Abstract
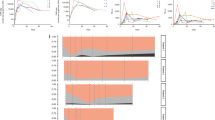
Diversity in the T cell receptor (TCR) repertoire provides a miniature defense ability for the T cell immune system that may be related to tumor initiation and progression. Understanding the T cell immune status of leukemia patients is critical for establishing specific immunotherapies. Previous studies have reported abnormal TCR repertoires and clonally expanded TCR Vβ T cells in chronic myeloid leukemia in chronic phase (CP-CML). In this study, we investigated the distribution and clonality of the TCR Vβ repertoire in 4 cases with imatinib-resistant CML in blast crisis (BC-CML) with abelson murine leukemia viral oncogene homolog 1 (ABL1) kinase domain mutations (KDMs). Examination of TCR V expression and clonality was performed by reverse transcription-polymerase chain reaction (RT-PCR) and GeneScan analysis. Significantly skewed TCR Vβ repertoires were observed in BC-CML patients with different KDMs, and 4 to 8 oligoclonally expanded TCR Vβ subfamilies could be identified in each sample. Intriguingly, a relatively highly expanded Vβ9 clone with the same length as complementarity- determining region 3 (CDR3) (139 bp) was found in all three CML patients in lymphoid blast crisis (LBC-CML) who had different KDMs, but the clone was not detected in the only CML patient in myeloid blast crisis (MBC-CML). In conclusion, restricted TCR Vβ repertoire expression and decreased clone complexity was a general phenomenon observed in the BC-CML patients with different KDMs, indicating the T-cell immunodeficiency of these patients. In addition, clonally expanded Vβ9 T cell clones may indicate a specific immune response to leukemia-associated antigens in LBC-CML patients.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Apperley JF. Chronic myeloid leukaemia. The Lancet, 2015, 385: 1447–1459
Li Y, Yang L, Chen S, Li R, Zhang Y, Lu Y, Luo G. Clonal expansion T cells identified in acute monoblastic leukemia by CDR3 size analysis of TCR T beta repertoire using RT-PCR and genescan. Chin Med J (Engl), 2002, 115: 69–71
Kantarjian H, Sawyers C, Hochhaus A, Guilhot F, Schiffer C, Gambacorti-Passerini C, Niederwieser D, Resta D, Capdeville R, Zoellner U, Talpaz M, Druker B, Goldman J, O’Brien SG, Russell N, Fischer T, Ottmann O, Cony-Makhoul P, Facon T, Stone R, Miller C, Tallman M, Brown R, Schuster M, Loughran T, Gratwohl A, Mandelli F, Saglio G, Lazzarino M, Russo D, Baccarani M, Morra E. Hematologic and cytogenetic responses to imatinib mesylate in chronic myelogenous leukemia. N Engl J Med, 2002, 346: 645–652
Usui N. Molecular targeted treatment—new treatment strategy for patients with chronic myeloid leukemia. Rinsho Byori, 2004, 52: 136–144
Kok KH, Tang HM, Jin DY. ‘One Health’ for the people of Hong Kong and the world. Sci China Life Sci, 2012, doi: 10.1007/s11427-012-4408-6
Jabbour E, Kantarjian H, Jones D, Talpaz M, Bekele N, O’Brien S, Zhou X, Luthra R, Garcia-Manero G, Giles F, Rios MB, Terstovsek S, Cortes J. Frequency and clinical significance of BCR-ABL mutations in patients with chronic myeloid leukemia treated with imatinib mesylate. Leukemia, 2006, 20: 1767–1773
Shah NP, Nicoll JM, Nagar B, Gorre ME, Paquette RL, Kuriyan J, Sawyers CL. Multiple BCR-ABL kinase domain mutations confer polyclonal resistance to the tyrosine kinase inhibitor imatinib (STI571) in chronic phase and blast crisis chronic myeloid leukemia. Cancer Cell, 2002, 2: 117–125
Ilaria RL Jr. Pathobiology of lymphoid and myeloid blast crisis and management issues. Hematology Am Soc Hematol Educ Program, 2005, 188–194
Saussele S, Silver RT. Management of chronic myeloid leukemia in blast crisis. Ann Hematol, 2015, 94 Suppl 2: S159–S165
Chereda B, Melo JT. Natural course and biology of CML. Ann Hematol, 2015, 94 Suppl 2: S107–S121
Apperley JF. Part I: Mechanisms of resistance to imatinib in chronic myeloid leukaemia. Lancet Oncol, 2007, 8: 1018–1029
Arstila TP, Casrouge A, Baron T, Even J, Kanellopoulos J, Kourilsky P. A direct estimate of the human alphabeta T cell receptor diversity. Science, 1999, 286: 958–961
Qi Q, Liu Y, Cheng Y, Glanville J, Zhang D, Lee JY, Olshen RA, Weyand CM, Boyd SD, Goronzy JJ. Diversity and clonal selection in the human T-cell repertoire. Proc Natl Acad Sci USA, 2014, 111: 13139–13144
Nikolich-Zugich J, Slifka MK, Messaoudi I. The many important facets of T-cell repertoire diversity. Nat Rev Immunol, 2004, 4: 123–132
Li Y, Chen S, Yang L, Yin Q, Geng S, Wu X, Schmidt CA, Przybylski GK. TRAT and TRBT repertoire, clonality and the proliferative history of umbilical cord blood T-cells. Transpl Immunol, 2007, 18: 151–158
Li Y, Yang L, Chen S, Zhang Y, Zhang X, Luo G. T cell receptor Tbeta repertoire usage and clonal expansion of T cells in chronic myelogenous leukemia. Chin Med J (Engl), 2004, 117: 840–843
Zha X, Chen S, Yang L, Li B, Chen Y, Yan X, Li Y. Characterization of the CDR3 structure of the Tbeta21 T cell clone in patients with P210(BCR-ABL)-positive chronic myeloid leukemia and B-cell acute lymphoblastic leukemia. Hum Immunol, 2011, 72: 798–804
Jin Z, Wu X, Chen S, Yang L, Liu Q, Li Y. Distribution and clonality of the vα and vβ T-cell receptor repertoire of regulatory T cells in leukemia patients with and without graft versus host disease. DNA Cell Biol, 2014, 33: 182–188
Li Y, Geng S, Du X, Chen S, Yang L, Wu X, Li B, Schmidt CA, Przybylski GK. Restricted TRBT repertoire in CD4+ and CD8+ T-cell subsets from CML patients. Hematology, 2011, 16: 43–49
Li Y. Leukemia associated clonal expansion of TCR Tbeta subfamily T cells. Hematology, 2003, 8: 375–384
Li Y, Yang L, Chen S, Zhang Y, Wu X. The TCR Tbeta repertoire usage of T-cells from cord blood induced by chronic myelogenous leukemia associated antigen. Hematology, 2005, 10: 387–392
thor Straten P, Guldberg P, Gronbaek K, Hansen MR, Kirkin AF, Seremet T, Zeuthen J, Becker JC. In situ T cell responses against melanoma comprise high numbers of locally expanded T cell clonotypes. J Immunol, 1999, 163: 443–447
Claret EJ, Alyea EP, Orsini E, Pickett CC, Collins H, Wang Y, Neuberg D, Soiffer RJ, Ritz J. Characterization of T cell repertoire in patients with graft-versus-leukemia after donor lymphocyte infusion. J Clin Invest, 1997, 100: 855–866
Marktel S, Marin D, Foot N, Szydlo R, Bua M, Karadimitris A, De Melo TA, Kotzampaltiris P, Dazzi F, Rahemtulla A, Olavarria E, Apperley JF, Goldman JM. Chronic myeloid leukemia in chronic phase responding to imatinib: the occurrence of additional cytogenetic abnormalities predicts disease progression. Haematologica, 2003, 88: 260–267
Wang L, Zhu K, Zha X, Chen S, Yang L, Chen S, Li Y. Evolution of T-cell clonality in a patient with Ph-negative acute lymphocytic leukemia occurring after interferon and imatinib therapy for Ph-positive chronic myeloid leukemia. J Hematol Oncol, 2010, 3: 14
Wang Q, Cheng T. Evidences for mutations in the histone modifying gene SETD2 as critical drivers in leukemia development. Sci China Life Sci, 2014, 57: 944–946
Author information
Authors and Affiliations
Corresponding author
Additional information
Contributed equally to this work
This article is published with open access at link.springer.com
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0), which permits use, duplication, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Xu, L., Lu, Y., Lai, J. et al. Characteristics of the TCR Vβ repertoire in imatinib-resistant chronic myeloid leukemia patients with ABL mutations. Sci. China Life Sci. 58, 1276–1281 (2015). https://doi.org/10.1007/s11427-015-4930-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-015-4930-4