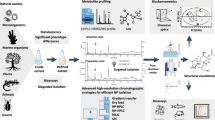
Abstract
Introduction
Lipidomics can reveal global alterations in a broad class of molecules whose functions are innately linked to physiology. Monitoring changes in the phospholipid composition of biological membranes in response to stressors can aid the development of targeted therapies. However, exact quantitation of cardiolipins is not a straightforward task due to low ionization efficiencies and poor chromatographic separation of these compounds.
Objective
The aim of this study was to develop a quantitative method for the detection of cardiolipins and other phospholipids using both a targeted and untargeted analyses with a Q-Exactive.
Methods
HILIC chromatography and high-resolution mass spectrometry with parallel reaction monitoring was used to measure changes in lipid concentration. Internal standards and fragmentation techniques allowed for the reliable quantitation of lipid species including: lysyl-phosphatidylglycerol, phosphatidylglycerol, and cardiolipin.
Results
The untargeted analysis was capable to detecting 6 different phospholipid classes as well as free fatty acids. The targeted analysis quantified up to 23 cardiolipins, 10 phosphatidylglycerols and 10 lysyl-phosphatidylglycerols with detection limits as low as 50 nM. Biological validation with Enterococcus faecalis demonstrates sensitivity in monitoring the incorporation of exogenously supplied free fats into membrane phospholipids. When supplemented with oleic acid, the amount of free oleic acid in the membrane was 100 times greater and the concentration of polyunsaturated cardiolipin increased to over 3.5 µM compared to controls.
Conclusions
This lipidomics method is capable of targeted quantitation for challenging biologically relevant cardiolipins as well as broad, untargeted lipid profiling.



Similar content being viewed by others
References
Bao, Y., Sakinc, T., Laverde, D., Wobser, D., Benachour, A., Theilacker, C., et al. (2012). Role of mprF1 and mprF2 in the pathogenicity of Enterococcus faecalis. PLoS ONE, 7, e38458.
Beate, F., & Jürgen, S. (2009). Application of MALDI-TOF mass spectrometry in lipidomics. European Journal of Lipid Science and Technology, 111, 83–98.
Beney, L., & Gervais, P. (2001). Influence of the fluidity of the membrane on the response of microorganisms to environmental stresses. Applied Microbiology and Biotechnology, 57, 34–42.
Bird, S. S., Marur, V. R., Sniatynski, M. J., Greenberg, H. K., & Kristal, B. S. (2011). Lipidomics profiling by high-resolution LC–MS and high-energy collisional dissociation fragmentation: Focus on characterization of mitochondrial cardiolipins and monolysocardiolipins. Analytical Chemistry, 83, 940–949.
Bligh, E. G., & Dyer, W. J. (1959). A rapid method of total lipid extraction and purification. Canadian Journal of Biochemistry and Physiology, 37, 911–917.
Chapman, D. J., De-Felice, J., & Barber, J. (1985). Characteristics of chloroplast thylakoid lipid composition associated with resistance to triazine herbicides. Planta, 166, 280–285.
Clasquin, M. F., Eugene, M., & Rabinowitz, D. J. (2012). LC-MS data processing with MAVEN: A metabolomic analysis and visualization engine. Current Protocols in Bioinformatics, 37, 14.11.1–14.11.23.
Cronan, J. E. (2003). Bacterial membrane lipids: Where do we stand? Annual Review of Microbiology, 57, 203–224.
Dörmann, P., & Benning, C. (2002). Galactolipids rule in seed plants. Trends in Plant Science, 7, 112–118.
Dowhan, W. (1997). Molecular basis for membrane phospholipid diversity: Why are there so many lipids? Annual Review of Biochemistry, 66, 199–232.
Fahy, E., Subramaniam, S., Murphy, R. C., Nishijima, M., Raetz, C. R. H., Shimizu, T., et al. (2009). Update of the LIPID MAPS comprehensive classification system for lipids. Journal of Lipid Research, 50, S9–S14.
Fozo, E. M., & Rucks, E. A. (2016) Chapter two—The making and taking of lipids: The role of bacterial lipid synthesis and the harnessing of host lipids in bacterial pathogenesis. In R. K. Poole (Ed.), Advances in microbial physiology (pp. 51–155) Academic Press.
Hamilton, J. A., & Kamp, F. (1999). How are free fatty acids transported in membranes? Is it by proteins or by free diffusion through the lipids? Diabetes, 48, 2255–2269.
Harp, J. R., Saito, H. E., Bourdon, A. K., Reyes, J., Arias, C. A., Campagna, S. R., et al. (2016). Exogenous fatty acids protect Enterococcus faecalis from daptomycin-induced membrane stress independently of the response regulator LiaR. Applied and Environmental Microbiology, 82, 4410–4420.
Hines, K. M., Waalkes, A., Penewit, K., Holmes, E. A., Salipante, S. J., Werth, B. J., et al. (2017). Characterization of the mechanisms of daptomycin resistance among gram-positive bacterial pathogens by multidimensional lipidomics. mSphere, 2, e00492.
Holman, J. D., Tabb, D. L., & Mallick, P. (2014). Employing proteowizard to convert raw mass spectrometry data. Current Protocols in Bioinformatics, 46, 13241–13249.
Iverson, S. J., Lang, S. L., & Cooper, M. H. (2001). Comparison of the bligh and dyer and folch methods for total lipid determination in a broad range of marine tissue. Lipids, 36, 1283–1287.
Joo, H.-S., & Otto, M. (2015). Mechanisms of resistance to antimicrobial peptides in staphylococci. Biochimica et Biophysica Acta (BBA)—Biomembranes, 1848, 3055–3061.
Kessner, D., Chambers, M., Burke, R., Agus, D., & Mallick, P. (2008). ProteoWizard: Open source software for rapid proteomics tools development. Bioinformatics, 24, 2534–2536.
Kilelee, E., Pokorny, A., Yeaman, M. R., & Bayer, A. S. (2010). Lysyl-phosphatidylglycerol attenuates membrane perturbation rather than surface association of the cationic antimicrobial peptide 6 W-RP-1 in a model membrane system: implications for daptomycin resistance. Antimicrobial Agents and Chemotherapy, 54, 4476–4479.
Klausner, R. D., Kleinfeld, A. M., Hoover, R. L., & Karnovsky, M. J. (1980). Lipid domains in membranes. Evidence derived from structural perturbations induced by free fatty acids and lifetime heterogeneity analysis. Journal of Biological Chemistry, 255, 1286–1295.
Minkler, P. E., & Hoppel, C. L. (2010). Separation and characterization of cardiolipin molecular species by reverse-phase ion pair high-performance liquid chromatography-mass spectrometry. Journal of Lipid Research, 51, 856–865.
Mishra, N. N., Bayer, A. S., Tran, T. T., Shamoo, Y., Mileykovskaya, E., Dowhan, W., et al. (2012). Daptomycin resistance in enterococci is associated with distinct alterations of cell membrane phospholipid content. PLoS ONE, 7, e43958.
Nishi, H., Komatsuzawa, H., Fujiwara, T., McCallum, N., & Sugai, M. (2004). Reduced content of lysyl-phosphatidylglycerol in the cytoplasmic membrane affects susceptibility to moenomycin, as well as vancomycin, gentamicin, and antimicrobial peptides, in Staphylococcus aureus. Antimicrobial Agents and Chemotherapy, 48, 4800–4807.
Oliver, P. M., Crooks, J. A., Leidl, M., Yoon, E. J., Saghatelian, A., & Weibel, D. B. (2014). Localization of anionic phospholipids in Escherichia coli cells. Journal of Bacteriology, 196, 3386–3398.
Olofsson, G., & Sparr, E. (2013). Ionization constants pKa of cardiolipin. PLoS ONE, 8, e73040.
Rashid, R., Cazenave-Gassiot, A., Gao, I. H., Nair, Z. J., Kumar, J. K., Gao, L., et al. (2017). Comprehensive analysis of phospholipids and glycolipids in the opportunistic pathogen Enterococcus faecalis. PLoS ONE, 12, e0175886.
Renner, L. D., & Weibel, D. B. (2011). Cardiolipin microdomains localize to negatively curved regions of Escherichia coli membranes. Proceedings of the National Academy of Sciences, 108, 6264–6269.
Romantsov, T., Guan, Z., & Wood, J. M. (2009). Cardiolipin and the osmotic stress responses of bacteria. Biochimica et Biophysica Acta (BBA)—Biomembranes, 1788, 2092–2100.
Saito, H. E., Harp, J. R., & Fozo, E. M. (2014). Incorporation of exogenous fatty acids protects Enterococcus faecalis from membrane-damaging agents. Applied and Environmental Microbiology, 80, 6527–6538.
Saito, H. E., Harp, J. R., & Fozo, E. M. (2018). Enterococcus faecalis responds to individual exogenous fatty acids independently of their degree of saturation or chain length. Applied and Environmental Microbiology, 84, e01633.
Scherer, M., Schmitz, G., & Liebisch, G. (2010). Simultaneous quantification of cardiolipin, bis(monoacylglycero)phosphate and their precursors by hydrophilic interaction LC–MS/MS including correction of isotopic overlap. Analytical Chemistry, 82, 8794–8799.
Schwalbe-Herrmann, M., Willmann, J., & Leibfritz, D. (2010). Separation of phospholipid classes by hydrophilic interaction chromatography detected by electrospray ionization mass spectrometry. Journal of Chromatography A, 1217, 5179–5183.
Sohlenkamp, C., & Geiger, O. (2016). Bacterial membrane lipids: Diversity in structures and pathways. FEMS Microbiology Reviews, 40, 133–159.
Sprott, G. D., Larocque, S., Cadotte, N., Dicaire, C. J., McGee, M., & Brisson, J. R. (2003). Novel polar lipids of halophilic eubacterium Planococcus H8 and archaeon Haloferax volcanii. Biochimica et Biophysica Acta (BBA)—Molecular and Cell Biology of Lipids, 1633, 179–188.
Sumner, L. W., Amberg, A., Barrett, D., Beale, M. H., Beger, R., Daykin, C. A., et al. (2007). Proposed minimum reporting standards for chemical analysis. Metabolomics, 3, 211–221.
Tran, T. T., Panesso, D., Mishra, N. N., Mileykovskaya, E., Guan, Z., Munita, J. M., et al. (2013). Daptomycin-resistant Enterococcus faecalis diverts the antibiotic molecule from the division septum and remodels cell membrane phospholipids. mBio, 4, e00281.
Valianpour, F., Wanders, R. J. A., Barth, P. G., Overmars, H., & van Gennip, A. H. (2002). Quantitative and compositional study of cardiolipin in platelets by electrospray ionization mass spectrometry: Application for the identification of barth syndrome patients. Clinical Chemistry, 48, 1390–1397.
Van Mooy, B. A. S., Fredricks, H. F., Pedler, B. E., Dyhrman, S. T., Karl, D. M., Koblížek, M., et al. (2009). Phytoplankton in the ocean use non-phosphorus lipids in response to phosphorus scarcity. Nature, 458, 69.
Watson, A. D. (2006). Thematic review series: Systems biology approaches to metabolic and cardiovascular disorders. Lipidomics: A global approach to lipid analysis in biological systems. Journal of Lipid Research, 47, 2101–2111.
Funding
The authors would like to acknowledge the National Institute of Health (NIH) Grant #R01 AI116571-03 for financial support.
Author information
Authors and Affiliations
Contributions
EDT, SRC, and EMF designed the project. EDT and BMW designed and conducted targeted analysis. EDT and ATF designed and conducted the untargeted analysis. JRH grew and extracted lipids from the bacteria. EDT, BMW, JRH, SRC, and EMF wrote the manuscript. All authors read and approved the manuscript. Mass spectrometric analyses were performed at the University of Tennessee, Knoxville Biological and Small Molecule Mass Spectrometry Core with the assistance of Dr. Hector F. Castro.
Corresponding author
Ethics declarations
Conflict of interest
The authors declares that he/she has no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tague, E.D., Woodall, B.M., Harp, J.R. et al. Expanding lipidomics coverage: effective ultra performance liquid chromatography-high resolution mass spectrometer methods for detection and quantitation of cardiolipin, phosphatidylglycerol, and lysyl-phosphatidylglycerol. Metabolomics 15, 53 (2019). https://doi.org/10.1007/s11306-019-1512-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-019-1512-7