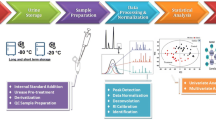
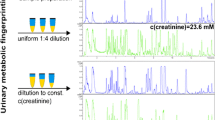
Abstract
Introduction
Metabolomics is a multi-discipline approach to systems biology that provides a snapshot of the metabolic status of a cell, tissue, or organism. Metabolomics uses mass spectroscopy (MS) and nuclear magnetic resonance (NMR) to analyze biological samples for low molecular weight metabolites.
Objective
Normalize urine sample pre-acquisition to perform a targeted quantitative analysis of selected metabolites in rat urine.
Methods
Urine samples were provided from rats on a control diet (n = 10) and moderate sucrose diet (n = 8) collected in a metabolic cage during an eight hour fast. Urine from each sample was prepared by two different methods. One sample was a non-normalized sample of 1200 µL and the second sample was a variable volume-normalized to the concentration of urobilin in a standard sample of urine. The urobilin concentration in all samples was determined by fluorescence. Ten metabolites for each non-normalized and normalized urine sample were quantified by integration to an internal standard of DSS.
Results
Both groups showed an improvement in pH range going from non-normalized to normalized samples. In the group on the control diet, eight metabolites had significant improvement in range, while the remaining two metabolites had insignificant improvement in range comparing the non-normalized sample to the normalized sample. In the group on the moderate sucrose diet all ten metabolites showed significant improvement in range going from non-normalized to normalized samples.
Conclusions
These findings describe a pre-acquisition method of urine normalization to adjust for differences in hydration state of each organism. This results in a narrower concentration range in a targeted analysis.




Similar content being viewed by others
6 References
Amiel, Aurelien, Tremblay-Franco, Marie, Gautier, Roselyne, Ducheix, Simon, Montagner, Alexandra, Polizzi, Arnaud, Debrauwer, Laurent, Guillou, Herve, Bertrand-Michel, Justine, & Canlet, Cecile. (2020). Proton NMR enables the absolute quantification of aqueous metabolites and lipid classes in unique mouse liver samples. Metabolites, 10(1), 9. https://doi.org/10.3390/metabo10010009
Chetwynd, A. J., Abdul-Sada, A., Holt, S. G., & Hill, E. M. (2016). Use of a pre-analysis osmolality normalisation method to correct for variable urine concentrations and for improved metabolomic analyses. Journal of Chromatography A, 1431, 103–110. https://doi.org/10.1016/j.chroma.2015.12.056
Edmands, W. M. B., Ferrari, P., & Scalbert, A. (2014). Normalization to specific gravity prior to analysis improves information recovery from high resolution mass spectrometry metabolomic profiles of human urine. Analytical Chemistry, 86, 10925–10931. https://doi.org/10.1021/ac503190m
Ellis, L. A., Yates, B. A., McKenzie, A. L., Munoz, C. X., Casa, D. J., & Armstrong, L. E. (2016). Effects of three oral nutritional supplements on human hydration indices. International Journal of Sport Nutrition and Exercise Metabolism, 26, 356–362. https://doi.org/10.1123/ijsnem.2015-0244
Gagnebin, Y., Tonoli, D., Lescuyer, P., Ponte, B., de Seigneux, S., Martin, P.-Y., Schappler, J., Boccard, J., & Rudaz, S. (2017). Metabolomic analysis of urine samples by UHPLC-QTOF-MS: Impact of normalization stratagies. Analytica Chimica Acta, 955, 27–35. https://doi.org/10.1016/j.aca.2016.12.029
Gholizadeh, N., Pundavela, J., Nagarajan, R., Dona, A., Quadrelli, S., Biswas, T., Greer, P. B., & Ramadan, S. (2020). Nuclear magnetic resonance spectroscopy of human body fluids and in vivo magnetic resonance spectroscopy potential role in the diagnosis and management of prostate cancer. Urological Oncology: Seminars and Original Investigations, 38, 150–173. https://doi.org/10.1016/j.urolonc.2019.10.019
Giraudeau, Patrick, Tea, Illa, Remaud, Gerald S., & Akoka, Serge. (2014). Reference and normalization methods: Essential tools for the intercomparison of NMR spectra. Journal of Pharmaceutical and Biomedical Analysis, 93, 3–16. https://doi.org/10.1016/j.jppa.2013.07.020
Gorling, Benjamine, Brase, Stefan, & Luy, Burkhard. (2016). NMR chemical shift ranges of urine metabolites in various organic solvents. Metabolites, 6, 27. https://doi.org/10.3390/metabo6030027
Kavouras, S. A., Johnson, E. C., Bougatsas, D., Arnaoutis, G., Panagiotakos, D. B., Perrier, E., & Klein, A. (2016). Validation of a urine color scale for assessment of urine osmolality in healthy children. European Journal of Nutrition, 55, 907–915. https://doi.org/10.1007/s00394-015-0905-2
Khamis, Mona M., Holt, Teagan, Awad, Hanan, El-Aneed, Anas, & Adamko, Daryl J. (2018). Comparative analysis of creatinine and osmolality as urine normalization strategies in targeted metabolimics for the differential diagnosis of asthma and COPD. Metabolomics, 14, 115. https://doi.org/10.1007/s11306-018-1418-9
Kohl, Stefanie M., Klein, Matthias S., Hochren, Jochen, Oefner, Peter J., Spang, Rainer, & Gronwald, Wolfram. (2012). State-of-the-art data normalization methods improve NMR-based metabolomic analysis. Metabolomics, 8(S1), S146–S160. https://doi.org/10.1007/s11306-011-0350-z
Li, Ning, Song, Yi Peng, Tang, Huiru, & Wang, Yulan. (2016). Recent developments in sample preparation and data pre-ttreatment in metabonomics research. Archives of Biochemistry and Biophysics, 589, 4–9. https://doi.org/10.1016/j.abb.2015.08.024
Lichardusova, L., Kusnir, J., Valko-Rokytovska, M., & Marekova, M. (2010). The factors influencing direct spectral fluorimetry of some urine metabolites. Prague Medical REport, 111(4), 272–278.
Lu, W., Xiaoyang, Su., Klein, M. S., Lewis, I. A., Fiehn, O., & Rabinowitz, J. D. (2017). Metabolite measurement: Pitfalls to avoid and practices to follow. Annual Review of Biochemistry, 86, 277–304. https://doi.org/10.1146/annurev-biochem-061516-044952
Ma, Y.-R., Zhou, Y., Huang, J., Qin, H.-Y., Wang, P., & Xin-an, Wu. (2018). The urinary excretion of metformin, ceftizoxime and ofloxacin in high serum creatinine rats: Can creatinine predict renal tubular elimination? Life Sciences, 196, 110–117. https://doi.org/10.1016/j.lfs.2018.01.017
McKenzie, James J., Donarski, James A., Wilson, Julie C., & Charlton, Adrian J. (2011). Analysis of complex mixtures using high-resolution nuclear magnetic resonance spectroscopy and chemometrics. Progress in Nuclear Magnetic Resonance Spectroscopy, 59, 336–359. https://doi.org/10.1016/j.pnmrs.2011.04.003
Monakhova, Y. B., & Diehl, B. W. K. (2017). Practical guide for selection of 1H qNMR acquisition and processing parameters confirmed by automated spectra evaluation. Magnetic Resonance in Chemistry, 55, 996–1005. https://doi.org/10.1002/mrc.4622
Nicholson, J. K., Lindon, J. C., & Holmes, E. (1999). ‘Metabonomics’: Understanding the metabolic responses of living system to pathophysiological stimuli via multivariate statistical analysis of biological NMR spectroscopic data. Xenobiotica, 29(11), 1181–1189. https://doi.org/10.1080/004982599238047
Perrier, E., Rondeau, P., Poupin, M., Le Bellego, L., Armstrong, L. E., Lang, F., Stookey, J., Tack, I., Vergne, S., & Klein, A. (2013). Relation between urinary hydration biomarkers and total fluid intake in healthy adults. European Journal of Clinical Nutrition, 67, 939–943. https://doi.org/10.1038/ejcn.2013.93
Puig-Castellvi, F., Perez, Y., Pina, B., Tauler, R., & Alfonso, I. (2018). Comparative analysis of 1H NMR and 1H–13C HSQC NMR metabolomics to understand the effects of medium composition in yeast growth. Analytical Chemistry, 90, 12422–12430. https://doi.org/10.1021/acs.analchem.8b01196
Rosato, A., Tenori, L., Cascante, M., Carulla, P. R. D. A., Vitor, A. P., dos Santos, M., & Saccenti, E. (2018). From correlation to causation: Analysis of metabolomics data using systems biology approaches. Metabolomics, 14, 37. https://doi.org/10.1007/s11306-018-1335-y
Saccenti, E. (2017). Correlation patterns in experimental data are affected by normalization procedures: Consequences for data analysis and network inference. Journal of Proteome Research, 16, 619–634. https://doi.org/10.1021/acs.jproteome.6b00704
Schmidt, N. A., & Scholtis, R. J. H. (1964). Urobilin in Urine. Clinica Chemica Acta, 10(6), 574–576. https://doi.org/10.1016/0009-8981(64)90201-3
Torgrip, R. J. O., Aberg, K. M., Alm, E., Schuppe-Koistinen, I., & Lindberg, J. (2008). A note on normalization of biofluid 1D 1H-NMR data. Metabolomics, 4, 114–121. https://doi.org/10.1007/s11306-007-0102-2
Tredwell, G. T., Bundy, J. G., De Iorio, M., & Ebbels, T. M. D. (2016). Modelling the acid/base 1H NMR chemical shift limits of metabolites in human urine. Metabolomics, 12, 152. https://doi.org/10.1007/s11306-016-1101-y
Vogl, Franziska C., Mehrl, Sebastian, Heizinger, Leonhard, Schlecht, Inga, Zacharias, Helena U., Ellman, Lisa, Nurnberger, Nadine, et al. (2016). Evaluation of dilution and normalization strategies to correct for urinary output in HPLC-HRTOFMS metabolomics. Analytical and Bioanalytical Chemistry, 408, 8483–8493. https://doi.org/10.1007/s00216-016-9974-1
Vu, T., Siemak, P., Bhinderwala, F., Yuhang, Xu., & Powers, R. (2019). Evaluation of multivariate classification models for analyzing NMR metabolomics data. Journal of Proteome Research, 18, 3282–3294. https://doi.org/10.1021/acs.jproteome.9b00227
Waikar, S. S., Sabbisetti, V. S., & Bonventre, J. V. (2010). Normalization of urinary biomarkers to creatinine curing changes in glomerular filtration rate. Kidney International, 78, 486–494. https://doi.org/10.1038/ki.2010.165
Wallmeier, J., Samol, C., Ellman, L., Zacharias, H. U., Vogl, F. C., Garcia, M., Dettmer, K., Oefner, P. J., & Gronwald, W. (2017). Quantification of metabolites by NMR spectroscopy in the presence of protein. Journal of Proteome Research. https://doi.org/10.1021/acs.jproteome.7b00057
Warrack, Bethanne M., Hnatyshyn, Serhiy, Ott, Karl-Heinz., Reily, Michael D., Sanders, Mark, Zhang, Haiying, & Drexler, Dieter M. (2009). Normalization strategies for metabonomic analysis of urine samples. Journal of Chromatography B, 877, 547–552. https://doi.org/10.1016/j.jchromb.2009.01.007
Wu, Y., & Li, L. (2016). Sample Normalization methods in quantitative metabolomics. Journal of Chromatography A, 1430, 80–95.
Zacharias, H. U., Rehberg, T., Mehrl, S., Richtmann, D., Wettig, T., Oefner, P. J., Spang, R., Gronwald, W., & Altenbuchinger, M. (2017). Scale-invariant biomarker discovery in urine and plasma metabolite fingerprints. Journal of Proteome Research Online. https://doi.org/10.1021/acs.jproteome.7b00325
Acknowledgements
The authors would like to thank Dr. John B. Vincent in the Department of Chemistry and Biochemistry at the University of Alabama, Tuscaloosa, for the loan of the metabolic cages used for urine collection. We would also like to thank Genoah Collins, Amelia Clopp, Luis Mercado, and Helen Gibson in Dr. Love-Rutledge’s lab for the rat urine samples used in our experiments.
Funding
This study was financially supported by the Army Education Outreach Program and the Research or Creative Experience for Undergraduates at the University of Alabama in Huntsville.
Author information
Authors and Affiliations
Contributions
JGW wrote the manuscript and prepared figures. SLR, BV contributed to the content of the paper. BV, EH and SB participated in data collection. BV developed NMR method participated in data interpretation, prepared figures. SLR contributed to the design of the study.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wolfsberger, J.G., Hunt, E.C., Bobba, S.S. et al. Metabolite quantification: A fluorescence-based method for urine sample normalization prior to 1H-NMR analysis. Metabolomics 18, 80 (2022). https://doi.org/10.1007/s11306-022-01939-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-022-01939-y