Abstract
Background and aims
Exclusive enteral nutrition is recommended as a first-line treatment in active pediatric Crohn’s Disease, but its mechanism of action is still not clear. We aimed to assess alterations in the metabolic profile of newly diagnosed pediatric Crohn’s Disease patients before and during exclusive enteral nutrition therapy.
Methods
Plasma samples from 14 pediatric Crohn’s Disease patients before and after 3–4 weeks on exclusive enteral nutrition were analyzed using mass spectrometry. T-test, fold change and orthogonal partial least squares discriminant analysis were used for mining significant features. Correlation analysis was performed between the annotated features and the weighted pediatric Crohn’s disease activity index using Pearson r distance.
Results
Among the 13 compounds which decreased during exclusive enteral nutrition, most are related to diet, while one is a bacterial metabolite, Bacteriohopane-32,33,34,35-tetrol. The phosphatidic acid metabolite PA(15:1/18:0) was significantly reduced and correlated with the weighted pediatric Crohn’s disease activity index. Lipids increased during exclusive enteral nutrition therapy included phosphatidylethanolamines; PE(24:1/24:1), PE(17:2/20:2) and one lactosylceramide; LacCer(d18:1/14:0).
Conclusion
Food additives and other phytochemicals were the major metabolites, which decreased following the exclusion of a regular diet during exclusive enteral nutrition. An alteration in bacterial biomarkers may reflect changes in intestinal microbiota composition and metabolism. Thus, metabolomics provides an opportunity to characterize the molecular mechanisms of dietary factors triggering Crohn’s Disease activity, and the mechanisms of action of exclusive enteral nutrition, thereby providing the basis for the development and evaluation of improved intervention strategies for prevention and treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Genetic and environmental factors, including the gut microbiota, play an important role in the pathophysiology of Crohn’s disease (CD) [1]. CD has been attributed to complex mechanisms triggering an dysregulated immune response to commensal gut microbiota [2]. Exclusive enteral nutrition (EEN) is an established therapy to induce remission in pediatric CD patients with efficacy in up to 80% of individuals [3], and it is recommended as first-line therapy in European guidelines [4,5,6]. A meta-analysis based on 18 studies concluded that there is no significant difference in efficacy using EEN or corticosteroids treatment to induce remission in pediatric CD, however, EEN seems to be superior in promoting mucosal healing and faster reduction of PCDAI (pediatric Chrohn’s disease activity index) [7]. The benefits of using EEN in pediatric patients extend beyond promoting mucosal healing, contributing to the improvement of nutritional status [8], bone metabolism, and muscle mass [9]. However, the mechanism of EEN action is still unclear. Several studies have focused on investigating how EEN may affect the microbiome, and most report an overall decrease in microbiome diversity during EEN therapy [10,11,12,13,14,15,16,17]. However, the use of different techniques to assess taxonomic shifts and the high diversity of microorganisms present in the human microbiota and its interindividual variation generates heterogenous results, especially at taxonomic resolution lower than phylum-level [18]. In addition to changes in microbiota composition, CD-associated dysbiosis affects microbial metabolic functions. Previous research demonstrated alterations in microbial functions with a shift in genetic abundance related to oxidative stress pathways, carbohydrate metabolism and amino acid biosynthesis, which was considered more disturbed than microbiota composition shifts [19]. In another observational study, the authors reported a reduction in metabolic activity of the intestinal microbiome during enteral feeding for two weeks, based on analyzing exhaled breath and fecal samples using gas chromatography/mass spectrometry [20]. Recently, Diederen et al. reported a reduction in microbiome diversity and changes in the fecal metabolome during EEN in pediatric CD patients, with alterations in amino acids, cadaverine, trimethylamine, and bile acids levels [21].
Over the past years, untargeted metabolomics using mass spectrometry has been applied as powerful tool for identification and tracking of biomarkers which help in understanding the system biology and treatment outcomes [22]. Metabolomics have provided new insights into metabolic alterations in CD patients versus healthy subjects using both serum and fecal samples [23,24]. However, only few studies were performed on pediatric population [25] and could not detect the differences induced by EEN therapy, which is one of the points of strength of our study.
2 Methods
2.1 Plasma samples
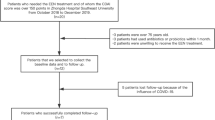
Plasma samples from 14 pediatric CD patients (age (mean ± SD) 13.5 ± 2.2 years, 8 boys, 12 newly diagnosed), before and after 25 ± 5 days on EEN treatment were analyzed. Thirteen patients received Modulen® IBD (Nestlé Nutrition) and one patient was treated with Neocate Junior® (Nutricia). Samples were obtained from a previously published study [15].
The sample preparation procedure was previously described 26. Briefly, after samples were thawed on ice, 100 µL of plasma were precipitated with the addition of 900 µL of ice-cold high-performance liquid chromatography (HPLC)-grade methanol in a 1.5 mL Eppendorf tube, vortexed and rested at -20 °C for 30 min. The tubes were then centrifuged, and the supernatant was filtered in a polytetrafluoroethylene (PTFE) 45 μm 96-well filter plate. Samples were kept at − 80 °C before analysis. Quality control (QC) samples were created by pooling aliquots from the study samples and used to create an inclusion list in the method development and to ensure reproducibility in the analysis.
2.2 LC-QTOF-MS(/MS)
The analysis was conducted on a 1290 Infinity II HPLC system using a Poroshell 120 EC-C18 column (2.1 × 150 mm, 2.7 μm) coupled to a 6545 Q-TOF (both from Agilent, Santa Clara, CA), as previously described [26]. Briefly, 6 µL of the quality control sample were injected in triplicate in full scan (MS) acquisition mode. Data from the MS experiment was then used to create an acquisition list to be used in the auto MS/MS acquisition mode. The analysis was performed with the instrument in the 2 GHz, extended dynamic range in the negative ionization (NEG) mode using an Agilent Jet Stream (AJS) electrospray ionization (ESI) ion source. Operation parameters were set as follows: capillary voltage: − 4000 V; nozzle voltage − 500 V; nebulizer pressure: 40 psi; gas temperature: 290 °C; sheath gas flow: 12 L/min; sheath gas temperature: 380 °C; fragmentor voltage: 170 V; and skimmer voltage: 65 V. The instrument was operated using MassHunter Acquisition B.09.00 software (Agilent). Chromatographic separation was achieved in 16 min run time using a mobile phase A, water (0.1% formic acid); and B, methanol (0.1% formic acid) at a flow rate of 0.4 mL/min. Gradient elution was performed with an initial mixture of 5% B and 95% A, then increased to 60% B throughout 4 min, to 99% B at 12 min, held until 14 min, returned to 5% B at 15.1 min, and held to 16 min.
2.2.1 LC-QTOF-MS analysis
The full-scan analysis was performed in triplicate using the QC samples acquired over the range of 100 to 1050 m/z in the NEG mode to create an inclusion list to further create the auto MS/MS acquisition mode. Data were extracted using batch recursive feature extraction algorithm in MassHunter Profinder B.08.00 software (Agilent) and after evaluation exported as CEF (Cluster Exchange Format) files. The features were aligned on Mass Profiler software (Agilent) using retention time (RT) tolerance of up to 0.3 min and mass tolerance of ± 15 ppm. Features with a 100% occurrence in the replicates were used to create a target MS/MS inclusion list.
2.2.2 LC-QTOF-MS/MS analysis
Analysis of the study samples was performed using data-dependent acquisition (DDA) (auto MS/MS) acquisition mode using the inclusion list as preferred ions for fragmentation, using delta m/z of 15 ppm and delta retention time (RT) of 0.15 min. The collision cell operates with fixed collision energies of 10, 20, and 40 eV using nitrogen (N2) as the collision gas. The acquisition parameters were set as follows: acquisition mass range: 100 to 1050 m/z at 4 spectra/s in the MS, and 50–800 m/z at 3 spectra/s in the time of flight (TOF).
2.3 Data processing and statistical analysis
Samples from 14 pediatric CD patients, from before and after at least 19 days of EEN were analyzed pairwise. Data were processed as described in our previous work [26]. Briefly, quantile normalization was applied on raw data for features filtered based on QC procedure, where features with less than 100% occurrence between the QC and with coefficient of variance higher than 25% were excluded. Principal component analysis was used to perform a QC on samples to exclude any outlier by visual inspection. Identification of features was performed by library search using Mass Hunter METLIN Personal Compound Database and Library (PCDL) (Agilent Technologies, Santa Clara, USA) at MS/MS level.
Then, Principal component analysis (PCA) was used to evaluate reproducibility across measurements by checking the location of the QC samples on the PCA plots. After quality control, annotated compounds were used in the statistical analysis using MetaboAnalyst 4.0 [27]. Fold change and t-test, using false discovery rate (FDR) to correct for multiple testing, were performed to detect significant changes in certain metabolites between the pairwise samples over time. Orthogonal partial least squares discriminant analysis (OPLS-DA) model was built, and significant metabolites related to the differences between the pairwise samples were identified using the S-plot, comprising the combination of magnitude (covariance), with the effect and reliability (correlation) for the model variables concerning model component scores. Correlation analysis was performed using the difference between the values of the annotated features and the weighted pediatric Crohn’s disease activity index wPCDAI [28] scores (after EEN - before EEN) using Pearson r distance measurement.
3 Results
An average of 8000 features were detected per sample from the total of samples analyzed, 6000–7000 features had formulas generated and 3000–4000 were putatively annotated. However, after a strict QC, 318 features were filtered with match score higher than 70 and are shown in the supplementary material containing their respective retention time and m/z (S1). PCA performed on those data showed clustering of all QCs together on obtained PCA models (Data not shown) which indicates the system’s stability and consistent performance throughout the analysis.
A volcano plot (Fig. 1) was built presenting the compounds filtered by pvalues and fold change (FC) and shows eight compounds found significant on the t-test with a p-value < 0.05 (Table 1). Albeit, after FDR correction, only 4 features remained significant at p < 0.05. Considering FC, with arbitrary cut-off of 1.3, two compounds (Bacteriohopane-32,33,34,35-tetrol, and PA(15:1/18:0)) were filtered as significant. Orthogonal PLS-DA model provided complete separation between the samples before and after EEN treatment. Figure 2 depicts the scores plot and the s-plot with features of importance showing the most significant compounds ordered by the covariance loading values obtained using an arbitrary cut-off value of 3.6, resulting in 10 annotated compounds with increased concentration after treatment and 13 compounds which decreased in concentration (Table 2).
Volcano plot with log2 fold change (FC) in the x-axis and –log10 of p values on the y axis. The lines indicate FC > 1.3. Box whisker plot showing the interquartile range of the significant metabolites before (red) and after EEN treatment (green)
* Entities significant after FDR correction for multiple testing
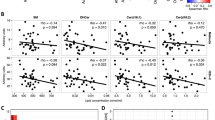
As previously reported [15], disease activity measured by wPCDAI decreased significantly (p < 0.001) over the course of EEN treatment. The average score (± SD) was 48.8 ± 18.6 before start of EEN and decreased to 16.4 ± 10.1 after 3–4 weeks on EEN therapy. wPCDAI was developed as a score to stratify the severity of Crohn’s disease in pediatric patients with ranges between 0 to 100 with higher scores signifying more active disease. A score of < 12,5 is consistent remission, > 40 indicates moderate disease, and > 57,5 severe disease. A 17.5-point decrease in PCDAI is taken as evidence of small improvement and 37,5 as moderate improvement [28]. Correlation analysis was performed between the annotated compounds and the wPCDAI scores. The most significant compounds with R values < -0.5 for the negative correlation and > 0.3 for the positive are shown in Fig. 3. The following compounds were found negatively correlated with wPCDAI scores: 1-Phosphatidyl-1D-myo-inositol 3-phosphate; Erythromycin ethylsuccinate; 1,2,4,5,7-Pentathiocane; dTDP / Thymidine 5’-diphosphate; PS(22:4/19:0); Medicagenic acid beta-maltoside; Phe Arg Val; Anisole; and PS(18:2/21:0) with R values of -0.71, -0.67, -0.66, -0.64, -0.58, -0.55, -0.53, -0.52, -0.52, respectively). Conversely, we filtered 9 compounds positively correlated with wPCDAI: CDP-DG(16:0/20:4), 1,2-bis(Chloromethoxy)ethane; Theophylline/Theobromine; LMST03020510; Navalioside; 2α-Fluoro-19-nor-22-oxa-1α,25-dihydroxyvitamin D3; PA(15:1/18:0); 3-Acetylthiophene; and LMST01080090 ( R = 0.30, 0.31, 0.35, 0.35, 0.35, 0.43, 0.50, 0.54, and 0.67, respectively).
4 Discussion
The use of EEN therapy clearly shows an impact on the plasma metabolome, with complete separation by orthogonal PLS-DA. Some features with a higher importance in the model appear related to the exclusion of a regular diet and they decreased most during EEN treatment. The compound annotated as Theophylline, for instance, presented the same match score as Theobromine, natural methylxanthines present in chocolate derivatives and beverages that are highly consumed by children and teenagers [29]. Other compounds related to diet that decreased during EEN therapy included: the food additive (+/-)-3-[(2-Methyl-3-furyl)thio]-2-Butanone [30]; organic compounds found in fruits: 3-(3,4-Dihydroxyphenyl)-1-Propanol 3’-Glucoside [31] and 2-Phenylethyl beta-D-glucopyranoside [32]; in tomatoes: N-Nitrosotomatidine [33] and other phytochemicals naturally occurring in fats and oils, green vegetables, herbs, and spices: (Methoxysulfinyl) Pentasulfide [34] and 4-(3-Hydroxy-7-phenyl-6-heptenyl)-1,2-benzenediol [35]. Violacene is a polyhalogenated monocyclic monoterpene [36] which is produced by diverse genera of bacterial strains which were isolated from various marine to freshwater and soil environments as well as marine algae and could probably reflect cessation of food intake [37]. A slight reduction shift was detected in Prostaglandin D2-1-glyceryl ester, a bioactive lipid involved in the endocannabinoid system with potential anti-inflammatory properties in vivo [38].
The compounds found elevated during EEN treatment in the multivariate analysis present two features annotated as phosphatidylethanolamine (PE), PE(24:1/24:1), and PE(17:2/20:2) and one ceramide phosphoethanolamine (PE-Cer): PE-Cer(d16:2/23:0). PE are estimated to comprise 15–25% of the total lipid content in mammalian cells and exert remarkable bio-activities [39]. In Gram-negative bacteria, PE comprises around 75% of the phospholipid cell envelop and are dynamic key compounds modulating metabolic activities [40].
Other elevated compounds were 3,5-Diiodo-L-tyrosine, involved in thyroid hormone synthesis [41] and LacCer(d18:1/14:0). Lactosylceramide (LacCer) was previously related with CD, however the role in the pathogenesis was unclear [42]. Another study indicated the potential of Lactosylceramide as a potential biomarker of inflammatory bowel disease in children [43]. LacCer is highly expressed in phagocytes and epithelial cells and may play an essential role in the human innate immune system, binding pathogenic microorganisms [44]. Furfuryl thioacetate, a naturally occurring aroma compound [45], and Furfuryl B, a secondary metabolite with antimicrobial activity produced by many plants source of edible vegetable oils [46] were also elevated. Two chemical entities appear with weighing importance in the OPLS-DA model, 2-Methylpropanoyl phosphate and 1,3-Benzenedisulfonamide; however, neither of them has been previously reported in the human metabolome. Interestingly, compounds containing the sulphonamide moiety present potential biological activities, such as carbonic anhydrase and COX-1/2 inhibition, as well as anti-inflammatory, and antitumor activities [47].
Some of the features with high importance in the multivariate model were found significant in the t-test and fold change, concomitantly. Bacteriohopane-32,33,34,35-tetrol, a bacterial metabolite, and biomarker of Burkholderia, Pseudomonas, and Ralstonia spp [48[, decreased during treatment. Burkholderia spp is a Proteobacteria known for causing dysfunction of GALT and gut microbiota in IBD [49], with the potential to invade intestinal epithelial cells [50]. An increase in the abundance of proteobacteria has been reported in IBD patients [51], in active or aggressive Crohn’s disease [52]. A decrease of Bacteriohopane-32,33,34,35-tetrol concentration after EEN treatment may indicate an effect on the gut microbiome with a decrease in Burkholderia and/or Pseudomonas population. Thus, the presence of elevated PE in plasma could be a marker of gram-negative bacterial cell membranes that underwent cell death.
Two other features annotated as Cohibin B and Chatenaytrienin 2 were significantly (p < 0.05) decreased with EEN treatment. Both compounds belong to the class of organic compounds known as annonaceous acetogenins, a class of natural compounds with a wide variety of biological activities and are powerful inhibitors of complex I (NADH : ubiquinone oxidoreductase) in mammalian and insect mitochondrial electron transport systems [53].
The most important metabolite in the OPLS-DA model was PA(15:1/18:0), also found significantly decreased in the t-test and fold change. PA is the simplest class of glycerophospholipids (GPL) present in virtually all organisms, from bacteria to higher plants. It is an intermediate in lipid membrane synthesis and storage and is also involved in many eukaryotic processes. Besides, PA influences membrane structure and interacts with diverse proteins due to its unique physicochemical properties in comparison to other GPL, thus acting as a lipid mediator in various signaling and cellular processes 54.
We correlated the metabolome with the patients’ wPCDAI scores to investigate what metabolites were associated with clinical improvement. The results show two metabolites in common with the list of features related to EEN treatment. There was a parallel decrease of PA(15:1/18:0), which correlated with wPCDAI scores, indicating that this metabolite could play a role in the CD pathophysiology. Another compound showing a similar trend was Theophylline/Theobromine reflecting exclusion of chocolate derivatives and beverages. These products were previously shown to modify the intestinal microbiome in a rat model, with a decrease in E. coli; Bifidobacterium spp.; Streptococcus spp.; and Clostridium histolyticum-C [55].
4.1 Strengths and limitations
The strengths of our study are the use of a novel sensitive analytical methodology with the capability of detection of a wide variety of endogenous and exogenous compounds and inclusion of mostly treatment naïve patients. The limitations are the putative annotation of compounds, the small sample size, a limited number of two samples per patient, and the absence of dietary data at baseline, and the associations with hypothesis raising nature without providing evidence on causality.
5 Conclusion
We report remarkable alterations in the plasma metabolic profile of pediatric CD patients treated with EEN. Markers reflecting the change of a mixed regular to a formula diet showed marked changes, including xenobiotics, such as food additives, other phytochemicals, and Theophylline/Theobromine which also correlated with a CD activity index. The decrease in the concentration of a biomarker of proteobacteria (Bacteriohopane-32,33,34,35-tetrol) and the increase in (PE) concentration, the main compound in cell membrane composition of gram-negative bacteria, together may indicate an alteration in potential pathogenic bacteria populations and metabolism. Thus, metabolomics provides an opportunity to characterize the molecular mechanisms of dietary factors triggering CD activity and unravel the mechanisms of action of EEN.
Data Availability
The data underlying this article will be shared at reasonable request to the corresponding author.
References
Sasson, A. N., Ananthakrishnan, A. N., & Raman, M. (2019). Diet in treatment of inflammatory bowel diseases.Clin Gastroenterol Hepatol
Fiocchi, C. (2015). Inflammatory bowel disease pathogenesis: Where are we? Journal Of Gastroenterology And Hepatology, 30(Suppl 1), 12–18.
Ashton, J. J., Gavin, J., & Beattie, R. M. (2019). Exclusive enteral nutrition in crohn’s disease: Evidence and practicalities. Clinical Nutrition, 38, 80–89.
Wall, C. L., Day, A. S., & Gearry, R. B. (2013). Use of exclusive enteral nutrition in adults with crohn’s disease: A review. World Journal Of Gastroenterology, 19, 7652–7660.
Frivolt, K., Schwerd, T., Werkstetter, K. J., et al. (2014). Repeated exclusive enteral nutrition in the treatment of paediatric crohn’s disease: Predictors of efficacy and outcome. Alimentary Pharmacology & Therapeutics, 39, 1398–1407.
Ruemmele, F. M., Veres, G., Kolho, K. L., et al. (2014). Consensus guidelines of ecco/espghan on the medical management of pediatric crohn’s disease. J Crohns Colitis, 8, 1179–1207.
Yu, Y., Chen, K. C., & Chen, J. (2019). Exclusive enteral nutrition versus corticosteroids for treatment of pediatric crohn’s disease: A meta-analysis. World Journal of Pediatrics, 15, 26–36.
Day, A. S., Whitten, K. E., Lemberg, D. A., et al. (2006). Exclusive enteral feeding as primary therapy for crohn’s disease in australian children and adolescents: A feasible and effective approach. Journal Of Gastroenterology And Hepatology, 21, 1609–1614.
Werkstetter, K. J., Schatz, S. B., Alberer, M., Filipiak-Pittroff, B., & Koletzko, S. (2013). Influence of exclusive enteral nutrition therapy on bone density and geometry in newly diagnosed pediatric crohn’s disease patients. Annals Of Nutrition & Metabolism, 63, 10–16.
Leach, S. T., Mitchell, H. M., Eng, W. R., Zhang, L., & Day, A. S. (2008). Sustained modulation of intestinal bacteria by exclusive enteral nutrition used to treat children with crohn’s disease. Alimentary Pharmacology & Therapeutics, 28, 724–733.
Shiga, H., Kajiura, T., Shinozaki, J., et al. (2012). Changes of faecal microbiota in patients with crohn’s disease treated with an elemental diet and total parenteral nutrition. Digestive And Liver Disease : Official Journal Of The Italian Society Of Gastroenterology And The Italian Association For The Study Of The Liver, 44, 736–742.
Lionetti, P., Callegari, M. L., Ferrari, S., et al. (2005). Enteral nutrition and microflora in pediatric crohn’s disease. Jpen. Journal Of Parenteral And Enteral Nutrition, 29, S173–S175. discussion S5-8.
Gerasimidis, K., Bertz, M., Hanske, L., et al. (2014). Decline in presumptively protective gut bacterial species and metabolites are paradoxically associated with disease improvement in pediatric crohn’s disease during enteral nutrition. Inflammatory Bowel Diseases, 20, 861–871.
Kaakoush, N. O., Day, A. S., Leach, S. T., et al. (2015). Effect of exclusive enteral nutrition on the microbiota of children with newly diagnosed crohn’s disease. Clin Transl Gastroenterol, 6, e71.
Schwerd, T., Frivolt, K., Clavel, T., et al. (2016). Exclusive enteral nutrition in active pediatric crohn disease: Effects on intestinal microbiota and immune regulation. The Journal Of Allergy And Clinical Immunology, 138, 592–596.
Quince, C., Ijaz, U. Z., Loman, N., et al. (2015). Extensive modulation of the fecal metagenome in children with crohn’s disease during exclusive enteral nutrition. American Journal Of Gastroenterology, 110, 1718–1729. quiz 30.
Guinet-Charpentier, C., Lepage, P., Morali, A., Chamaillard, M., & Peyrin-Biroulet, L. (2017). Effects of enteral polymeric diet on gut microbiota in children with crohn’s disease. Gut, 66, 194–195.
MacLellan, A. C., Grant, J., Cahill, S., Langille, L., & Van Limbergen, M. G. I. (2017). The impact of exclusive enteral nutrition (een) on the gut microbiome in crohn’s disease: A review. Nutrients, 9, 447.
Morgan, X. C., Tickle, T. L., Sokol, H., et al. (2012). Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biology, 13, R79.
Walton, C., Montoya, M. P., Fowler, D. P., et al. (2016). Enteral feeding reduces metabolic activity of the intestinal microbiome in crohn’s disease: An observational study. European Journal Of Clinical Nutrition, 70, 1052–1056.
Diederen, K., Li, J. V., Donachie, G. E., et al. (2020). Exclusive enteral nutrition mediates gut microbial and metabolic changes that are associated with remission in children with crohn’s disease. Scientific reports, 10, 18879.
Johnson, C. H., Ivanisevic, J., Siuzdak, G., & Metabolomics (2016). Beyond biomarkers and towards mechanisms. Nature Reviews Molecular Cell Biology, 17, 451–459.
Kolho, K. L., Pessia, A., Jaakkola, T., de Vos, W. M., & Velagapudi, V. (2016). Faecal and serum metabolomics in paediatric inflammatory bowel disease. Journal of Crohn’s and Colitis, 11, 321–334.
Liu, H., Xu, M., He, Q., et al. (2022). Untargeted serum metabolomics reveals specific metabolite abnormalities in patients with crohn’s disease. Front Med (Lausanne), 9, 814839.
Daniluk, U., Daniluk, J., Kucharski, R., et al. (2019). Untargeted metabolomics and inflammatory markers profiling in children with crohn’s disease and ulcerative colitis-a preliminary study. Inflammatory Bowel Diseases, 25, 1120–1128.
Marques, J. G., Shokry, E., Frivolt, K., et al. Metabolomic signatures in pediatric crohn’s disease patients with mild or quiescent disease treated with partial enteral nutrition: A feasibility study.SLAS TECHNOLOGY: Translating Life Sciences Innovation;0:2472630320969147.
Jasmine Chong, D. S. W., & Jianguo Xia. (2019). Using metaboanalyst 4.0 for comprehensive and integrative metabolomics data analysis. Current protocols in bioinformatics, 68, 1–128.
Turner, D., Griffiths, A. M., Walters, T. D., et al. (2011). Mathematical weighting of the pediatric crohn’s disease activity index (pcdai) and comparison with its other short versions. Inflammatory Bowel Diseases, 18, 55–62.
Shively, C. A., & Tarka, S. M. (1984). Jr. Methylxanthine composition and consumption patterns of cocoa and chocolate products. Progress In Clinical And Biological Research, 158, 149–178.
Wishart, D. S., Tzur, D., Knox, C., et al. (2007). Hmdb: The human metabolome database. Showing metabocard for (+/-)-3-[(2-methyl-3-furyl)thio]-2-butanone (hmdb0032401. Nucleic Acids Research, 35, D521–D526.
Wishart, D. S., Tzur, D., Knox, C., et al. (2007). Hmdb: The human metabolome database. Showing metabocard for 3-(3,4-dihydroxyphenyl)-1-propanol 3’-glucoside (hmdb0038304). Nucleic Acids Research, 35, D521–D526.
Wishart, D. S., Tzur, D., Knox, C., et al. (2007). Hmdb: The human metabolome database. Nucleic Acids Research, 35, D521–D526.
Wishart, D. S., Tzur, D., Knox, C., et al. (2007). Hmdb: The human metabolome database. Showing metabocard for n-nitrosotomatidine (hmdb0034102). Nucleic Acids Research, 35, D521–D526.
Wishart, D. S., Tzur, D., Knox, C., et al. (2007). Hmdb: The human metabolome database. Showing metabocard for amino (methoxysulfinyl) pentasulfide (hmdb0031980). Nucleic Acids Research, 35, D521–D526.
Wishart, D. S., Tzur, D., Knox, C., et al. (2007). Hmdb: The human metabolome database. Showing metabocard for 4-(3-hydroxy-7-phenyl-6-heptenyl)-1,2-benzenediol (hmdb0041089). Nucleic Acids Research, 35, D521–D526.
Van Engen, D., Clardy, J., Kho-Wiseman, E., et al. (1978). Violacene: A reassignment of structure. Tetrahedron Letters ;19:29–32.
Choi, S. Y., Yoon, K. H., Lee, J. I., & Mitchell, R. J. (2015). Violacein: Properties and production of a versatile bacterial pigment. Biomed Research International, 2015, 465056.
Alhouayek, M., Masquelier, J., Cani, P. D., Lambert, D. M., & Muccioli, G. G. (2013). Implication of the anti-inflammatory bioactive lipid prostaglandin d2-glycerol ester in the control of macrophage activation and inflammation by abhd6. Proceedings of the National Academy of Sciences of the United States of America, 110, 17558–17563.
Patel, D., & Witt, S. N. (2017). Ethanolamine and phosphatidylethanolamine: Partners in health and disease. Oxidative Medicine and Cellular Longevity, 2017, 4829180.
Bogdanov, M., Pyrshev, K., Yesylevskyy, S., et al. (2020). Phospholipid distribution in the cytoplasmic membrane of gram-negative bacteria is highly asymmetric, dynamic, and cell shape-dependent. Science Advances, 6, eaaz6333.
Wishart, D. S., Tzur, D., Knox, C., et al. (2007). Hmdb: The human metabolome database. Showing metabocard for 3,5-diiodo-l-tyrosine (hmdb0003474). Nucleic Acids Research, 35, D521–D526.
Stevens, C. R., Oberholzer, V. G., Walker-Smith, J. A., & Phillips, A. D. (1988). Lactosylceramide in inflammatory bowel disease: A biochemical study. Gut, 29, 580–587.
Filimoniuk, A., Blachnio-Zabielska, A., Imierska, M., Lebensztejn, D. M., & Daniluk, U. (2020). Sphingolipid analysis indicate lactosylceramide as a potential biomarker of inflammatory bowel disease in children.Biomolecules;10.
Iwabuchi, K. (2018). [lactosylceramide-enriched lipid raft-mediated infection immunity]. Med Mycol J, 59, J51–j61.
Láng, J., Rákász, V., Magyar, A., Pállinger, É., & Kőhidai, L. (2011). Chemotactic effect of odorants and tastants on the ciliate tetrahymena pyriformis. Journal of Receptors and Signal Transduction, 31, 423–433.
Pedras, M. S. C., Zheng, Q., & Sarma-Mamillapalle, V. K. (2007). The phytoalexins from brassicaceae: Structure, biological activity, synthesis and biosynthesis. Natural Product Communications, 2, 1934578X0700200315.
Abdel-Aziz, A. A. M., El-Azab, A. S., AlSaif, N. A., et al. (2020). Synthesis, anti-inflammatory, cytotoxic, and cox-1/2 inhibitory activities of cyclic imides bearing 3-benzenesulfonamide, oxime, and β-phenylalanine scaffolds: A molecular docking study. Journal of Enzyme Inhibition and Medicinal Chemistry, 35, 610–621.
Cvejic, J. H., Putra, S. R., El-Beltagy, A., et al. (2000). Bacterial triterpenoids of the hopane series as biomarkers for the chemotaxonomy of burkholderia, pseudomonas and ralstonia spp. Fems Microbiology Letters, 183, 295–299.
Dinakaran, V., Mandape, S. N., Shuba, K., et al. (2019). Identification of specific oral and gut pathogens in full thickness colon of colitis patients: Implications for colon motility. Frontiers In Microbiology, 9, 3220.
Armstrong, H., Alipour, M., Valcheva, R., et al. (2019). Host immunoglobulin g selectively identifies pathobionts in pediatric inflammatory bowel diseases. Microbiome, 7, 1.
Frank, D. N., St Amand, A. L., Feldman, R. A., et al. (2007). Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proceedings of the National Academy of Sciences of the United States of America, 104, 13780–13785.
Vester-Andersen, M. K., Mirsepasi-Lauridsen, H. C., Prosberg, M. V., et al. (2019). Increased abundance of proteobacteria in aggressive crohn’s disease seven years after diagnosis. Scientific reports, 9, 13473.
Alali, F. Q., Liu, X. X., & McLaughlin, J. L. (1999). Annonaceous acetogenins: Recent progress. Journal of Natural Products, 62, 504–540.
Kooijman, E. E., & Burger, K. N. J. (2009). Biophysics and function of phosphatidic acid: A molecular perspective. Biochimica et Biophysica Acta (BBA) - Molecular and Cell Biology of Lipids, 1791, 881–888.
Martín-Peláez, S., Camps-Bossacoma, M., Massot-Cladera, M., et al. (2017). Effect of cocoa’s theobromine on intestinal microbiota of rats.Molecular nutrition & food research;61.
Acknowledgements
We thank all IBD patients and families for their study participation. We thank Klara Frivolt for her wonderful work and enthusiastic support of the clinical trial.
Funding
This work was supported by Agilent Technologies, Waldbronn, Germany, granting an internship in their facilities and providing chemicals and instruments for the analytical measurements. The clinical study was supported by a young investigator grant of the Ludwig-Maximilian-University Munich (FöFoLe 742 to T.S.). TS acknowledges support from the Munich Clinician Scientist Program (MCSP) at LMU. BK is the Else Kröner Seniorprofessor of Paediatrics at LMU – University of Munich, financially supported by Else Kröner Fresenius Foundation, LMU Medical Faculty, and LMU University Hospitals.
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
TS, PB, SK, and BK coordinated the project and critically revised the manuscript; TS and PB recruited patients and collected clinical data; JM acquired mass spectrometry data, performed the statistical analysis, data interpretation, and drafted the manuscript; All co-authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
JGM has nothing to declare. SK reports personal fees from Nestle, Danone, Biocodex, Shire, Abbvie, Pharmacosmos, Janssen, Pfizer, Takeda, Mead-Johnson, and research grants paid to the institution from Mead Johnson outside the submitted work. BK benefitted from support for scientific or educational activities from Barilla, Bayer, Cheplapharm, Cogitando, Danone, DSM, DGC, Hipp, Nestlé, and Reckitt Benkiser. TS received speaker’s fees from MSD and Nutricia. PB received personal fees (lectures, advisory board) from AbbVie, Orphalan, Mirum, Amgen, Nestlé Nutrition Institute, Nutricia, and Albireo with no activities connected to the presented work.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Marques, J.G., Schwerd, T., Bufler, P. et al. Metabolic changes during exclusive enteral nutrition in pediatric Crohn’s disease patients. Metabolomics 18, 96 (2022). https://doi.org/10.1007/s11306-022-01953-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-022-01953-0