Abstract
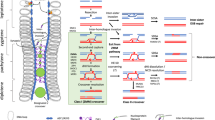
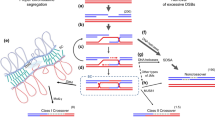
Meiotic recombination is a deeply conserved process within eukaryotes that has a profound effect on patterns of natural genetic variation. During meiosis homologous chromosomes pair and undergo DNA double strand breaks generated by the Spo11 endonuclease. These breaks can be repaired as crossovers that result in reciprocal exchange between chromosomes. The frequency of recombination along chromosomes is highly variable, for example, crossovers are rarely observed in heterochromatin and the centromeric regions. Recent work in plants has shown that crossover hotspots occur in gene promoters and are associated with specific chromatin modifications, including H2A.Z. Meiotic chromosomes are also organized in loop-base arrays connected to an underlying chromosome axis, which likely interacts with chromatin to organize patterns of recombination. Therefore, epigenetic information exerts a major influence on patterns of meiotic recombination along chromosomes, genetic variation within populations and evolution of plant genomes.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Villeneuve AM, Hillers KJ. Whence meiosis? Cell, 2001, 106: 647–650
Wilkins AS, Holliday R. The evolution of meiosis from mitosis. Genetics, 2009, 181: 3–12
Cavalier-Smith T. Origins of the machinery of recombination and sex. Heredity (Edinb), 2002, 88: 125–141
Barton NH, Charlesworth B. Why sex and recombination? Science, 1998, 281: 1986–1990
Hamilton WD. Narrow Roads of Gene Land: the Collected Papers of W.D Hamilton. Volume 2 The Evolution of Sex. Oxford: Oxford University Press, 2001
Higgins JD, Osman K, Jones GH, Franklin FCH. Factors underlying restricted crossover localization in barley meiosis. Annu Rev Genet, 2014, 48: 29–47
Keeney S, Giroux CN, Kleckner N. Meiosis-specific DNA double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell, 1997, 88: 375–384
Bergerat A, de Massy B, Gadelle D, Varoutas PC, Nicolas A, Forterre P. An atypical topoisomerase II from Archaea with implications for meiotic recombination. Nature, 1997, 386: 414–417
Pan J, Sasaki M, Kniewel R, Murakami H, Blitzblau HG, Tischfield SE, Zhu X, Neale MJ, Jasin M, Socci ND, Hochwagen A, Keeney S. A hierarchical combination of factors shapes the genome-wide topography of yeast meiotic recombination initiation. Cell, 2011, 144: 719–731
Neale MJ, Pan J, Keeney S. Endonucleolytic processing of covalent protein-linked DNA double-strand breaks. Nature, 2005, 436: 1053–1057
Garcia V, Phelps SE, Gray S, Neale MJ. Bidirectional resection of DNA double-strand breaks by Mre11 and Exo1. Nature, 2011, 479: 241–244
Bishop DK, Park D, Xu L, Kleckner N. DMC1: a meiosis-specific yeast homolog of E. coli recA required for recombination, synaptonemal complex formation, and cell cycle progression. Cell, 1992, 69: 439–456
Shinohara A, Ogawa H, Ogawa T. Rad51 protein involved in repair and recombination in S. cerevisiae is a RecA-like protein. Cell, 1992, 69: 457–470
Grelon M, Vezon D, Gendrot G, Pelletier G. AtSPO11-1 is necessary for efficient meiotic recombination in plants. EMBO J, 2001, 20: 589–600
Couteau F, Belzile F, Horlow C, Grandjean O, Vezon D, Doutriaux MP. Random chromosome segregation without meiotic arrest in both male and female meiocytes of a dmc1 mutant of Arabidopsis. Plant Cell, 1999, 11: 1623–1634
Youds JL, Boulton SJ. The choice in meiosis-defining the factors that influence crossover or non-crossover formation. J Cell Sci, 2011, 124: 501–513
McMahill MS, Sham CW, Bishop DK. Synthesis-dependent strand annealing in meiosis. PLoS Biol, 2007, 5: e299
Keeney S, Neale MJ. Initiation of meiotic recombination by formation of DNA double-strand breaks: mechanism and regulation. Biochem Soc Trans, 2006, 34: 523–525
Chelysheva L, Vezon D, Chambon A, Gendrot G, Pereira L, Lemhemdi A, Vrielynck N, Le Guin S, Novatchkova M, Grelon M. The Arabidopsis HEI10 is a new ZMM protein related to Zip3. PLoS Genet, 2012, 8: e1002799
Ferdous M, Higgins JD, Osman K, Lambing C, Roitinger E, Mechtler K, Armstrong SJ, Perry P, Pradillo M, Cunado N, Franklin FC. Inter-homolog crossing-over and synapsis in Arabidopsis meiosis are dependent on the chromosome axis protein AtASY3. PLoS Genet, 2012, 8: e1002507
Giraut L, Falque M, Drouaud J, Pereira L, Martin OC, Mézard C. Genome-wide crossover distribution in Arabidopsis thaliana meiosis reveals sex-specific patterns along chromosomes. PLoS Genet, 2011, 7: e1002354
Salomé PA, Bomblies K, Fitz J, Laitinen RA, Warthmann N, Yant L, Weigel D. The recombination landscape in Arabidopsis thaliana F2 populations. Heredity, 2012, 108: 447–455
Lynn A, Soucek R, Börner GV. ZMM proteins during meiosis: crossover artists at work. Chromosome Res, 2007, 15: 591–605
Muller HJ. The mechanism of crossing-over. Am Nat, 1916, 50: 193–221
Sturtevant AH. The behavior of the chromosomes as studied through linkage. Zeitschrift fur Indukt. Abstammungs und Vererbungslehre, 1915, 13: 234–287
Copenhaver GP, Housworth EA, Stahl FW. Crossover interference in Arabidopsis. Genetics, 2002, 160: 1631–1639
Chelysheva L, Grandont L, Vrielynck N, Le Guin S, Mercier R, Grelon M. An easy protocol for studying chromatin and recombination protein dynamics during Arabidopsis thaliana meiosis: immunodetection of cohesins, histones and MLH1. Cytogenet Genome Res, 2010, 129: 143–153
Chelysheva L, Gendrot G, Vezon D, Doutriaux MP, Mercier R, Grelon M. Zip4/Spo22 is required for class I CO formation but not for synapsis completion in Arabidopsis thaliana. PLoS Genet, 2007, 3: e83
Chen C, Zhang W, Timofejeva L, Gerardin Y, Ma H. The Arabidopsis ROCK-N-ROLLERS gene encodes a homolog of the yeast ATP-dependent DNA helicase MER3 and is required for normal meiotic crossover formation. Plant J, 2005, 43: 321–334
Higgins JD, Armstrong SJ, Franklin FCH, Jones GH. The Arabidopsis MutS homolog AtMSH4 functions at an early step in recombination: evidence for two classes of recombination in Arabidopsis. Genes Dev, 2004, 18: 2557–2570
Higgins JD, Vignard J, Mercier R, Pugh AG, Franklin FC, Jones GH. AtMSH5 partners AtMSH4 in the class I meiotic crossover pathway in Arabidopsis thaliana, but is not required for synapsis. Plant J, 2008, 55: 28–39
Macaisne N, Novatchkova M, Peirera L, Vezon D, Jolivet S, Froger N, Chelysheva L, Grelon M, Mercier R. SHOC1, an XPF endonuclease-related protein, is essential for the formation of class I meiotic crossovers. Curr Biol, 2008, 18: 1432–1437
Mercier R, Jolivet S, Vezon D, Huppe E, Chelysheva L, Giovanni M, Noqué F, Doutriaux MP, Horlow C, Grelon M, Mézard C. Two meiotic crossover classes cohabit in Arabidopsis: one is dependent on MER3,whereas the other one is not. Curr Biol, 2005, 15: 692–701
Wijeratne AJ, Chen C, Zhang W, Timofejeva L, Ma H. The Arabidopsis thaliana PARTING DANCERS gene encoding a novel protein is required for normal meiotic homologous recombination. Mol Biol Cell, 2006, 17: 1331–1343
Higgins JD, Sanchez-Moran E, Armstrong SJ, Jones GH, Franklin FCH. The Arabidopsis synaptonemal complex protein ZYP1 is required for chromosome synapsis and normal fidelity of crossing over. Genes Dev, 2005, 19: 2488–2500
Higgins JD, Buckling EF, Franklin FCH, Jones GH. Expression and functional analysis of AtMUS81 in Arabidopsis meiosis reveals a role in the second pathway of crossing-over. Plant J, 2008, 54: 152–162
Berchowitz LE, Francis KE, Bey AL, Copenhaver GP. The role of AtMUS81 in interference-insensitive crossovers in A. thaliana. PLoS Genet, 2007, 3: e132
Crismani W, Girard C, Froger N, Pradillo M, Santos JL, Chelysheva L, Copenhaver GP, Horlow C, Mercier R. FANCM limits meiotic crossovers. Science, 2012, 336: 1588–1590
Knoll A, Higgins JD, Seeliger K, Reha SJ, Dangel NJ, Bauknecht M, Schröpfer S, Franklin FC, Puchta H. The Fanconi anemia ortholog FANCM ensures ordered homologous recombination in both somatic and meiotic cells in Arabidopsis. Plant Cell, 2012, 24: 1448–1464
Kauppi L, Jeffreys AJ, Keeney S. Where the crossovers are: recombination distributions in mammals. Nat Rev Genet, 2004, 5: 413–424
Law JA, Jacobsen SE. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat Rev Genet, 2010, 11: 204–220
Henderson IR, Jacobsen SE. Epigenetic inheritance in plants. Nature, 2007, 447: 418–424
Yelina NE, Choi K, Chelysheva L, Macaulay M, de Snoo B, Wijnker E, Miller N, Drouaud J, Grelon M, Copenhaver GP, Mezard C, Kelly KA, Henderson IR. Epigenetic remodeling of meiotic crossover frequency in Arabidopsis thaliana DNA methyltransferase mutants. PLoS Genet, 2012, 8: e1002844
Choi K, Zhao X, Kelly KA, Venn O, Higgins JD, Yelina NE, Hardcastle TJ, Ziolkowski PA, Copenhaver GP, Franklin FC, McVean G, Henderson IR. Arabidopsis meiotic crossover hot spots overlap with H2A.Z nucleosomes at gene promoters. Nat Genet, 2013, 45: 1327–1336
Wijnker E, Velikkakam JG, Ding J, Becker F, Klasen JR, Rawat V, Rowan BA, de Jong DF, de Snoo CB, Zapata L, Huettel B, de Jong H, Ossowski S, Weigel D, Koornneef M, Keurentjes JJ, Schneeberger K. The genomic landscape of meiotic crossovers and gene conversions in Arabidopsis thaliana. eLIFE, 2013, 2: e01426
Melamed-Bessudo C, Levy AA. Deficiency in DNA methylation increases meiotic crossover rates in euchromatic but not in heterochromatic regions in Arabidopsis. Proc Natl Acad Sci USA, 2012, 109: E981–988
Colomé-Tatché M, Cortijo S, Wardenaar R, Morgado L, Lahouze B, Sarazin A, Etcheverry M, Martin A, Feng S, Duvernois-Berthet E, Labadie K, Wincker P, Jacobsen SE, Jansen RC, Colot V, Johannes F. Features of the Arabidopsis recombination landscape resulting from the combined loss of sequence variation and DNA methylation. Proc Natl Acad Sci USA, 2012, 109: 16240–16245
Mirouze M, Lieberman-Lazarovich M, Aversano R, Bucher E, Nicolet J, Reinders J, Paszkowski J. Loss of DNA methylation affects the recombination landscape in Arabidopsis. Proc Natl Acad Sci USA, 2012, 109: 5880–5885
Gore MA, Chia JM, Elshire RJ, Sun Q, Ersoz ES, Hurwitz BL, Peiffer JA, McMullen MD, Grills GS, Ross-Ibarra J, Ware DH, Buckler ES. A first-generation haplotype map of maize. Science, 2009, 326: 1115–1117
International Barley Genome Sequencing Consortium. A physical, genetic and functional sequence assembly of the barley genome. Nature, 2012, 491: 711–716
Tomato Genome Consortium. The tomato genome sequence provides insights into fleshy fruit evolution. Nature, 2012, 485: 635–641
Wei F, Zhang J, Zhou S, He R, Schaeffer M, Collura K, Kudrna D, Faga BP, Wissotski M, Golser W, Rock SM, Graves TA, Fulton RS, Coe E, Schnable PS, Schwartz DC, Ware D, Clifton SW, Wilson RK, Wing RA. The physical and genetic framework of the maize B73 genome. PLoS Genet, 2009, 5: e1000715
International Wheat Genome Sequencing Consortium (IWGSC). A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science, 2014, 345: 1251788-1–1251788-11
Dooner HK, Martínez-Férez IM. Recombination occurs uniformly within the bronze gene, a meiotic recombination hotspot in the maize genome. Plant Cell, 1997, 9: 1633–1646
Dooner HK, He L. Maize genome structure variation: interplay between retrotransposon polymorphisms and genic recombination. Plant Cell, 2008, 20: 249–258
Fu H, Zheng Z, Dooner HK. Recombination rates between adjacent genic and retrotransposon regions in maize vary by 2 orders of magnitude. Proc Natl Acad Sci USA, 2002, 99: 1082–1087
Fu H, Park W, Yan X, Zheng Z, Shen B, Dooner HK. The highly recombinogenic bz locus lies in an unusually gene-rich region of the maize genome. Proc Natl Acad Sci USA, 2001, 98: 8903–8908
Liu S, Yeh CT, Ji T, Ying K, Wu H, Tang HM, Fu Y, Nettleton D, Schnable PS. Mu transposon insertion sites and meiotic recombination events co-localize with epigenetic marks for open chromatin across the maize genome. PLoS Genet, 2009, 5: e1000733
Yao H, Zhou Q, Li J, Smith H, Yandeau M, Nikolau BJ, Schnable PS. Molecular characterization of meiotic recombination across the 140-kb multigenic a1-sh2 interval of maize. Proc Natl Acad Sci USA, 2002, 99: 6157–6162
Drouaud J, Khademian H, Giraut L, Zanni V, Bellalou S, Henderson IR, Falque M, Mézard C. Contrasted patterns of crossover and non-crossover at Arabidopsis thaliana meiotic recombination hotspots. PLoS Genet, 2013, 9: e1003922
Hellsten U, Wright KM, Jenkins J, Shu S, Yuan Y, Wessler SR, Schmutz J, Willis JH, Rokhsar DS. Fine-scale variation in meiotic recombination in Mimulus inferred from population shotgun sequencing. Proc Natl Acad Sci USA, 2013, 110: 19478–19482
Venters BJ, Pugh BF. How eukaryotic genes are transcribed. Crit Rev Biochem Mol Biol, 2009, 44: 117–141
Chodavarapu RK, Feng S, Bernatavichute YV, Chen PY, Stroud H, Yu Y, Hetzel JA, Kuo F, Kim J, Cokus SJ, Casero D, Bernal M, Huijser P, Clark AT, Krämer U, Merchant SS, Zhang X, Jacobsen SE, Pellegrini M. Relationship between nucleosome positioning and DNA methylation. Nature, 2010, 466: 388–392
Zilberman D, Coleman-Derr D, Ballinger T, Henikoff S. Histone H2A.Z and DNA methylation are mutually antagonistic chromatin marks. Nature, 2008, 456: 125–129
Coleman-Derr D, Zilberman D. Deposition of histone variant H2A.Z within gene bodies regulates responsive genes. PLoS Genet, 2012, 8: e1002988
Kumar SV, Wigge PA. H2A.Z-containing nucleosomes mediate the thermosensory response in Arabidopsis. Cell, 2010, 140: 136–147
Zhang X, Bernatavichute YV, Cokus S, Pellegrini M, Jacobsen SE. Genome-wide analysis of mono-, di- and trimethylation of histone H3 lysine 4 in Arabidopsis thaliana. Genome Biol, 2009, 10: R62
Wu TC, Lichten M. Meiosis-induced double-strand break sites determined by yeast chromatin structure. Science, 1994, 263: 515–518
Borde V, Robine N, Lin W, Bonfils S, Géli V, Nicolas A. Histone H3 lysine 4 trimethylation marks meiotic recombination initiation sites. EMBO J, 2009, 28: 99–111
Acquaviva L, Székvölgyi L, Dichtl B, Dichtl BS, de La Roche Saint-André C, Nicolas A, Géli V. The COMPASS subunit Spp1 links histone methylation to initiation of meiotic recombination. Science, 2013, 339: 215–218
Sommermeyer V, Béneut C, Chaplais E, Serrentino ME, Borde V. Spp1, a member of the Set1 Complex, promotes meiotic DSB formation in promoters by tethering histone H3K4 methylation sites to chromosome axes. Mol Cell, 2013, 49: 43–54
Sollier J, Lin W, Soustelle C, Suhre K, Nicolas A, Géli V, de La Roche Saint-André C. Set1 is required for meiotic S-phase onset, double-strand break formation and middle gene expression. EMBO J, 2004, 23: 1957–1967
Kleckner N. Chiasma formation: chromatin/axis interplay and the role(s) of the synaptonemal complex. Chromosoma, 2006, 115: 175–194
Panizza S, Mendoza MA, Berlinger M, Huang L, Nicolas A, Shirahige K, Klein F. Spo11-accessory proteins link double-strand break sites to the chromosome axis in early meiotic recombination. Cell, 2011, 146: 372–383
Blat Y, Protacio RU, Hunter N, Kleckner N. Physical and functional interactions among basic chromosome organizational features govern early steps of meiotic chiasma formation. Cell, 2002, 111: 791–802
Perrella G, Consiglio MF, Aiese-Cigliano R, Cremona G, Sanchez-Moran E, Barra L, Errico A, Bressan RA, Franklin FC, Conicella C. Histone hyperacetylation affects meiotic recombination and chromosome segregation in Arabidopsis. Plant J, 2010, 62: 796–806
Jeffreys AJ, Allen MJ, Armour JAL, Collick A, Dubrova Y, Fretwell N, Guram T, Jobling M, May CA, Neil DL, Neumann R. Mutation processes at human minisatellites. Electrophoresis, 1995, 16: 1577–1585
Paape T, Zhou P, Branca A, Briskine R, Young N, Tiffin P. Fine-scale population recombination rates, hotspots, and correlates of recombination in the Medicago truncatula genome. Genome Biol Evol, 2012, 4: 726–737
Cao J, Schneeberger K, Ossowski S, Günther T, Bender S, Fitz J, Koenig D, Lanz C, Stegle O, Lippert C, Wang X, Ott F, Müller J, Alonso-Blanco C, Borgwardt K, Schmid KJ, Weigel D. Whole-genome sequencing of multiple Arabidopsis thaliana populations. Nat Genet, 2011, 43: 956–963
Duret L, Galtier N. Biased gene conversion and the evolution of mammalian genomic landscapes. Annu Rev Genomics Hum Genet, 2009, 10: 285–311
Baudat F, Buard J, Grey C, Fledel-Alon A, Ober C, Przeworski M, Coop G, de Massy B. PRDM9 is a major determinant of meiotic recombination hotspots in humans and mice. Science, 2010, 327: 836–840
Myers S, Bowden R, Tumian A, Bontrop RE, Freeman C, MacFie TS, McVean G, Donnelly P. Drive against hotspot motifs in primates implicates the PRDM9 gene in meiotic recombination. Science, 2010, 327: 876–879
Parvanov ED, Petkov PM, Paigen K. Prdm9 controls activation of mammalian recombination hotspots. Science, 2010, 327: 835
Brick K, Smagulova F, Khil P, Camerini-Otero RD, Petukhova GV. Genetic recombination is directed away from functional genomic elements in mice. Nature, 2012, 485: 642–645
Oliver PL, Goodstadt L, Bayes JJ, Birtle Z, Roach KC, Phadnis N, Beatson SA, Lunter G, Malik HS, Ponting CP. Accelerated evolution of the Prdm9 speciation gene across diverse metazoan taxa. PLoS Genet, 2009, 5: e1000753
Haag JR, Pikaard CS. Multisubunit RNA polymerases IV and V: purveyors of non-coding RNA for plant gene silencing. Nat Rev Mol Cell Biol, 2011, 12: 483–492
Zhong X, Hale CJ, Law JA, Johnson LM, Feng S, Tu A, Jacobsen SE. DDR complex facilitates global association of RNA polymerase V to promoters and evolutionarily young transposons. Nat Struct Mol Biol, 2012, 19: 870–875
Law JA, Du J, Hale CJ, Feng S, Krajewski K, Palanca AM, Strahl BD, Patel DJ, Jacobsen SE. Polymerase IV occupancy at RNA-directed DNA methylation sites requires SHH1. Nature, 2013, 498: 385–389
Johnson LM, Du J, Hale CJ, Bischof S, Feng S, Chodavarapu RK, Zhong X, Marson G, Pellegrini M, Segal DJ, Patel DJ, Jacobsen SE. SRA- and SET-domain-containing proteins link RNA polymerase V occupancy to DNA methylation. Nature, 2014, 507: 124–128
Lippman Z, Gendrel AV, Black M, Vaughn MW, Dedhia N, McCombie WR, Lavine K, Mittal V, May B, Kasschau KD, Carrington JC, Doerge RW, Colot V, Martienssen R. Role of transposable elements in heterochromatin and epigenetic control. Nature, 2004, 430: 471–476
Cokus SJ, Feng S, Zhang X, Chen Z, Merriman B, Haudenschild CD, Pradhan S, Nelson SF, Pellegrini M, Jacobsen SE. Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature, 2008, 452: 215–219
Saze H, Shiraishi A, Miura A, Kakutani T. Control of genic DNA methylation by a jmjC domain-containing protein in Arabidopsis thaliana. Science, 2008, 319: 462–465
Zhang X, Yazaki J, Sundaresan A, Cokus S, Chan SW, Chen H, Henderson IR, Shinn P, Pellegrini M, Jacobsen SE, Ecker JR. Genome-wide high-resolution mapping and functional analysis of DNA methylation in Arabidopsis. Cell, 2006, 126: 1189–1201
Copenhaver GP, Nickel K, Kuromori T, Benito MI, Kaul S, Lin X, Bevan M, Murphy G, Harris B, Parnell LD, McCombie WR, Martienssen RA, Marra M, Preuss D. Genetic definition and sequence analysis of Arabidopsis centromeres. Science, 1999, 286: 2468–2474
Lamb NE, Sherman SL, Hassold TJ. Effect of meiotic recombination on the production of aneuploid gametes in humans. Cytogenet Genome Res, 2005, 111: 250–255
Maloisel L, Rossignol JL. Suppression of crossing-over by DNA methylation in Ascobolus. Genes Dev, 1998, 12: 1381–1389
Kankel MW, Ramsey DE, Stokes TL, Flowers SK, Haag JR, Jeddeloh JA, Riddle NC, Verbsky ML, Richards EJ. Arabidopsis MET1 cytosine methyltransferase mutants. Genetics, 2003, 163: 1109–1122
Mathieu O, Reinders J, Caikovski M, Smathajitt C, Paszkowski J. Transgenerational stability of the Arabidopsis epigenome is coordinated by CG methylation. Cell, 2007, 130: 851–862
Singer T, Yordan C, Martienssen RA. Robertson’s Mutator transposons in A. thaliana are regulated by the chromatin-remodeling gene decrease in DNA Methylation (DDM1). Genes Dev, 2001, 15: 591–602
Tariq M, Saze H, Probst AV, Lichota J, Habu Y, Paszkowski J. Erasure of CpG methylation in Arabidopsis alters patterns of histone H3 methylation in heterochromatin. Proc Natl Acad Sci USA, 2003, 100: 8823–8827
Vongs A, Kakutani T, Martienssen RA, Richards EJ. Arabidopsis thaliana DNA methylation mutants. Science, 1993, 260: 1926–1928
Martini E, Diaz RL, Hunter N, Keeney S. Crossover homeostasis in yeast meiosis. Cell, 2006, 126: 285–295
Thacker D, Mohibullah N, Zhu X, Keeney S. Homologue engagement controls meiotic DNA break number and distribution. Nature, 2014, 510: 241–246
Komiya R, Ohyanagi H, Niihama M, Watanabe T, Nakano M, Kurata N, Nonomura K. Rice germline-specific Argonaute MEL1 protein binds to phasiRNAs generated from more than 700 lincRNAs. Plant J, 2014, 78: 385–397
Nonomura K, Morohoshi A, Nakano M, Eiguchi M, Miyao A, Hirochika H, Kurata N. A germ cell specific gene of the ARGONAUTE family is essential for the progression of premeiotic mitosis and meiosis during sporogenesis in rice. Plant Cell, 2007, 19: 2583–2594
Kleckner N, Zickler D, Jones GH, Dekker J, Padmore R, Henle J, Hutchinson J. A mechanical basis for chromosome function. Proc Natl Acad Sci USA, 2004, 101: 12592–12597
Kim KP, Weiner BM, Zhang L, Jordan A, Dekker J, Kleckner N. Sister cohesion and structural axis components mediate homolog bias of meiotic recombination. Cell, 2010, 143: 924–937
Schwacha A, Kleckner N. Interhomolog bias during meiotic recombination: meiotic functions promote a highly differentiated interhomolog-only pathway. Cell, 1997, 90: 1123–1135
Xu L, Weiner BM, Kleckner N. Meiotic cells monitor the status of the interhomolog recombination complex. Genes Dev, 1997, 11: 106–118
Kugou K, Fukuda T, Yamada S, Ito M, Sasanuma H, Mori S, Katou Y, Itoh T, Matsumoto K, Shibata T, Shirahige K, Ohta K. Rec8 guides canonical Spo11 distribution along yeast meiotic chromosomes. Mol Biol Cell, 2009, 20: 3064–3076
Mao-Draayer Y, Galbraith AM, Pittman DL, Cool M, Malone RE. Analysis of meiotic recombination pathways in the yeast Saccharomyces cerevisiae. Genetics, 1996, 144: 71–86
Börner GV, Barot A, Kleckner N. Yeast Pch2 promotes domainal axis organization, timely recombination progression, and arrest of defective recombinosomes during meiosis. Proc Natl Acad Sci USA, 2008, 105: 3327–3332
Deshong AJ, Ye AL, Lamelza P, Bhalla N. A quality control mechanism coordinates meiotic prophase events to promote crossover assurance. PLoS Genet, 2014, 10: e1004291
Dernburg AF, McDonald K, Moulder G, Barstead R, Dresser M, Villeneuve AM. Meiotic recombination in C. elegans initiates by a conserved mechanism and is dispensable for homologous chromosome synapsis. Cell, 1998, 94: 387–398
Giroux CN, Dresser ME, Tiano HF. Genetic control of chromosome synapsis in yeast meiosis. Genome, 1989, 31: 88–94
McKim KS, Green-Marroquin BL, Sekelsky JJ, Chin G, Steinberg C, Khodosh R, Hawley RS. Meiotic synapsis in the absence of recombination. Science, 1998, 279: 876–878
Romanienko PJ, Camerini-Otero RD. The mouse Spo11 gene is required for meiotic chromosome synapsis. Mol Cell, 2000, 6: 975–987
Page SL, Hawley RS. Chromosome choreography: the meiotic ballet. Science, 2003, 301: 785–789
Libuda DE, Uzawa S, Meyer BJ, Villeneuve AM. Meiotic chromosome structures constrain and respond to designation of crossover sites. Nature, 2013, 502: 703–706
Osman K, Higgins JD, Sanchez-Moran E, Armstrong SJ, Franklin FCH. Pathways to meiotic recombination in Arabidopsis thaliana. New Phytol, 2011, 190: 523–544
Sanchez-Moran E, Santos JL, Jones GH, Franklin FCH. ASY1 mediates AtDMC1-dependent interhomolog recombination during meiosis in Arabidopsis. Genes Dev, 2007, 21: 2220–2233
Caryl AP, Armstrong SJ, Jones GH, Franklin FC. A homologue of the yeast HOP1 gene is inactivated in the Arabidopsis meiotic mutant asy1. Chromosoma, 2000, 109: 62–71
Armstrong SJ, Caryl AP, Jones GH, Franklin FCH. Asy1, a protein required for meiotic chromosome synapsis, localizes to axis-associated chromatin in Arabidopsis and Brassica. J Cell Sci, 2002, 115: 3645–3655
Cai X, Dong F, Edelmann RE, Makaroff CA. The Arabidopsis SYN1 cohesin protein is required for sister chromatid arm cohesion and homologous chromosome pairing. J Cell Sci, 2003, 116: 2999–3007
Bhatt AM, Lister C, Page T, Fransz P, Findlay K, Jones GH, Dickinson HG, Dean C. The DIF1 gene of Arabidopsis is required for meiotic chromosome segregation and belongs to the REC8/RAD21 cohesin gene family. Plant J, 1999, 19: 463–472
Bai X, Peirson BN, Dong F, Xue C, Makaroff CA. Isolation and characterization of SYN1, a RAD21-like gene essential for meiosis in Arabidopsis. Plant Cell, 1999, 11: 417–430
Chelysheva L, Diallo S, Vezon D, Gendrot G, Vrielynck N, Belcram K, Rocques N, Márquez-Lema A, Bhatt AM, Horlow C, Mercier R, Mézard C, Grelon M. AtREC8 and AtSCC3 are essential to the monopolar orientation of the kinetochores during meiosis. J Cell Sci, 2005, 118: 4621–4632
Lam WS, Yang X, Makaroff CA. Characterization of Arabidopsis thaliana SMC1 and SMC3: evidence that AtSMC3 may function beyond chromosome cohesion. J Cell Sci, 2005, 118: 3037–3048
Glynn EF, Megee PC, Yu HG, Mistrot C, Unal E, Koshland DE, DeRisi JL, Gerton JL. Genome-wide mapping of the cohesin complex in the yeast Saccharomyces cerevisiae. PLoS Biol, 2004, 2: E259
Watanabe Y. Shugoshin: guardian spirit at the centromere. Curr Opin Cell Biol, 2005, 17: 590–595
Buonomo SB, Clyne RK, Fuchs J, Loidl J, Uhlmann F, Nasmyth K. Disjunction of homologous chromosomes in meiosis I depends on proteolytic cleavage of the meiotic cohesin Rec8 by separin. Cell, 2000, 103: 387–398
Kitajima TS, Miyazaki Y, Yamamoto M, Watanabe Y. Rec8 cleavage by separase is required for meiotic nuclear divisions in fission yeast. EMBO J, 2003, 22: 5643–5653
Zamariola L, De Storme N, Vannerum K, Vandepoele K, Armstrong SJ, Franklin FC, Geelen D. SHUGOSHINs and PATRONUS protect meiotic centromere cohesion in Arabidopsis thaliana. Plant J, 2014, 77: 782–794
Cromer L, Jolivet S, Horlow C, Chelysheva L, Heyman J, De Jaeger G, Koncz C, De Veylder L, Mercier R. Centromeric cohesion is protected twice at meiosis, by SHUGOSHINs at anaphase I and by PATRONUS at interkinesis. Curr Biol, 2013, 23: 2090–2099
Kitajima TS, Kawashima SA, Watanabe Y. The conserved kinetochore protein shugoshin protects centromeric cohesion during meiosis. Nature, 2004, 427: 510–517
Hamant O, Golubovskaya I, Meeley R, Fiume E, Timofejeva L, Schleiffer A, Nasmyth K, Cande WZ. A REC8-dependent plant Shugoshin is required for maintenance of centromeric cohesion during meiosis and has no mitotic functions. Curr Biol, 2005, 15: 948–954
Hollister JD, Arnold BJ, Svedin E, Xue KS, Dilkes BP, Bomblies K. Genetic adaptation associated with genome-doubling in autotetr-aploid Arabidopsis arenosa. PLoS Genet, 2012, 8: e1003093
Yant L, Hollister JD, Wright KM, Arnold BJ, Higgins JD, Franklin FC, Bomblies K. Meiotic adaptation to genome duplication in Arabidopsis arenosa. Curr Biol, 2013, 23: 2151–2156
Sun Y, Ambrose JH, Haughey BS, Webster TD, Pierrie SN, Muñoz DF, Wellman EC, Cherian S, Lewis SM, Berchowitz LE, Copenhaver GP. Deep genome-wide measurement of meiotic gene conversion using tetrad analysis in Arabidopsis thaliana. PLoS Genet, 2012, 8: e1002968
Yang S, Yuan Y, Wang L, Li J, Wang W, Liu H, Chen JQ, Hurst LD, Tian D. Great majority of recombination events in Arabidopsis are gene conversion events. Proc Natl Acad Sci USA, 2012, 109: 20992–20997
Lu P, Han X, Qi J, Yang J, Wijeratne AJ, Li T, Ma H. Analysis of Arabidopsis genome-wide variations before and after meiosis and meiotic recombination by resequencing Landsberg erecta and all four products of a single meiosis. Genome Res, 2012, 22: 508–518
Qi J, Chen Y, Copenhaver GP, Ma H. Detection of genomic variations and DNA polymorphisms and impact on analysis of meiotic recombination and genetic mapping. Proc Natl Acad Sci USA, 2014, 111: 10007–10012
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at link.springer.com
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0), which permits use, duplication, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Yelina, N., Diaz, P., Lambing, C. et al. Epigenetic control of meiotic recombination in plants. Sci. China Life Sci. 58, 223–231 (2015). https://doi.org/10.1007/s11427-015-4811-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-015-4811-x