Abstract
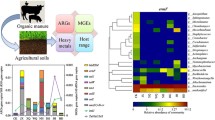
The practice of utilizing animal manures on land is widespread in agriculture, but it has raised concerns about the possible spread of antibiotic resistance genes (ARGs) and the potential risk it poses to public health through food production. Fermentation bed culture is an effective circular agricultural practice commonly utilized in pig farming that minimizes the environmental impact of livestock farming. However, this method generates a significant amount of fermentation bed waste (FBW), which can be turned into organic fertilizer for land application. The objective of this research was to examine the impacts of amending agricultural soil samples with swine manure–derived FBW on microbial communities, mobile genetic elements (MGEs), and ARG profiles over different periods. The study findings indicated that the amendment of swine manure–derived FBW significantly increased the diversity and abundance of ARGs and MGEs during the early stages of amendment, but this effect diminished over time, and after 12 months of FBW amendments, the levels returned to those comparable to control samples. The shift in the bacterial communities played a significant role in shaping the patterns of ARGs. Actinobacteriota, Proteobacteria, and Bacteroidetes were identified as the primary potential hosts of ARGs through metagenomic binning analysis. Furthermore, the pH of soil samples was identified as the most important property in driving the composition of the bacterial community and soil resistome. These findings provided valuable insights into the temporal patterns and dissemination risks of ARGs in FBW-amended agriculture soil, which could contribute to the development of effective strategies to manage the dissemination risks of FBW-derived ARGs.






Similar content being viewed by others
Data availability
The raw metagenomic sequence data were publicly available in the NCBI BioProject database with the accession number of PRJNA953804.
References
Ben W, Wang J, Pan X, Qiang Z (2017) Dissemination of antibiotic resistance genes and their potential removal by on-farm treatment processes in nine swine feedlots in shandong province, china. Chemosphere 167:262–268. https://doi.org/10.1016/j.chemosphere.2016.10.013
Chaumeil PA, Mussig AJ, Hugenholtz P, Parks DH (2020) Gtdb-tk: A toolkit to classify genomes with the genome taxonomy database. Bioinformatics 36:1925–1927. https://doi.org/10.1093/bioinformatics/btz848
Checcucci A, Trevisi P, Luise D, Modesto M, Blasioli S, Braschi I, Mattarelli P (2020) Exploring the animal waste resistome: The spread of antimicrobial resistance genes through the use of livestock manure. Front Microbiol 11:1416. https://doi.org/10.3389/fmicb.2020.01416
Chen C, Pankow CA, Oh M, Heath LS, Zhang L, Du P, Xia K, Pruden A (2019) Effect of antibiotic use and composting on antibiotic resistance gene abundance and resistome risks of soils receiving manure-derived amendments. Environ Int 128:233–243. https://doi.org/10.1016/j.envint.2019.04.043
Chen Q, An X, Li H, Su J, Ma Y, Zhu Y-G (2016) Long-term field application of sewage sludge increases the abundance of antibiotic resistance genes in soil. Environ Int 92:1–10. https://doi.org/10.1016/j.envint.2016.03.026
Chen Q, Liu B, Wang J, Che J, Liu G, Guan X (2017) Diversity and dynamics of the bacterial community involved in pig manure biodegradation in a microbial fermentation bed system. Annals Microbiol 67:491–500. https://doi.org/10.1007/s13213-017-1278-y
Chen SF, Zhou YQ, Chen YR, Gu J (2018) Fastp: An ultra-fast all-in-one fastq preprocessor. Bioinformatics 34:884–890. https://doi.org/10.1093/bioinformatics/bty560
Cheng JH, Tang XY, Cui JF (2019) Effect of long-term manure slurry application on the occurrence of antibiotic resistance genes in arable purple soil (entisol). Sci Total Environ 647:853–861. https://doi.org/10.1016/j.scitotenv.2018.08.028
Christou A, Agüera A, Bayona JM, Cytryn E, Fotopoulos V, Lambropoulou D, Manaia CM, Michael C, Revitt M, Schröder P (2017) The potential implications of reclaimed wastewater reuse for irrigation on the agricultural environment: The knowns and unknowns of the fate of antibiotics and antibiotic resistant bacteria and resistance genes–a review. Water Res 123:448–467. https://doi.org/10.1016/j.watres.2017.07.004
D’Costa VM, King CE, Kalan L, Morar M, Sung WWL, Schwarz C, Froese D, Zazula G, Calmels F, Debruyne R, Golding GB, Poinar HN, Wright GD (2011) Antibiotic resistance is ancient. Nature 477:457–461. https://doi.org/10.1038/nature10388
Du S, Shen J-P, Hu H-W, Wang J-T, Han L-L, Sheng R, Wei W-X, Fang Y-T, Zhu Y-G, Zhang L-M (2020) Large-scale patterns of soil antibiotic resistome in chinese croplands. Sci Total Environ 712:136418. https://doi.org/10.1016/j.scitotenv.2019.136418
Ezugworie FN, Igbokwe VC, Onwosi CO (2021) Proliferation of antibiotic-resistant microorganisms and associated genes during composting: An overview of the potential impacts on public health, management and future. Sci Total Environ 784:147191. https://doi.org/10.1016/j.scitotenv.2021.147191
Fu LM, Niu BF, Zhu ZW, Wu ST, Li WZ (2012) Cd-hit: Accelerated for clustering the next-generation sequencing data. Bioinformatics 28:3150–3152. https://doi.org/10.1093/bioinformatics/bts565
Gu Q, Sun M, Lin T, Zhang Y, Wei X, Wu S, Zhang S, Pang R, Wang J, Ding Y, Liu Z, Chen L, Chen W, Lin X, Zhang J, Chen M, Xue L, Wu Q (2021) Characteristics of antibiotic resistance genes and antibiotic-resistant bacteria in full-scale drinking water treatment system using metagenomics and culturing. Front Microbiol 12:798442. https://doi.org/10.3389/fmicb.2021.798442
Guan Y, Xue X, Jia J, Li X, Xing H, Wang Z (2022) Metagenomic assembly and binning analyses the prevalence and spread of antibiotic resistome in water and fish gut microbiomes along an environmental gradient. J Environ Manage 318:115521. https://doi.org/10.1016/j.jenvman.2022.115521
Han XM, Hu HW, Chen QL, Yang LY, Li HL, Zhu YG, Li XZ, Ma YB (2018) Antibiotic resistance genes and associated bacterial communities in agricultural soils amended with different sources of animal manures. Soil Biol Biochem 126:91–102. https://doi.org/10.1016/j.soilbio.2018.08.018
He Y, Yuan Q, Mathieu J, Stadler L, Senehi N, Sun R, Alvarez PJJ (2020) Antibiotic resistance genes from livestock waste: Occurrence, dissemination, and treatment. NPJ Clean Water 3:4. https://doi.org/10.1038/s41545-020-0051-0
Heuer H, Schmitt H, Smalla K (2011) Antibiotic resistance gene spread due to manure application on agricultural fields. Curr Opin Microbiol 14:236–243. https://doi.org/10.1016/j.mib.2011.04.009
Hyatt D, Chen GL, LoCascio PF, Land ML, Larimer FW, Hauser LJ (2010) Prodigal: Prokaryotic gene recognition and translation initiation site identification. Bmc Bioinformatics 11. https://doi.org/10.1186/1471-2105-11-119
Kang DD, Li F, Kirton E, Thomas A, Egan R, An H, Wang Z (2019) Metabat 2: An adaptive binning algorithm for robust and efficient genome reconstruction from metagenome assemblies. PeerJ 7:e7359. https://doi.org/10.7717/peerj.7359
Kumar K, Gupta SC, Baidoo SK, Chander Y, Rosen CJ (2005) Antibiotic uptake by plants from soil fertilized with animal manure. J Environ Qual 34:2082–2085. https://doi.org/10.2134/jeq2005.0026
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with bowtie 2. Nat Methods 9:357-U354. https://doi.org/10.1038/Nmeth.1923
Li DH, Liu CM, Luo RB, Sadakane K, Lam TW (2015) Megahit: An ultra-fast single-node solution for large and complex metagenomics assembly via succinct de bruijn graph. Bioinformatics 31:1674–1676. https://doi.org/10.1093/bioinformatics/btv033
Li LG, Xia Y, Zhang T (2017) Co-occurrence of antibiotic and metal resistance genes revealed in complete genome collection. Isme J 11:651–662. https://doi.org/10.1038/ismej.2016.155
Li T, Li R, Cao Y, Tao C, Deng X, Ou Y, Liu H, Shen Z, Li R, Shen Q (2022) Soil antibiotic abatement associates with the manipulation of soil microbiome via long-term fertilizer application. J Hazard Mater 439:129704. https://doi.org/10.1016/j.jhazmat.2022.129704
Mbanga J, Amoako DG, Abia ALK, Allam M, Ismail A, Essack SY (2021) Genomic analysis of enterococcus spp. Isolated from a wastewater treatment plant and its associated waters in umgungundlovu district, south africa. Front Microbiol 12:648454. https://doi.org/10.3389/fmicb.2021.648454
Morrison RS, Johnston LJ, Hilbrands AM (2007) The behaviour, welfare, growth performance and meat quality of pigs housed in a deep-litter, large group housing system compared to a conventional confinement system. Appl Anim Behav Sci 103:12–24. https://doi.org/10.1016/j.applanim.2006.04.002
Mulchandani R, Wang Y, Gilbert M, Van Boeckel TP (2023) Global trends in antimicrobial use in food-producing animals: 2020 to 2030. PLOS Global Public Health 3:e0001305. https://doi.org/10.1371/journal.pgph.0001305
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW (2015) Checkm: Assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res 25:1043–1055. https://doi.org/10.1101/gr.186072.114
Patro R, Duggal G, Love MI, Irizarry RA, Kingsford C (2017) Salmon provides fast and bias-aware quantification of transcript expression. Nat Methods 14:417-+. https://doi.org/10.1038/nmeth.4197
Pérez P, Solís-Tejeda MÁ, Lango-Reynoso F, Díaz-Rivera PD-R, Aguilar-Ávila J, Asiain-Hoyos A (2022) Deep litter pig production system as a sustainable alternative for small farmers. Agrociencia. https://doi.org/10.47163/agrociencia.v56i6.2755
Peterson E, Kaur P (2018) Antibiotic resistance mechanisms in bacteria: Relationships between resistance determinants of antibiotic producers, environmental bacteria, and clinical pathogens. Front Microbiol 9:2928. https://doi.org/10.3389/fmicb.2018.02928
Van Boeckel TP, Brower C, Gilbert M, Grenfell BT, Levin SA, Robinson TP, Teillant A, Laxminarayan R (2015) Global trends in antimicrobial use in food animals. P Natl Acad Sci USA 112:5649–5654. https://doi.org/10.1073/pnas.1503141112
Van Boeckel TP, Glennon EE, Chen D, Gilbert M, Robinson TP, Grenfell BT, Levin SA, Bonhoeffer S, Laxminarayan R (2017) Reducing antimicrobial use in food animals. Science 357:1350–1352. https://doi.org/10.1126/science.aao1495
Van Boeckel TP, Pires J, Silvester R, Zhao C, Song J, Criscuolo NG, Gilbert M, Bonhoeffer S, Laxminarayan R (2019) Global trends in antimicrobial resistance in animals in low- and middle-income countries. Science 365:1266-+. https://doi.org/10.1126/science.aaw1944
Van den Meersche T, Rasschaert G, Vanden Nest T, Haesebrouck F, Herman L, Van Coillie E, Van Weyenberg S, Daeseleire E, Heyndrickx M (2020) Longitudinal screening of antibiotic residues, antibiotic resistance genes and zoonotic bacteria in soils fertilized with pig manure. Environ Sci Pollut R 27:28016–28029. https://doi.org/10.1007/s11356-020-09119-y
Wang J, Ben W, Yang M, Zhang Y, Qiang Z (2016) Dissemination of veterinary antibiotics and corresponding resistance genes from a concentrated swine feedlot along the waste treatment paths. Environ Int 92–93:317–323. https://doi.org/10.1016/j.envint.2016.04.020
Williams-Nguyen J, Sallach JB, Bartelt-Hunt S, Boxall AB, Durso LM, McLain JE, Singer RS, Snow DD, Zilles JL (2016) Antibiotics and antibiotic resistance in agroecosystems: State of the science. J Environ Qual 45:394–406. https://doi.org/10.2134/jeq2015.07.0336
Xu Y, Li P, Gao Y, Li Y, Li S, Li Y, Sui P, Wang X, Gao W, Chen Y (2022) Sustainability evaluation for a circular maize-pig system driven by indigenous microbes: A case study in northeast china. Ecosyst Health Sustain 8:2080591. https://doi.org/10.1080/20964129.2022.2080591
Yang Y, Jiang X, Chai B, Ma L, Li B, Zhang A, Cole JR, Tiedje JM, Zhang T (2016) Args-oap: Online analysis pipeline for antibiotic resistance genes detection from metagenomic data using an integrated structured arg-database. Bioinformatics 32:2346–2351. https://doi.org/10.1093/bioinformatics/btw136
Yin XL, Jiang XT, Chai BL, Li LG, Yang Y, Cole JR, Tiedje JM, Zhang T (2018) Args-oap v2.0 with an expanded sarg database and hidden markov models for enhancement characterization and quantification of antibiotic resistance genes in environmental metagenomes. Bioinformatics 34:2263–2270. https://doi.org/10.1093/bioinformatics/bty053
Yu P, Dong P, Zou Y, Wang H (2023) Effect of ph on the mitigation of extracellular/intracellular antibiotic resistance genes and antibiotic resistance pathogenic bacteria during anaerobic fermentation of swine manure. Bioresour Technol 373:128706. https://doi.org/10.1016/j.biortech.2023.128706
Zhang AN, Gaston JM, Dai CL, Zhao S, Poyet M, Groussin M, Yin X, Li LG, van Loosdrecht MCM, Topp E, Gillings MR, Hanage WP, Tiedje JM, Moniz K, Alm EJ, Zhang T (2021) An omics-based framework for assessing the health risk of antimicrobial resistance genes. Nat Commun 12:4765. https://doi.org/10.1038/s41467-021-25096-3
Zhao RX, Yu K, Zhang JY, Zhang GJ, Huang J, Ma LP, Deng CF, Li XY, Li B (2020) Deciphering the mobility and bacterial hosts of antibiotic resistance genes under antibiotic selection pressure by metagenomic assembly and binning approaches. Water Research 186. https://doi.org/10.1016/j.watres.2020.116318
Zheng D, Yin G, Liu M, Hou L, Yang Y, Van Boeckel TP, Zheng Y, Li Y (2022) Global biogeography and projection of soil antibiotic resistance genes. Sci Adv 8:eabq8015. https://doi.org/10.1126/sciadv.abq8015
Zhou SY, Zhu D, Giles M, Yang XR, Daniell T, Neilson R, Zhu YG (2019) Phyllosphere of staple crops under pig manure fertilization, a reservoir of antibiotic resistance genes. Environ Pollut 252:227–235. https://doi.org/10.1016/j.envpol.2019.05.098. (Barking, Essex : 1987)
Zhou SY, Zhu D, Giles M, Daniell T, Neilson R, Yang XR (2020) Does reduced usage of antibiotics in livestock production mitigate the spread of antibiotic resistance in soil, earthworm guts, and the phyllosphere? Environ Int 136:105359. https://doi.org/10.1016/j.envint.2019.105359
Zhu L, Yuan L, Shuai XY, Lin ZJ, Sun YJ, Zhou ZC, Meng LX, Ju F, Chen H (2023) Deciphering basic and key traits of antibiotic resistome in influent and effluent of hospital wastewater treatment systems. Water Res 231:119614. https://doi.org/10.1016/j.watres.2023.119614
Zou Y, Xiao Y, Wang H, Fang T, Dong P (2020) New insight into fates of sulfonamide and tetracycline resistance genes and resistant bacteria during anaerobic digestion of manure at thermophilic and mesophilic temperatures. J Hazard Mater 384:121433. https://doi.org/10.1016/j.jhazmat.2019.121433
Acknowledgements
This work was supported by Fujian Special Fund for Scientific Research Institutes in the Public Interest (2020R1034002); Natural Science Foundation of Xiamen, China (3502Z202272400); and external cooperation project of Fujian Academy of Agricultural Sciences (DWHZ2021-15). The authors thank Yunfeng Agricultural and Animal Husbandry Cooperative, Maiping Li from the Baota District Animal Husbandry and Veterinary Service Center, and Xiyong Zhang for assisting with the sampling process.
Author information
Authors and Affiliations
Contributions
Zhizhen Pan, conceptualization, methodology, investigation, writing (original draft), visualization, data curation; Zheng Chen, methodology, investigation, resources; Liting Zhu, methodology, software, formal analysis; Ricardo David Avellán-Llaguno, methodology, software, formal analysis; Bo Liu, writing (review and editing), funding acquisition; Qiansheng Huang: conceptualization, methodology, writing (original draft), writing (review and editing), visualization, supervision, funding acquisition. All authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Diane Purchase
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pan, Z., Chen, Z., Zhu, L. et al. Antibiotic resistome and associated bacterial communities in agricultural soil following the amendments of swine manure–derived fermentation bed waste. Environ Sci Pollut Res 30, 104520–104531 (2023). https://doi.org/10.1007/s11356-023-29691-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-023-29691-3