Abstract
Introduction
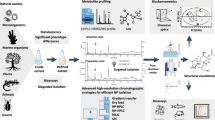
Despite the availability of several pre-processing software, poor peak integration remains a prevalent problem in untargeted metabolomics data generated using liquid chromatography high–resolution mass spectrometry (LC–MS). As a result, the output of these pre-processing software may retain incorrectly calculated metabolite abundances that can perpetuate in downstream analyses.
Objectives
To address this problem, we propose a computational methodology that combines machine learning and peak quality metrics to filter out low quality peaks.
Methods
Specifically, we comprehensively and systematically compared the performance of 24 different classifiers generated by combining eight classification algorithms and three sets of peak quality metrics on the task of distinguishing reliably integrated peaks from poorly integrated ones. These classifiers were compared to using a residual standard deviation (RSD) cut-off in pooled quality-control (QC) samples, which aims to remove peaks with analytical error.
Results
The best performing classifier was found to be a combination of the AdaBoost algorithm and a set of 11 peak quality metrics previously explored in untargeted metabolomics and proteomics studies. As a complementary approach, applying our framework to peaks retained after filtering by 30% RSD across pooled QC samples was able to further distinguish poorly integrated peaks that were not removed from filtering alone. An R implementation of these classifiers and the overall computational approach is available as the MetaClean package at https://CRAN.R-project.org/package=MetaClean.
Conclusion
Our work represents an important step forward in developing an automated tool for filtering out unreliable peak integrations in untargeted LC–MS metabolomics data.




Similar content being viewed by others
Data availability
The metabolomics and metadata analyzed in this paper are available via the Metabolomics Workbench (https://www.metabolomicsworkbench.org/) Study IDs ST000726 and ST000695, and via MetaboLights (https://www.ebi.ac.uk/metabolights/) Study IDs MTBLS354 and MTBLS306.
Code availability
The MetaClean R package developed in this study is available at https://CRAN.R-project.org/package=MetaClean.
References
Alpaydin, E. (2014). Introduction to machine learning (3rd ed.). London: The MIT Press.
Arlot, S., & Celisse, A. (2010). A survey of cross-validation procedures for model selection. Statistics Surveys, 4, 40–79. https://doi.org/10.1214/09-SS054
Borgsmüller, N., Gloaguen, Y., Opialla, T., Blanc, E., Sicard, E., Royer, A.-L., et al. (2019). WiPP: Workflow for improved peak picking for gas chromatography-mass spectrometry (GC-MS) data. Metabolites, 9(9), 171. https://doi.org/10.3390/metabo9090171
Broadhurst, D., Goodacre, R., Reinke, S. N., Kuligowski, J., Wilson, I. D., Lewis, M. R., & Dunn, W. B. (2018). Guidelines and considerations for the use of system suitability and quality control samples in mass spectrometry assays applied in untargeted clinical metabolomic studies. Metabolomics, 14(6), 72. https://doi.org/10.1007/s11306-018-1367-3
Calvo, B., & Santafé, G. (2016). Guidelines and considerations for the use of system suitability and quality control samples in mass spectrometry assays applied in untargeted clinical metabolomic studies. The R Journal, 8(1), 248.
Chong, J., Wishart, D. S., & Xia, J. (2019). Using MetaboAnalyst 4.0 for comprehensive and integrative metabolomics data analysis. Current Protocols in Bioinformatics, 68(1), e86. https://doi.org/10.1002/cpbi.86
Coble, J. B., & Fraga, C. G. (2014). Comparative evaluation of preprocessing freeware on chromatography/mass spectrometry data for signature discovery. Journal of Chromatography A, 1358, 155–164. https://doi.org/10.1016/j.chroma.2014.06.100
Demsˇar, J. (2006). Statistical comparisons of classifiers over multiple data sets. Journal of Machine Learning Research, 7, 1–30. https://doi.org/10.5555/1248547.1248548
Dunn, W. B., Broadhurst, D., Begley, P., Zelena, E., Francis-McIntyre, S., Anderson, N., et al. (2011). Procedures for large-scale metabolic profiling of serum and plasma using gas chromatography and liquid chromatography coupled to mass spectrometry. Nature Protocols, 6(7), 1060–1083. https://doi.org/10.1038/nprot.2011.335
Eshghi, S. T., Auger, P., & Mathews, W. R. (2018). Quality assessment and interference detection in targeted mass spectrometry data using machine learning. Clinical Proteomics. https://doi.org/10.1186/s12014-018-9209-x
Haug, K., Cochrane, K., Nainala, V. C., Williams, M., Chang, J., Jayaseelan, K. V., & Oonovan, C. (2019). MetaboLights: A resource evolving in response to the needs of its scientific community. Nucleic Acids Research. https://doi.org/10.1093/nar/gkz1019
Kuhn, M. (2008). Building predictive models in R using the caret package. Journal of Statistical Software, 28(1), 1–26.
Lever, J., Krzywinski, M., & Altman, N. S. (2016). Points of Significance: Classification evaluation. Nature methods, 13(8), 603–604. https://doi.org/10.1038/nmeth.3945
Libiseller, G., Dvorzak, M., Kleb, U., Gander, E., Eisenberg, T., Madeo, F., et al. (2015). IPO: A tool for automated optimization of XCMS parameters. BMC Bioinformatics, 16(1), 118. https://doi.org/10.1186/s12859-015-0562-8
Mahieu, N. G., Spalding, J. L., & Patti, G. J. (2016). Warpgroup: Increased precision of metabolomic data processing by consensus integration bound analysis. Bioinformatics (Oxford, England), 32(2), 268–275. https://doi.org/10.1093/bioinformatics/btv564
MetaboLights. (2016a). MTBLS354: Lipid metabolites as potential diagnostic and prognostic biomarkers for acute community acquired pneumonia. Retrieved March 4, 2020, from https://www.ebi.ac.uk/metabolights/MTBLS354.
MetaboLights. (2016b). MTBLS306:Metabolic profiling of submaximal exercise at a standardised relative intensity in healthy adults. Retrieved September 4, 2020, from https://www.ebi.ac.uk/metabolights/MTBLS306.
Metabolomics Workbench. (2017a). PR000523, ST000726. https://doi.org/10.21228/M82D6X
Metabolomics Workbench. (2017b). PR000492, ST000625. https://doi.org/10.21228/M8G31N
Muhsen Ali, A., Burleigh, M., Daskalaki, E., Zhang, T., Easton, C., & Watson, D. G. (2016). Metabolomic profiling of submaximal exercise at a standardised relative intensity in healthy adults. Metabolites, 6(1), 9. https://doi.org/10.3390/metabo6010009
Myers, O. D., Sumner, S. J., Li, S., Barnes, S., & Du, X. (2017). Detailed investigation and comparison of the XCMS and MZmine 2 chromatogram construction and chromatographic peak detection methods for preprocessing mass spectrometry metabolomics data. Analytical Chemistry, 89(17), 8689–8695. https://doi.org/10.1021/acs.analchem.7b01069
Pluskal, T., Castillo, S., Villar-Briones, A., & Orešič, M. (2010). MZmine 2: Modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinformatics, 11(1), 395. https://doi.org/10.1186/1471-2105-11-395
Rafiei, A., & Sleno, L. (2015). Comparison of peak-picking workflows for untargeted liquid chromatography/high-resolution mass spectrometry metabolomics data analysis. Rapid Communications in Mass Spectrometry, 29, 119–127. https://doi.org/10.1002/rcm.7094
Schiffman, C., Petrick, L., Perttula, K., Yano, Y., Carlsson, H., Whitehead, T., et al. (2019). Filtering procedures for untargeted LC-MS metabolomics data. BMC Bioinformatics, 20(1), 334. https://doi.org/10.1186/s12859-019-2871-9
Smith, C. A., Want, E. J., O’Maille, G., Abagyan, R., & Siuzdak, G. (2006). XCMS: Processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Analytical Chemistry, 78(3), 779–787. https://doi.org/10.1021/ac051437y
Sud, M., Fahy, E., Cotter, D., Azam, K., Vadivelu, I., Burant, C., et al. (2016). Metabolomics Workbench: An international repository for metabolomics data and metadata, metabolite standards, protocols, tutorials and training, and analysis tools. Nucleic Acids Research, 44, D463–D470. https://doi.org/10.1093/nar/gkv1042
To, K. K. W., Lee, K.-C., Wong, S. S. Y., Sze, K.-H., Ke, Y.-H., Lui, Y.-M., et al. (2016). Lipid metabolites as potential diagnostic and prognostic biomarkers for acute community acquired pneumonia. Diagnostic Microbiology and Infectious Disease, 85(2), 249–254. https://doi.org/10.1016/j.diagmicrobio.2016.03.012
Uppal, K., Soltow, Q. A., Strobel, F. H., Pittard, W. S., Gernert, K. M., Yu, T., & Jones, D. P. (2013). xMSanalyzer: Automated pipeline for improved feature detection and downstream analysis of large-scale, non-targeted metabolomics data. BMC Bioinformatics, 14(1), 15. https://doi.org/10.1186/1471-2105-14-15
Want, E. J., Wilson, I. D., Gika, H., Theodoridis, G., Plumb, R. S., Shockcor, J., et al. (2010). Global metabolic profiling procedures for urine using UPLC–MS. Nature Protocols, 5(6), 1005–1018. https://doi.org/10.1038/nprot.2010.50
Whalen, S., Pandey, O. P., & Pandey, G. (2016). Predicting protein function and other biomedical characteristics with heterogeneous ensembles. Methods, 93, 92–102. https://doi.org/10.1016/j.ymeth.2015.08.016
Yang, P., Yang, Y. H., Zhou, B. B., & Zomaya, A. Y. (2010). A review of ensemble methods in bioinformatics. Current Bioinformatics, 5(4), 296–308. https://doi.org/10.2174/157489310794072508
Yu, T., Park, Y., Johnson, J. M., & Jones, D. P. (2009). apLCMS—adaptive processing of high-resolution LC/MS data. Bioinformatics, 25(15), 1930–1936. https://doi.org/10.1093/bioinformatics/btp291
Zhang, W., & Zhao, P. X. (2014). Quality evaluation of extracted ion chromatograms and chromatographic peaks in liquid chromatography/mass spectrometry-based metabolomics data. BMC Bioinformatics, 15(Suppl 11), S5. https://doi.org/10.1186/1471-2105-15-S11-S5
Acknowledgements
The authors would like to thank Miao Yu and Gabriel Hoffman for helping develop, and Yan-chak Li for testing the MetaClean R package. They also thank the authors of the Zhang et al. (2014) study for sharing the Matlab code implementing their peak quality metrics.
Funding
This work was supported by the National Institute of Environmental Health Sciences (NIEHS) Grant Nos. U2CES030859, P30ES23515, R01ES031117, and R21ES030882, and the National Institute of General Medical Sciences (NIGMS) Grant No. R01GM114434.
Author information
Authors and Affiliations
Contributions
KC performed research, analyzed data, and wrote the paper. LP conceived and designed the study, curated data, interpreted the results, and wrote the paper. GP conceived and designed study, analyzed data, interpreted the results, and wrote the paper. All the authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflicts of interest
All authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human and/or animal participants performed by any of the authors, and only utilizes publicly available data and software.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chetnik, K., Petrick, L. & Pandey, G. MetaClean: a machine learning-based classifier for reduced false positive peak detection in untargeted LC–MS metabolomics data. Metabolomics 16, 117 (2020). https://doi.org/10.1007/s11306-020-01738-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-020-01738-3