Abstract
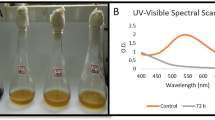
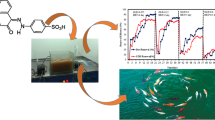
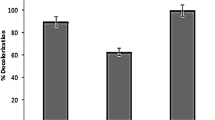
In this study, performance of hydrolysis acidification process treating simulated dyeing wastewater containing azo and anthraquinone dyes in different stages was investigated. The decolorization ratio, CODCr removal ratio, BOD5/CODCr value, and volatile fatty acids (VFAs) production were almost better in stage 1 than that in stage 2. Fourier transform infrared spectroscopy (FTIR) and gas chromatography-mass spectrometry (GC-MS) confirmed the biodegradation of Reactive Black 5 (RB5) and Remazol Brilliant Blue R (RBBR) in hydrolysis acidification process. Polymerase chain reaction-denaturing gradient gel electrophoresis (PCR-DGGE) analyses revealed that significant difference of microbial community structures existed in stage 1 and 2. The dominant species in stage 1 was related to Bacteroidetes group, while the dominant species in stage 2 was related to Bacteroidetes and Firmicutes groups. From the results, it could be speculated that different dyes’ structures might have significant influence on the existence and function of different bacterial species, which might supply information for bacteria screening and acclimation in the treatment of actual dyeing wastewater.





Similar content being viewed by others
References
Allen WPWB, Derby RE, Garland CE, Peret JM, Saltzman M (1973) Determination of color of water and wastewater by means of ADMI color values, vol 142 . Purdue Unive, Eng Ext SerProc 28th Ind Conf. edn
APHA, AWWA, WEF (1999) Standard Methods for the Examination of Water and Wastewater. 20 th edn., Washington DC.
Bai J, Xu H, Zhang Y, Peng Z, Xu G (2013) Combined industrial and domestic wastewater treatment by periodic allocating water hybrid hydrolysis acidification reactor followed by SBR. Biochem Eng J 70:115–119. doi:10.1016/j.bej.2012.10.009
Cui D, Li G, Zhao D, Gu X, Wang C, Zhao M (2012) Microbial community structures in mixed bacterial consortia for azo dye treatment under aerobic and anaerobic conditions. J Hazard Mater 221-222:185–192. doi:10.1016/j.jhazmat.2012.04.032
Cui D, Li G, Zhao M, Han S (2014) Decolourization of azo dyes by a newly isolated sp. strain Y3, and effects of various factors on biodegradation. Biotechnol Biotechnol Equip 28:478–486. doi:10.1080/13102818.2014.926053
Environmental statistics annual report (2013) (2014) http://zls.mep.gov.cn/hjtj/nb/2013tjnb/201411/t20141124_291868.htm.
Fanchiang JM, Tseng DH (2009) Degradation of anthraquinone dye C.I. Reactive blue 19 in aqueous solution by ozonation. Chemosphere 77:214–221. doi:10.1016/j.chemosphere.2009.07.038
Fernández A, Huang SY, Seston S, Xing J, Hickey R, Criddle C, Tiedje J (1999) How stable is stable? Function versus community composition. Appl Environ Microbiol 65:3697–3704
Forss J, Pinhassi J, Lindh M, Welander U (2013) Microbial diversity in a continuous system based on rice husks for biodegradation of the azo dyes reactive red 2 and reactive black 5. Bioresour Technol 130:681–688. doi:10.1016/j.biortech.2012.12.097
Huang Y, Cui C, Zhang D, Li L, Pan D (2015) Heterogeneous catalytic ozonation of dibutyl phthalate in aqueous solution in the presence of iron-loaded activated carbon. Chemosphere 119:295–301. doi:10.1016/j.chemosphere.2014.06.060
Jain K, Shah V, Chapla D, Madamwar D (2012) Decolorization and degradation of azo dye--reactive violet 5R by an acclimatized indigenous bacterial mixed cultures-SB4 isolated from anthropogenic dye contaminated soil. J Hazard Mater 213-214:378–386. doi:10.1016/j.jhazmat.2012.02.010
Khouni I, Marrot B, Amar RB (2012) Treatment of reconstituted textile wastewater containing a reactive dye in an aerobic sequencing batch reactor using a novel bacterial consortium. Sep Purif Technol 87:110–119. doi:10.1016/j.seppur.2011.11.030
Lane DJ (1991) 16S/23S rDNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. John Wiley & Sons, Chichester, England
Lei G, Ren H, Ding L, Wang F, Zhang X (2010) A full-scale biological treatment system application in the treated wastewater of pharmaceutical industrial park. Bioresour Technol 101:5852–5861. doi:10.1016/j.biortech.2010.03.025
McMullan G et al (2001) Microbial decolourisation and degradation of textile dyes. Appl Microbiol Biotechnol 56:81–87. doi:10.1007/s002530000587
Ministry of Environmental Protection of the People’s republic of China (2014) Environmental statistics annual report (2013). http://zls.mep.gov.cn/hjtj/nb/2013tjnb/201411/t20141124_291868.htm
Monserrat Castillo DB (2001) Characterisation of organic pollutants in textile wastewaters and landfill leachate by using toxicity-based fractionation methods followed by liquid and gas chromatography coupled to mass spectrometric detectionP.pdf. Anal Chim Acta 426:253–264
Muyzer G, Dewaal EC, Uitterlinden AG (1993) Profiling of complex microbial-populations by denaturing gradient gel-electrophoresis analysis of polymerase chain reaction-amplified genes-coding for 16S ribosomal-RNA. Appl Environ Microbiol 59:695–700
Novotny C, Dias N, Kapanen A, Malachova K, Vandrovcova M, Itavaara M, Lima N (2006) Comparative use of bacterial, algal and protozoan tests to study toxicity of azo- and anthraquinone dyes. Chemosphere 63:1436–1442. doi:10.1016/j.chemosphere.2005.10.002
Osma JF, Toca-Herrera JL, Rodriguez-Couto S (2010) Transformation pathway of Remazol brilliant blue R by immobilised laccase. Bioresour Technol 101:8509–8514. doi:10.1016/j.biortech.2010.06.074
Pathak H, Soni D, Chauhan K (2014) Evaluation of in vitro efficacy for decolorization and degradation of commercial azo dye RB-B by Morganella sp. HK-1 isolated from dye contaminated industrial landfill. Chemosphere 105:126–132. doi:10.1016/j.chemosphere.2014.01.004
Pavan B, Biondi C, Ferretti ME, Lunghi L, Paganetto G (2001) Phthalic acid mimics 17b-estradiol actions in WISH cells. Toxicol Lett 115:157–164
Prigione V, Tigini V, Pezzella C, Anastasi A, Sannia G, Varese GC (2008) Decolourisation and detoxification of textile effluents by fungal biosorption. Water Res 42:2911–2920. doi:10.1016/j.watres.2008.03.003
Salas-Veizaga DM, Morales-Belpaire I, Terrazas-Siles E (2013) Evaluation of the genotoxic potential of reactive black 5 solutions subjected to decolorizing treatments by three fungal strains. Ecotoxicol Environ Saf 89:125–129. doi:10.1016/j.ecoenv.2012.11.034
Sani RK, Banerjee UC (1999) Decolorization of triphenylmethane dyes and textile and dye-stuff effluent by Kurthia sp. Enzym Microb Technol 24:433–437
Saratale RG, Saratale GD, Chang JS, Govindwar SP (2011) Bacterial decolorization and degradation of azo dyes: a review. J Taiwan Inst Chem E 42:138–157. doi:10.1016/j.jtice.2010.06.006
Si J, Cui B-K, Dai Y-C (2012) Decolorization of chemically different dyes by white-rot fungi in submerged cultures. Ann Microbiol 63:1099–1108. doi:10.1007/s13213-012-0567-8
Silva MC, Torres JA, Vasconcelos de Sá LR, Chagas PMB, Ferreira-Leitão VS, Corrêa AD (2013) The use of soybean peroxidase in the decolourization of Remazol brilliant blue R and toxicological evaluation of its degradation products. J Mol Catal B Enzym 89:122–129. doi:10.1016/j.molcatb.2013.01.004
Soriano JJ, Mathieu-Denoncourt J, Norman G, de Solla SR, Langlois VS (2014) Toxicity of the azo dyes acid red 97 and Bismarck Brown Y to western clawed frog (Silurana tropicalis. Environ Sci Pollut Res Int 21:3582–3591. doi:10.1007/s11356-013-2323-4
Tony BD, Goyal D, Khanna S (2009) Decolorization of textile azo dyes by aerobic bacterial consortium. Int Biodeterior Biodegrad 63:462–469. doi:10.1016/j.ibiod.2009.01.003
Wang H, Li Q, Lu Y, He N, Hong J, Zou X (2007) Performance of batch-operated combined hydrolytic–aerobic biofilm process in treating anthraquinone reactive dye wastewater. Environ Eng Sci 24:483–492. doi:10.1089/ees.2006.0088
Wang H et al (2009) Removal of anthraquinone reactive dye from wastewater by batch hydrolytic–aerobic recycling process. Sep Purif Technol 67:180–186. doi:10.1016/j.seppur.2009.03.018
Wang X, Wen X, Yan H, Ding K, Zhao F, Hu M (2011a) Bacterial community dynamics in a functionally stable pilot-scale wastewater treatment plant. Bioresour Technol 102:2352–2357. doi:10.1016/j.biortech.2010.10.095
Wang Y, Zhu K, Zheng Y, Wang H, Dong G, He N, Li Q (2011b) The effect of recycling flux on the performance and microbial community composition of a biofilm hydrolytic-aerobic recycling process treating anthraquinone reactive dyes. Molecules 16:9838–9849. doi:10.3390/molecules16129838
Yang Q et al (2012) Evolution of the microbial community in a full-scale printing and dyeing wastewater treatment system. Bioresour Technol 117:155–163. doi:10.1016/j.biortech.2012.04.059
Acknowledgments
The authors acknowledge the financial support by the National Natural Science Foundation of China (21377023, 51508083), Shanghai Natural Science Foundation (13ZR1401000), the Fundamental Research Funds for the Central Universities (2232015D3-22), and Chinese Universities Scientific Fund (CUSF-DH-D-2015040). This work was partially supported by Shanghai Leading Academic Discipline Project (B604).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Responsible editor: Gerald Thouand
Rights and permissions
About this article
Cite this article
Liu, N., Xie, X., Yang, B. et al. Performance and microbial community structures of hydrolysis acidification process treating azo and anthraquinone dyes in different stages. Environ Sci Pollut Res 24, 252–263 (2017). https://doi.org/10.1007/s11356-016-7705-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-016-7705-y