Q (2017) Female breast cancer incidence and mortality in China. Thoracic Cancer 8(3):214–218
Reshma V, Arya N, Ahmad SS, Wattar I, Mekala S, Joshi S, Krah D et al (2022) Detection of breast cancer using histopathological image classification dataset with deep learning techniques. BioMed Research International 2022
Boyd NF, Martin LJ, Bronskill M, Yaffe MJ, Duric N, Minkin S (2010) Breast tissue composition and susceptibility to breast cancer. J Natl Cancer Inst 102(16):1224–1237
Urbaniak C, Gloor GB, Brackstone M, Scott L, Tangney M, Reid G (2016) The microbiota of breast tissue and its association with breast cancer. Appl Environ Microbiol 8(16):5039–5048
Wang H, Altemus J, Niazi F, Green H, Calhoun BC, Sturgis C, Grobmyer SR, Eng C (2017) Breast tissue, oral and urinary microbiomes in breast cancer. Oncotarget 8(50):88122
Selvaraju RR, Cogswell M, Das A, Vedantam R, Parikh D, Batra D (2017) Grad-CAM: visual explanations from deep networks via gradient-based localization. In: Proceedings of the IEEE international conference on computer vision, pp 618–626
Michael E, Ma H, Li H, Kulwa F, Li J (2021) Breast cancer segmentation methods: current status and future potentials. BioMed Res Int 2021:1–29
Al-Amri SS, Kalyankar NV et al (2010) Image segmentation by using threshold techniques. arXiv:1005.4020
Medina-Carnicer R, Carmona-Poyato A, Muñoz-Salinas R, Madrid-Cuevas FJ (2009) Determining hysteresis thresholds for edge detection by combining the advantages and disadvantages of thresholding methods. IEEE Trans Image Proc 19(1):165–173
Scales N, Kerry C, Prize M (2004) Automated image segmentation for breast analysis using infrared images. In: The 26th annual international conference of the IEEE engineering in medicine and biology society, vol 1, pp 1737–1740 IEEE
Ramani R, Suthanthiravanitha S, Valarmathy S (2012) A survey of current image segmentation techniques for detection of breast cancer. Intl J Eng Res Appl (IJERA) 2(5):1124–1129
Ahmed L, Iqbal MM, Aldabbas H, Khalid S, Saleem Y, Saeed S (2020) Images data practices for semantic segmentation of breast cancer using deep neural network. J Ambient Intell Humanized Comput, 1–17
Ronneberger O, Fischer P, Brox T (2015) U-Net: convolutional networks for biomedical image segmentation. In: Medical image computing and computer-assisted intervention–MICCAI 2015: 18th International Conference, Munich, Germany, Proceedings, Part III 18, pp 234–241, Springer October 5-9, 2015
Oktay O, Schlemper J, Folgoc LL, Lee M, Heinrich M, Misawa K, Mori K, McDonagh S, Hammerla NY, Kainz B et al (2018) Attention U-Net: learning where to look for the pancreas. arXiv:1804.03999
Su H, Liu F, Xie Y, Xing F, Meyyappan S, Yang L (2015) Region segmentation in histopathological breast cancer images using deep convolutional neural network. In: 2015 IEEE 12th international symposium on biomedical imaging (ISBI), pp 55–58 IEEE
Lai Z, Wang C, Oliveira LC, Dugger BN, Cheung S-C, Chuah C-N (2021) Joint semi-supervised and active learning for segmentation of gigapixel pathology images with cost-effective labeling. In: Proceedings of the IEEE/CVF international conference on computer vision, pp 591–600
Hu M, Li Y, Yang X (2023) BreastSAM: a study of segment anything model for breast tumor detection in ultrasound images. arXiv:2305.12447
Zhai D, Hu B, Gong X, Zou H, Luo J (2022) ASS-GAN: asymmetric semi-supervised GAN for breast ultrasound image segmentation. Neurocomput 493:204–216
Amgad M, Elfandy H, Hussein H, Atteya LA, Elsebaie MA, Abo Elnasr LS, Sakr RA, Salem HS, Ismail AF, Saad AM et al (2019) Structured crowdsourcing enables convolutional segmentation of histology images. Bioinforma 35(18):3461–3467
Spanhol FA, Oliveira LS, Petitjean C, Heutte L (2015) A dataset for breast cancer histopathological image classification. IEEE Trans Biomed Eng 63(7):1455–14623
Yu F, Koltun V, Funkhouser T (2017) Dilated residual networks. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 472–480
He K, Zhang X, Ren S, Sun J (2016) Deep residual learning for image recognition. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 770–778
Touvron H, Bojanowski P, Caron M, Cord M, El-Nouby A, Grave E, Izacard G, Joulin A, Synnaeve G, Verbeek J et al (2022) ResMLP: feedforward networks for image classification with data-efficient training. IEEE transactions on pattern analysis and machine intelligence (2022)
Touvron H, Cord M, Sablayrolles A, Synnaeve G, Jégou H (2021) Going deeper with image transformers. In: Proceedings of the IEEE/CVF international conference on computer vision, pp 32–42
Bao H, Dong L, Piao S, Wei F (2021) BEIT: BERT pre-training of image transformers. arXiv:2106.08254
Touvron H, Cord M, Douze M, Massa F, Sablayrolles A, Jégou H (2021) Training data-efficient image transformers & distillation through attention. In: International conference on machine learning, pp 10347–10357 PMLR
Zhou Z, Rahman Siddiquee MM, Tajbakhsh N, Liang J (2018) UNet++: a nested U-Net architecture for medical image segmentation. In: Deep learning in medical image analysis and multimodal learning for clinical decision support: 4th International Workshop, DLMIA 2018, and 8th International Workshop, ML-CDS 2018, Held in Conjunction with MICCAI 2018, Granada, Spain, September 20, 2018, Proceedings 4, pp 3–11 Springer
Fan T, Wang G, Li Y, Wang H (2020) MA-Net: a multi-scale attention network for liver and tumor segmentation. IEEE Access 8:179656–179665
Vu, T-H, Jain H, Bucher M, Cord M, Pérez P (2019) ADVENT: adversarial entropy minimization for domain adaptation in semantic segmentation. In: Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, pp 2517–2526
Tarvainen A, Valpola H (2017) Mean teachers are better role models: weight-averaged consistency targets improve semi-supervised deep learning results. Adv Neural Inf Proc Syst 30
Luo X, Chen J, Song T, Wang G (2021) Semi-supervised medical image segmentation through dual-task consistency. Proceedings of the AAAI conference on artificial intelligence 35:8801–8809
Ouali Y, Hudelot C, Tami M (2020) Semi-supervised semantic segmentation with cross-consistency training. In: Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, pp 12674–12684
Funding
This work is supported by the National Nature Science Foundation of China (no. 62272283), the Major Basic Research Project of Shandong Natural Science Foundation (no. ZR2019ZD04), and the New Twentieth Items of Universities in Jinan (2021GXRC049).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sun, K., Zheng, Y., Yang, X. et al. Semi-supervised breast cancer pathology image segmentation based on fine-grained classification guidance. Med Biol Eng Comput 62, 901–912 (2024). https://doi.org/10.1007/s11517-023-02970-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-023-02970-4