Abstract
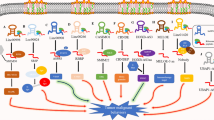
Long noncoding RNAs (lncRNAs) have been extensively identified in eukaryotic genomes and have been shown to play critical roles in the development of multiple cancers. Through the application and development of ribosome analysis and sequencing technologies, advanced studies have discovered the translation of lncRNAs. Although lncRNAs were originally defined as noncoding RNAs, many lncRNAs actually contain small open reading frames that are translated into peptides. This opens a broad area for the functional investigation of lncRNAs. Here, we introduce prospective methods and databases for screening lncRNAs with functional polypeptides. We also summarize the specific lncRNA-encoded proteins and their molecular mechanisms that promote or inhibit cancerous. Importantly, the role of lncRNA-encoded peptides/proteins holds promise in cancer research, but some potential challenges remain unresolved. This review includes reports on lncRNA-encoded peptides or proteins in cancer, aiming to provide theoretical basis and related references to facilitate the discovery of more functional peptides encoded by lncRNA, and to further develop new anti-cancer therapeutic targets as well as clinical biomarkers of diagnosis and prognosis.
Similar content being viewed by others
References
Anastasiadou, E., Jacob, L.S., and Slack, F.J. (2018). Non-coding RNA networks in cancer. Nat Rev Cancer 18, 5–18.
Anderson, D.M., Anderson, K.M., Chang, C.L., Makarewich, C.A., Nelson, B.R., McAnally, J.R., Kasaragod, P., Shelton, J.M., Liou, J., Bassel-Duby, R., et al. (2015). A micropeptide encoded by a putative long noncoding RNA regulates muscle performance. Cell 160, 595–606.
Anderson, K.M., Anderson, D.M., McAnally, J.R., Shelton, J.M., Bassel-Duby, R., and Olson, E.N. (2016). Transcription of the non-coding RNA upperhand controls Hand2 expression and heart development. Nature 539, 433–136.
Baralle, F.E., and Giudice, J. (2017). Alternative splicing as a regulator of development and tissue identity. Nat Rev Mol Cell Biol 18, 437–451.
Bazzini, A.A., Johnstone, T.G., Christiano, R., Mackowiak, S.D., Obermayer, B., Fleming, E.S., Vejnar, C.E., Lee, M.T., Rajewsky, N., Walther, T.C., et al. (2014). Identification of small ORFs in vertebrates using ribosome footprinting and evolutionary conservation. EMBO J 33, 981–993.
Beermann, J., Piccoli, M.T., Viereck, J., and Thum, T. (2016). Non-coding RNAs in development and disease: background, mechanisms, and therapeutic approaches. Physiol Rev 96, 1297–1325.
Brar, G.A., and Weissman, J.S. (2015). Ribosome profiling reveals the what, when, where and how of protein synthesis. Nat Rev Mol Cell Biol 16, 651–664.
Bridges, M.C., Daulagala, A.C., and Kourtidis, A. (2021). LNCcation: lncRNA localization and function. J Cell Biol 220.
Cabrita, L.D., Cassaignau, A.M.E., Launay, H.M.M., Waudby, C.A., Wlodarski, T., Camilloni, C., Karyadi, M.E., Robertson, A.L., Wang, X., Wentink, A.S., et al. (2016). A structural ensemble of a ribosomenascent chain complex during cotranslational protein folding. Nat Struct Mol Biol 23, 278–285.
Cai, T., Zhang, Q., Wu, B., Wang, J., Li, N., Zhang, T., Wang, Z., Luo, J., Guo, X., Ding, X., et al. (2021). LncRNA-encoded microproteins: a new form of cargo in cell culture-derived and circulating extracellular vesicles. J Extracell Vesicles 10, e12123.
Calviello, L., Mukherjee, N., Wyler, E., Zauber, H., Hirsekorn, A., Selbach, M., Landthaler, M., Obermayer, B., and Ohler, U. (2016). Detecting actively translated open reading frames in ribosome profiling data. Nat Methods 13, 165–170.
Cech, T.R., and Steitz, J.A. (2014). The noncoding RNA revolution—trashing old rules to forge new ones. Cell 157, 77–94.
Choi, S.W., Kim, H.W., and Nam, J.W. (2019). The small peptide world in long noncoding RNAs. Brief Bioinform 20, 1853–1864.
Choteau, S.A., Wagner, A., Pierre, P., Spinelli, L., and Brun, C. (2021). MetamORF: a repository of unique short open reading frames identified by both experimental and computational approaches for gene and metagene analyses. Database 2021.
D’Lima, N.G., Ma, J., Winkler, L., Chu, Q., Loh, K.H., Corpuz, E.O., Budnik, B.A., Lykke-Andersen, J., Saghatelian, A., and Slavoff, S.A. (2017). A human microprotein that interacts with the mRNA decapping complex. Nat Chem Biol 13, 174–180.
de Rie, D., Abugessaisa, I., Alam, T., Arner, E., Arner, P., Ashoor, H., Åström, G., Babina, M., Bertin, N., Burroughs, A.M., et al. (2017). An integrated expression atlas of miRNAs and their promoters in human and mouse. Nat Biotechnol 35, 872–878.
Dragomir, M.P., Manyam, G.C., Ott, L.F., Berland, L., Knutsen, E., Ivan, C., Lipovich, L., Broom, B.M., and Calin, G.A. (2020). FuncPEP: a database of functional peptides encoded by non-coding RNAs. ncRNA 6, 41.
Eastman, G., Smircich, P., and Sotelo-Silveira, J.R. (2018). Following ribosome footprints to understand translation at a genome wide level. Comput Struct Biotechnol J 16, 167–176.
Eißmann, M., Gutschner, T., Hämmerle, M., Günther, S., Caudron-Herger, M., Groß, M., Schirmacher, P., Rippe, K., Braun, T., Zörnig, M., et al. (2012). Loss of the abundant nuclear non-coding RNA MALAT1 is compatible with life and development. RNA Biol 9, 1076–1087.
Ge, Q., Jia, D., Cen, D., Qi, Y., Shi, C., Li, J., Sang, L., Yang, L., He, J., Lin, A., et al. (2021). Micropeptide ASAP encoded by LINC00467 promotes colorectal cancer progression by directly modulating ATP synthase activity. J Clin Invest 131.
Ghafouri-Fard, S., Khoshbakht, T., Hussen, B.M., Taheri, M., and Hajiesmaeili, M. (2022). A review on the role of LINC00467 in the carcinogenesis. Cancer Cell Int 22, 319.
Guo, B., Wu, S., Zhu, X., Zhang, L., Deng, J., Li, F., Wang, Y., Zhang, S., Wu, R., Lu, J., et al. (2020). Micropeptide CIP2A-BP encoded by LINC00665 inhibits triple-negative breast cancer progression. EMBO J 39, e102190.
Guo, Y., Chen, J., Zhang, X., Fang, M., Xu, M., Zhang, L., Rao, E., and Xin, Y. (2021). Recombinant human adenovirus-p53 therapy for the treatment of cervical cancer: a meta-analysis. Front Oncol 11, 748681.
Hanyu-Nakamura, K., Sonobe-Nojima, H., Tanigawa, A., Lasko, P., and Nakamura, A. (2008). Drosophila Pgc protein inhibits P-TEFb recruitment to chromatin in primordial germ cells. Nature 451, 730–733.
Heiman, M., Kulicke, R., Fenster, R.J., Greengard, P., and Heintz, N. (2014). Cell type-specific mRNA purification by translating ribosome affinity purification (TRAP). Nat Protoc 9, 1282–1291.
Hinnebusch, A.G. (2014). The scanning mechanism of eukaryotic translation initiation. Annu Rev Biochem 83, 779–812.
Huang, J.Z., Chen, M., Chen, D., Gao, X.C., Zhu, S., Huang, H., Hu, M., Zhu, H., and Yan, G.R. (2017). A peptide encoded by a putative lncRNA HOXB-AS3 suppresses colon cancer growth. Mol Cell 68, 171–184.e6.
Huang, Y., Wang, J., Zhao, Y., Wang, H., Liu, T., Li, Y., Cui, T., Li, W., Feng, Y., Luo, J., et al. (2021). cncRNAdb: a manually curated resource of experimentally supported RNAs with both protein-coding and noncoding function. Nucleic Acids Res 49, D65–D70.
Huarte, M., Guttman, M., Feldser, D., Garber, M., Koziol, M.J., Kenzelmann-Broz, D., Khalil, A.M., Zuk, O., Amit, I., Rabani, M., et al. (2010). A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 142, 409–419.
Ingolia, N.T. (2014). Ribosome profiling: new views of translation, from single codons to genome scale. Nat Rev Genet 15, 205–213.
Ingolia, N.T., Brar, G.A., Stern-Ginossar, N., Harris, M.S., Talhouarne, G.J.S., Jackson, S.E., Wills, M.R., and Weissman, J.S. (2014). Ribosome profiling reveals pervasive translation outside of annotated protein-coding genes. Cell Rep 8, 1365–1379.
Ingolia, N.T., Ghaemmaghami, S., Newman, J.R.S., and Weissman, J.S. (2009). Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223.
Ji, Z. (2018). Rfoot: transcriptome-scale identification of RNA-protein complexes from ribosome profiling data. Curr Protoc Mol Biol 124, e66.
Kahles, A., Lehmann, K.V., Toussaint, N.C., Hüser, M., Stark, S.G., Sachsenberg, T., Stegle, O., Kohlbacher, O., Sander, C., Rätsch, G., et al. (2018). Comprehensive analysis of alternative splicing across tumors from 8,705 patients. Cancer Cell 34, 211–224.e6.
Kang, Y.J., Yang, D.C., Kong, L., Hou, M., Meng, Y.Q., Wei, L., and Gao, G. (2017). CPC2: a fast and accurate coding potential calculator based on sequence intrinsic features. Nucleic Acids Res 45, W12–W16.
Khan, M.R., Wellinger, R.J., and Laurent, B. (2021). Exploring the alternative splicing of long noncoding RNAs. Trends Genet 37, 695–698.
Kopp, F., and Mendell, J.T. (2018). Functional classification and experimental dissection of long noncoding RNAs. Cell 172, 393–407.
Lauria, F., Tebaldi, T., Bernabo, P., Groen, E.J.N., Gillingwater, T.H., and Viero, G. (2018). riboWaltz: Optimization of ribosome P-site positioning in ribosome profiling data. PLoS Comput Biol 14, e1006169, doi: https://doi.org/10.1371/journal.pcbi.1006169.
Lee, S., Kopp, F., Chang, T.C., Sataluri, A., Chen, B., Sivakumar, S., Yu, H., Xie, Y., and Mendell, J.T. (2016). Noncoding RNA NORAD regulates genomic stability by sequestering PUMILIO proteins. Cell 164, 69–80.
Li, A., Zhang, J., and Zhou, Z. (2014). PLEK: a tool for predicting long non-coding RNAs and messenger RNAs based on an improved k-mer scheme. BMC Bioinformatics 15, 311.
Li, J., Huang, C., Zou, Y., Ye, J., Yu, J., and Gui, Y. (2020a). CircTLK1 promotes the proliferation and metastasis of renal cell carcinoma by sponging miR-136-5p. Mol Cancer 19, 103.
Li, Q., Chen, Q., Klauser, P.C., Li, M., Zheng, F., Wang, N., Li, X., Zhang, Q., Fu, X., Wang, Q., et al. (2020b). Developing covalent protein drugs via proximity-enabled reactive therapeutics. Cell 182, 85–97.e16.
Li, W., Wang, W., Uren, P.J., Penalva, L.O.F., and Smith, A.D. (2017). Riborex: fast and flexible identification of differential translation from Ribo-seq data. Bioinformatics 33, 1735–1737.
Li, Y., Zhou, H., Chen, X., Zheng, Y., Kang, Q., Hao, D., Zhang, L., Song, T., Luo, H., Hao, Y., et al. (2021). SmProt: a reliable repository with comprehensive annotation of small proteins identified from ribosome profiling. Genomics Proteomics Bioinf 19, 602–610.
Liang, Y., Song, X., Li, Y., Chen, B., Zhao, W., Wang, L., Zhang, H., Liu, Y., Han, D., Zhang, N., et al. (2020). RETRACTED ARTICLE: LncRNA BCRT1 promotes breast cancer progression by targeting miR-1303/PTBP3 axis. Mol Cancer 19, 85.
Lin, M.F., Jungreis, I., and Kellis, M. (2011). PhyloCSF: a comparative genomics method to distinguish protein coding and non-coding regions. Bioinformatics 27, i275–i282.
Lin, X., Wu, Z., Hu, H., Luo, M.L., and Song, E. (2021). Non-coding RNAs rewire cancer metabolism networks. Semin Cancer Biol 75, 116–126.
Liu, H., Zhou, X., Yuan, M., Zhou, S., Huang, Y., Hou, F., Song, X., Wang, L., and Jiang, W. (2020). ncEP: a manually curated database for experimentally validated ncRNA-encoded proteins or peptides. J Mol Biol 432, 3364–3368.
Liu, Y., Cheng, Z., Pang, Y., Cui, L., Qian, T., Quan, L., Zhao, H., Shi, J., Ke, X., and Fu, L. (2019). Role of microRNAs, circRNAs and long noncoding RNAs in acute myeloid leukemia. J Hematol Oncol 12, 51.
Liu, Y., Liu, X., Lin, C., Jia, X., Zhu, H., Song, J., and Zhang, Y. (2021). Noncoding RNAs regulate alternative splicing in cancer. J Exp Clin Cancer Res 40, 11.
Lu, S., Zhang, J., Lian, X., Sun, L., Meng, K., Chen, Y., Sun, Z., Yin, X., Li, Y., Zhao, J., et al. (2019). A hidden human proteome encoded by ‘non-coding’ genes. Nucleic Acids Res 47, 8111–8125.
Lun, Y.Z., Pan, Z.P., Liu, S.A., Sun, J., Han, M., Liu, B., Dong, W., Pan, L. H., and Cheng, J. (2020). The peptide encoded by a novel putative lncRNA HBVPTPAP inducing the apoptosis of hepatocellular carcinoma cells by modulating JAK/STAT signaling pathways. Virus Res 287, 198104.
Luo, X., Huang, Y., Li, H., Luo, Y., Zuo, Z., Ren, J., and Xie, Y. (2022). SPENCER: a comprehensive database for small peptides encoded by noncoding RNAs in cancer patients. Nucleic Acids Res 50, D1373–D1381.
Maacha, S., Bhat, A.A., Jimenez, L., Raza, A., Haris, M., Uddin, S., and Grivel, J.C. (2019). Extracellular vesicles-mediated intercellular communication: roles in the tumor microenvironment and anti-cancer drug resistance. Mol Cancer 18, 55.
Mackowiak, S.D., Zauber, H., Bielow, C., Thiel, D., Kutz, K., Calviello, L., Mastrobuoni, G., Rajewsky, N., Kempa, S., Selbach, M., et al. (2015). Extensive identification and analysis of conserved small ORFs in animals. Genome Biol 16, 179.
Magny, E.G., Pueyo, J.I., Pearl, F.M.G., Cespedes, M.A., Niven, J.E., Bishop, S.A., and Couso, J.P. (2013). Conserved regulation of cardiac calcium uptake by peptides encoded in small open reading frames. Science 341, 1116–1120.
Makarewich, C.A., Baskin, K.K., Munir, A.Z., Bezprozvannaya, S., Sharma, G., Khemtong, C., Shah, A.M., McAnally, J.R., Malloy, C. R., Szweda, L.I., et al. (2018). MOXI is a mitochondrial micropeptide that enhances fatty acid β-oxidation. Cell Rep 23, 3701–3709.
Malone, B., Atanassov, I., Aeschimann, F., Li, X., Großhans, H., and Dieterich, C. (2017). Bayesian prediction of RNA translation from ribosome profiling. Nucleic Acids Res gkw1350.
Mašek, T., Valášek, L., and Pospíšek, M. (2011). Polysome analysis and RNA purification from sucrose gradients. In: Nielsen, H., ed. RNA. Methods in Molecular Biology. New York: Humana Press. 293–309.
Matsumoto, A., Pasut, A., Matsumoto, M., Yamashita, R., Fung, J., Monteleone, E., Saghatelian, A., Nakayama, K.I., Clohessy, J.G., and Pandolfi, P.P. (2017). mTORC1 and muscle regeneration are regulated by the LINC00961-encoded SPAR polypeptide. Nature 541, 228–232.
Meng, N., Chen, M., Chen, D., Chen, X., Wang, J., Zhu, S., He, Y., Zhang, X., Lu, R., and Yan, G. (2020). Small protein hidden in lncRNA LOC90024 promotes “cancerous” RNA splicing and tumorigenesis. Adv Sci 7, 1903233.
Nelson, B.R., Makarewich, C.A., Anderson, D.M., Winders, B.R., Troupes, C.D., Wu, F., Reese, A.L., McAnally, J.R., Chen, X., Kavalali, E.T., et al. (2016). A peptide encoded by a transcript annotated as long noncoding RNA enhances SERCA activity in muscle. Science 351, 271–275.
Olexiouk, V., Crappé, J., Verbruggen, S., Verhegen, K., Martens, L., and Menschaert, G. (2016). sORFs.org: a repository of small ORFs identified by ribosome profiling. Nucleic Acids Res 44, D324–D329.
Pang, Y., Liu, Z., Han, H., Wang, B., Li, W., Mao, C., and Liu, S. (2020). Peptide SMIM30 promotes HCC development by inducing SRC/YES1 membrane anchoring and MAPK pathway activation. J Hepatol 73, 1155–1169.
Pauli, A., Norris, M.L., Valen, E., Chew, G.L., Gagnon, J.A., Zimmerman, S., Mitchell, A., Ma, J., Dubrulle, J., Reyon, D., et al. (2014). Toddler: an embryonic signal that promotes cell movement via Apelin receptors. Science 343, 1248636.
Qi, X., Chen, S., He, H., Wen, W., and Wang, H. (2021). The role and potential application of extracellular vesicles in liver cancer. Sci China Life Sci 64, 1281–1294.
Raj, A., Wang, S.H., Shim, H., Harpak, A., Li, Y.I., Engelmann, B., Stephens, M., Gilad, Y., and Pritchard, J.K. (2016). Thousands of novel translated open reading frames in humans inferred by ribosome footprint profiling. eLife 5, e13328.
Röhrig, H., Schmidt, J., Miklashevichs, E., Schell, J., and John, M. (2002). Soybean ENOD40 encodes two peptides that bind to sucrose synthase. Proc Natl Acad Sci USA 99, 1915–1920.
Rombel, I.T., Sykes, K.F., Rayner, S., and Johnston, S.A. (2002). ORF-FINDER: a vector for high-throughput gene identification. Gene 282, 33–41.
Rossi, M., Bucci, G., Rizzotto, D., Bordo, D., Marzi, M.J., Puppo, M., Flinois, A., Spadaro, D., Citi, S., Emionite, L., et al. (2019). LncRNA EPR controls epithelial proliferation by coordinating Cdkn1a transcription and mRNA decay response to TGF-β. Nat Commun 10, 1969.
Ruiz-Orera, J., and Albà, M.M. (2019). Translation of small open reading frames: roles in regulation and evolutionary innovation. Trends Genet 35, 186–198.
Saw, P.E., and Song, E.W. (2020). siRNA therapeutics: a clinical reality. Sci China Life Sci 63, 485–500.
Saw, P.E., Xu, X., Chen, J., and Song, E.W. (2021). Non-coding RNAs: the new central dogma of cancer biology. Sci China Life Sci 64, 22–50.
Shi, S.W., Li, B., Dong, Y., Ge, Y., Qu, X., Lu, L.G., Yuan, Y.H., Li, L.J., and Li, Y. (2019). In vitro and clinical studies of gene therapy with recombinant human adenovirus-p5_? injection for malignant melanoma. Hum Gene Ther Clin Dev 30, 7–18.
Spencer, H.L., Sanders, R., Boulberdaa, M., Meloni, M., Cochrane, A., Spiroski, A.M., Mountford, J., Emanueli, C., Caporali, A., Brittan, M., et al. (2020). The LINC00961 transcript and its encoded micropeptide, small regulatory polypeptide of amino acid response, regulate endothelial cell function. Cardiovasc Res 116, 1981–1994.
St. Laurent, G., Wahlestedt, C., and Kapranov, P. (2015). The landscape of long noncoding RNA classification. Trends Genet 31, 239–251.
Stein, C.S., Jadiya, P., Zhang, X., McLendon, J.M., Abouassaly, G.M., Witmer, N.H., Anderson, E.J., Elrod, J.W., and Boudreau, R.L. (2018). Mitoregulin: a lncRNA-encoded microprotein that supports mitochondrial supercomplexes and respiratory efficiency. Cell Rep 23, 3710–3720.e8.
Stern-Ginossar, N., and Ingolia, N.T. (2015). Ribosome profiling as a tool to decipher viral complexity. Annu Rev Virol 2, 335–349.
Su, P.P., Liu, D.W., Zhou, S.J., Chen, H., Wu, X.M., and Liu, Z.S. (2022). Down-regulation of Risa improves podocyte injury by enhancing autophagy in diabetic nephropathy. Mil Med Res 9, 23.
Tang, L., Zheng, Y., Melo, M.B., Mabardi, L., Castaño, A.P., Xie, Y.Q., Li, N., Kudchodkar, S.B., Wong, H.C., Jeng, E.K., et al. (2018). Enhancing T cell therapy through TCR-signaling-responsive nanoparticle drug delivery. Nat Biotechnol 36, 707–716.
Wang, L., Fan, J., Han, L., Qi, H., Wang, Y., Wang, H., Chen, S., Du, L., Li, S., Zhang, Y., et al. (2020a). The micropeptide LEMP plays an evolutionarily conserved role in myogenesis. Cell Death Dis 11, 357.
Wang, L., Park, H.J., Dasari, S., Wang, S., Kocher, J.P., and Li, W. (2013). CPAT: Coding-Potential Assessment Tool using an alignment-free logistic regression model. Nucleic Acids Res 41, e74.
Wang, Y., Wu, S., Zhu, X., Zhang, L., Deng, J., Li, F., Guo, B., Zhang, S., Wu, R., Zhang, Z., et al. (2020b). LncRNA-encoded polypeptide ASRPS inhibits triple-negative breast cancer angiogenesis. J Exp Med 217.
Wu, A.H., Chen, X.L., Guo, L.Y., Lu, D.F., Lu, S., Wang, A.A., and Liang, X.F. (2021a). Downregulation of lncRNA IGF2-AS-encoded peptide induces trophoblast—cycle arrest. Reprod Biomed Online 43, 598–606.
Wu, M., Xu, G., Han, C., Luan, P.F., Xing, Y.H., Nan, F., Yang, L.Z., Huang, Y., Yang, Z.H., Shan, L., et al. (2021b). lncRNA SLERT controls phase separation of FC/DFCs to facilitate Pol I transcription. Science 373, 547–555.
Wu, P., Mo, Y., Peng, M., Tang, T., Zhong, Y., Deng, X., Xiong, F., Guo, C., Wu, X., Li, Y., et al. (2020a). Emerging role of tumor-related functional peptides encoded by lncRNA and circRNA. Mol Cancer 19, 22.
Wu, S., Zhang, L., Deng, J., Guo, B., Li, F., Wang, Y., Wu, R., Zhang, S., Lu, J., and Zhou, Y. (2020b). A novel micropeptide encoded by Y-linked LINC00278 links cigarette smoking and AR signaling in male esophageal squamous cell carcinoma. Cancer Res 80, 2790–2803.
Xiang, X., Fu, Y., Zhao, K., Miao, R., Zhang, X., Ma, X., Liu, C., Zhang, N., and Qu, K. (2021). Cellular senescence in hepatocellular carcinoma induced by a long non-coding RNA-encoded peptide PINT87aa by blocking FOXM1-mediated PHB2. Theranostics 11, 4929–4944.
Xiao, Z., Huang, R., Xing, X., Chen, Y., Deng, H., and Yang, X. (2018). De novo annotation and characterization of the translatome with ribosome profiling data. Nucleic Acids Res 46, e61.
Xiao, Z., Zou, Q., Liu, Y., and Yang, X. (2016). Genome-wide assessment of differential translations with ribosome profiling data. Nat Commun 7, 11194.
Xin, Y., Huang, Q., Tang, J.Q., Hou, X.Y., Zhang, P., Zhang, L.Z., and Jiang, G. (2016). Nanoscale drug delivery for targeted chemotherapy. Cancer Lett 379, 24–31.
Xu, Z., Hu, L., Shi, B., Geng, S.S., Xu, L., Wang, D., and Lu, Z.J. (2018). Ribosome elongating footprints denoised by wavelet transform comprehensively characterize dynamic cellular translation events. Nucleic Acids Res 46, e109.
Xue, Y., Chen, R., Qu, L., and Cao, X. (2020). Noncoding RNA: from dark matter to bright star. Sci China Life Sci 63, 463–468.
Yu, B., Qi, Y., Li, R., Shi, Q., Satpathy, A.T., and Chang, H.Y. (2021). B cell-specific XIST complex enforces X-inactivation and restrains atypical B cells. Cell 184, 1790–1803.e17.
Zhang, Q., Vashisht, A.A., O’Rourke, J., Corbel, S.Y., Moran, R., Romero, A., Miraglia, L., Zhang, J., Durrant, E., Schmedt, C., et al. (2017). The microprotein Minion controls cell fusion and muscle formation. Nat Commun 8, 15664.
Zhong, Y., Karaletsos, T., Drewe, P., Sreedharan, V.T., Kuo, D., Singh, K., Wendel, H.-G., and Rätsch, G. (2017). RiboDiff: detecting changes of mRNA translation efficiency from ribosome footprints. Bioinformatics 33, 139–141.
Zhou, D.D., Bai, W.Q., Zhai, X.T., Sun, L.P., Zhen, Y.S., Li, Z.R., and Miao, Q.F. (2021). Excellent effects and possible mechanisms of action of a new antibody-drug conjugate against EGFR-positive triple-negative breast cancer. Mil Med Res 8, 63.
Zhou, J., Liu, Z., and Li, F. (2012). Upconversion nanophosphors for small-animal imaging. Chem Soc Rev 41, 1323–1349.
Zhu, S., Wang, J.Z., Chen, D., He, Y.T., Meng, N., Chen, M., Lu, R.X., Chen, X.H., Zhang, X.L., and Yan, G.R. (2020). An oncopeptide regulates m6A recognition by the m6A reader IGF2BP1 and tumorigenesis. Nat Commun 11, 1685.
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2021YFF1201300, 2022YFE0103600), the National Natural Science Foundation of China (82073094), the CAMS Innovation Fund for Medical Sciences (CIFMS) (2021-I2M-014), the Open Issue of State Key Laboratory of Molecular Oncology (SKL-KF-2021-16), and the Independent Issue of State Key Laboratory of Molecular Oncology (SKL-2021-16).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Compliance and ethics The author(s) declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Wang, J., Wang, W., Ma, F. et al. A hidden translatome in tumors—the coding lncRNAs. Sci. China Life Sci. 66, 2755–2772 (2023). https://doi.org/10.1007/s11427-022-2289-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-022-2289-6