Abstract
Nanobodies are the smallest known antigen-binding molecules to date. Their small size, good tissue penetration, high stability and solubility, ease of expression, refolding ability, and negligible immunogenicity in the human body have granted them excellence over conventional antibodies. Those exceptional attributes of nanobodies make them promising candidates for various applications in biotechnology, medicine, protein engineering, structural biology, food, and agriculture. This review presents an overview of their structure, development methods, advantages, possible challenges, and applications with special emphasis on infectious diseases-related ones. A showcase of how nanobodies can be harnessed for applications including neutralization of viruses and combating antibiotic-resistant bacteria is detailed. Overall, the impact of nanobodies in vaccine design, rapid diagnostics, and targeted therapies, besides exploring their role in deciphering microbial structures and virulence mechanisms are highlighted. Indeed, nanobodies are reshaping the future of infectious disease prevention and treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The discovery of monoclonal antibodies (mAbs) has significantly influenced the field of biological industries. This was implemented by Orthoclone, the first Food and Drug Administration (FDA) approved mAb which has a crucial role in preventing rejection in organ transplantation (Starzl and Fung 1986). Since then, an enormous number of mAbs have been well-established and marketed for their beneficial clinical applications including targeted treatment and enhanced therapeutic precision. However, mAbs use was restricted owing to the sophisticated structure and large size which affect their binding specificity, tissue penetration, and clearance time in certain diseases (Buss et al. 2012). Additionally, the synthesis and production of mAbs are costly and time-consuming.
Coincidence plays a vital role in most of the scientific breakthroughs and the same narrative transpired with the first observation of a peculiar antibody molecule that later became a defining milestone in history, currently known as nanobodies (Nbs). In 1993, the Hamers’ lab serendipitously discovered naturally occurring heavy-chain antibodies in the serum of the camel (Hamers-Casterman et al. 1993). Later in 1995, Greenberg and co-workers detected single-domain antibodies from nurse sharks (Ginglymostoma cirratum) (Greenberg et al. 1995). These molecules differ from their conventional ones in their composition which includes only the heavy-chain variable dimers while missing their light-chain counterparts. Nevertheless, they possess an extensive antigen-binding repertoire.
Nanobodies (Nbs) “also referred to as single-domain antibodies (sdAb)’’ are the antigen-binding molecules engineered from the camelid or sharks heavy chain antigen-binding domain that are called the camelid variable heavy-chain region (VHH) and immunoglobulin new antigen receptor (VNAR), respectively (Schrankel et al. 2019). While the human IgG immunoglobulin weighs ~ 150 kDa, the heavy-chain antibody weighs ~ 80 to 90 kDa, and the derived Nbs are ~ 12 to 15 kDa (Pillay and Muyldermans 2021; Vincke and Muyldermans 2012). They are one-tenth the size of a normal antibody (Schrankel et al. 2019), making their production and utilization far more applicable. They also possess low immunogenicity owing to their small size, which is around 110 amino acids (~ 4.4 nm high; ~ 2.5 to 2.8 nm diameter) (Cortez-Retamozo et al. 2004; Sánchez-García et al. 2021). Moreover, Nbs can bind to embedded epitopes that are not accessible to complete antibodies and have a greater affinity and selectivity in targeting the active sites of enzymes and receptors. It is worth noting that Nbs exhibit remarkable stability, demonstrated by their ability to withstand some drastic conditions of pH, pressure, and temperature while maintaining their antigen-binding capacity. They can tolerate extreme pHs (pH 3.0–9.0), and high pressure (500–750 MPa) (Jovčevska and Muyldermans 2020). Nbs are also known to exhibit long shelf-life with high storage stability at different temperatures; 4 °C and -20 °C for long storage periods (months), and 37°C for shorter ones (weeks). Moreover, some studies reported their heat tolerance to higher temperatures (60–80 °C). Yet improper Nbs refolding by heat denaturation represents a great concern. Furthermore, Nbs demonstrate high stability against proteolytic enzymes and some chemical denaturants like urea (De Vos et al. 2013; Jovčevska and Muyldermans 2020). On another front, the specificity of the Nb can be generated from cell-based microbial expression systems such as Escherichia coli, yeasts, or cell-free platforms (using ribosomes) (Schrankel et al. 2019). This simple yet critical approach can have a significant effect on the reduction of Nbs production costs.
Nbs have already been used in diverse fields and particularly notable is the first Nb approved for a therapeutic indication in 2018, named Caplacizumab, which is used for acquired thrombotic thrombocytopenic purpura (Duggan 2018). Nbs are tested in a wide range of prospective innovations, such as investigating the viability of the VHHs in phage display, testing its potential in shampoos for dandruff reduction and introducing the first evidence of Nbs inhibiting the cell-free and cell-to-cell transmission in hepatitis C infection (Dolk et al. 2005; Tarr et al. 2013). In addition, Nbs are also tested to serve in identifying tumor cells by targeting human growth factor cell receptors (HER2) and carbonic anhydrase IX (CAIX) (Keyaerts et al. 2016; Kijanka et al. 2016). The wide array of possible revolutionary applications offered by these small biomolecules will inevitably boost Nbs utilization. In the current review, the Nbs’ structure, methods of production, advantages, disadvantages and potential applications will be discussed with emphasis on their potential role in infectious diseases.
The structure of different forms of antibodies
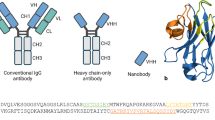
To properly understand the distinctiveness of Nbs, a structural comparative overview of Nbs, conventional antibodies, and the parent heavy chain antibodies is presented (Fig. 1) and discussed as follows.
Comparison of Nbs’ structure to other antigen-binding moieties. A schematic diagram showing the difference between Nbs and other antigen-binding moieties. A Conventional antibody with its heavy chain (VH) (pink color) and light chain (VL) (gray color). B Single chain fragment variable (scFv), which contains a pair of VH and VL domains connected by an oligopeptide bond. C Three different heavy chain antibodies; the camelid heavy chains contain VHH segment, hinge, CH2 and CH3 with long (IgG2) or short hinge (IgG3), and the shark heavy chain containing one variable domain and five constant domains. The three heavy chains exhibit single domain antibody (sdAb). The sdAb is formed of three hypervariable sections surrounded by nine β-sheet strands connected by disulfide bond
The binding specificity of the full-length antibodies is determined by variable regions in their heavy (VH) and light chains (VL). The two light chains are composed of a variable domain (VL) and a constant domain (CL). The VH, CH1, hinge, CH2, and CH3 domains make up the two heavy chains (VH), with the CH1 domain serving as a key connection between the heavy and light chains (Muyldermans 2013; Wanner et al. 2021). Collectively, they generate a diversity of at least 1015 B-cell receptors (BCRs) in humans (Mitchell and Colwell 2018). The linkage of the CH2 and the CH3 makes the crystallizable fragment (Fc) portion of the antibody while the antigen-binding (Fab) region is composed of the heavy chain's outer domains (CH1 & VH) as well as the light chain's variable and constant domains (CL & VL). The pairing of the VH-VL by an oligopeptide generates the smallest functional antigen-binding unit, known as the single-chain fragment variable (scFv), with a size of ~ 30 kDa that can be created from the full-size antibodies (Muyldermans 2013). However, unlike VHHs, scFvs have lower affinities, reduced half-life, and stability, as well as lower thermostability when compared to their parent antibodies. As a result, there is a higher probability of aggregation and subsequent risk of immunogenicity (Bates and Power 2019).
The camelid heavy-chain antibodies on the other hand lack both the light chains and the CH1, which gives them an advantageous small size, with a molecular weight of ~ 90 kDa. The dromedary heavy-chain antibodies carry only the VHH segment, hinge, CH2 and CH3 fragments with a direct connection of the rearranged VHH exon to the hinge region belonging to one of two types of hinge isotypes: long (IgG2) or short (IgG3), referring to the fraction's hinge length. In this case, antigen recognition is through the variable domain of the heavy chain. Accordingly, the compact design of Nbs allows better adaptability for hidden targets (Arbabi-Ghahroudi 2017; Muyldermans 2013). Similarly, the antibodies devoid of light chains found in cartilaginous fish consist of one variable domain followed by five constant domains [(V-NAR)-5(C-NAR)] (Deffar et al. 2009; Zielonka et al. 2015).
The VHHs molecules derived from the camelid heavy chains restrict the antigen binding to a single domain of about 110 amino acids. These molecules comprise three hypervariable sections (HV) that localize the sequence variation of the variable domains (V) and are surrounded by a conserved framework (FR). Nine β-sheet strands (A-B-C–C'-C'-D-D-E–F-G) make up the folded variable domain, which is arranged into four-stranded β-sheets and five-stranded β-sheets joined by loops and a conserved disulfide bond. The HV regions are arranged into three loops (H1, H2, and H3) that connect the stranded β-sheets. A continuous surface is formed by the cluster at the N-terminal that is complementary to the surface of the epitopes or antigens (paratope) and this area is referred to as the complementarity-determining region (CDR). The sequence within the loops is highly variable, but the extent of the variation is limited except for the H3 loop (CDR3) (Muyldermans 2013). Controversially, the conventional antibodies were thought to have wider diversity compared to Nbs as the latter have paratopes of smaller size. However, this notion was disproven by the large H1 loop (CDR1) that is responsible for antigen recognition and was found to be longer than those in the variable domain of the conventional antibody’s heavy chain, subsequently serving in largening the paratope size and exhibiting diverse loop architectures (Nguyen et al. 2000). Within the conserved FR2, the highly conserved hydrophobic amino acids normally found in the full-size antibodies, are replaced in VHHs with more hydrophilic amino acids, rendering them more soluble (Asaadi et al. 2021; Muyldermans 2013).
The VNAR domains, on the other hand, are members of the immunoglobulin’s superfamily and hence they have a β-sandwich structure. The VNARs lack the hinge region yet there is a wide space for interacting with multiple epitopes which is enhanced by the dimerization between C1 and C3 domains. Unlike the mammalian variable domain counterpart, the β-sandwich fold in the VNAR only has eight strands instead of ten. With a size of roughly 11–12 kDa, the VNARs are believed to be the smallest antibody-like antigen-binding domains known in the animal kingdom (Stanfield et al. 2004; Zielonka et al. 2015). This structure results in fewer antigen-binding loops (CDR1 & CDR3) compared to antibodies, but the elongated CDR3 compensates for this (Feige et al. 2014; Könning et al. 2017; Zielonka et al. 2015). Still, the VNARs' diversity, like that of the VHHs, is predominantly seen in the CDR3 sequences. Two cysteines in FR1 and FR3 form a stabilizing disulfide bond, and additional ones in CDR3 can provide extra stability (English et al. 2020; Feige et al. 2014; Feng et al. 2019).
In conclusion, compared to the standard antibody binding sites, antigen-binding sites in VHHs and the VNARs are smaller in terms of molecular surface area and diameters. They differ from the typical canonical structures of the full-length antibody in their non-canonical CDR1 and CDR2 structures, as well as an elongated CDR3 loop length distribution. However, they have similar amino acid compositions and as a group they appear to be no longer in the distance measured from the CDR base to the tip than the conventional antibodies (Henry and MacKenzie 2018). For protein-binding, rather than operating six-loop configurations like typical antibodies, Nbs only use their three CDR loops. They exert their expanded CDR3 loop to penetrate the active site or the CDR2 loop in circumstances where the Nb's standard CDR3 loop is insufficient to protrude to the antigen. (Desmyter et al. 2002; Henry and MacKenzie 2018; Sela-Culang et al. 2013).
Production of nanobodies
The production of sdAb fragments traditionally entails the amplification of VHH or VNAR gene segments at an affordable low cost. They are then cloned into a display system, whether it is a bacteria, yeast, phage, or ribosome, followed by the generation of a large collection of clones "library" accompanied by biopanning of the high-affinity antigen-specific clones and their retrieval (Fig. 2).
Inherently, for immune library generation, the stages of Nbs production generally commence by immunizing healthy young adults including dromedaries, camels, llamas, alpacas, or sharks with a protein cocktail to generate a library of at least 106–108 individual clones (Müller et al. 2012; Muyldermans 2021b). Over the course of a few months, the animals can be routinely injected with the target immunogen. Since the used animals are outbred, it is recommended that more than one is immunized at this early stage. Each animal is thought to elicit a different immune response, with a subsequent large repertoire of Nbs from which the best-performing clone is selected (Muyldermans 2021b). Affinity maturation and class switch recombination are induced by deliberate repeated immunization which leads to boosting the odds of detecting VHHs with the targeted functional features that may not be existent in naïve libraries (Ingram et al. 2018). Extraction of mRNA is done from the blood acquired after the immunization step, then the mRNA is transformed into cDNA and utilized to amplify the VHH gene segments (Muyldermans 2021b). Constructing libraries through animal immunization has some limitations such as being time-consuming and costly, and it may also generate redundant subpopulations of certain antigens. Additionally, when it comes to non-immunogenic molecules like RNA or DNA, which fail to elicit an immune response, they are not the best choice (Muyldermans 2021b; Sabir et al. 2014).
Another significant limitation of the immunization libraries is the limited target space for sensitive proteins. Targets, such as many human membrane transporters, easily unfold upon injection primarily due to the adjuvants used and the dromedary's high body temperature. Additionally, unless their affinities are extremely high, non-covalent ligands dissociate from the protein shortly after injection, making it difficult to promote target conformations (Zimmermann et al. 2018). It is worth noting that immunization requires access to animal facilities, and this may not always be feasible. On another front, and from an ethical point of view, animal usage to that end is strongly discouraged for compounds that are poisonous, contagious, or harmful to both animals and environment. Hence, steering directions are currently implemented towards the use of other Nb repertoires such as naïve and synthetic libraries which do not require animals to be immunized against bacteria, viruses, or toxoids (Gray et al. 2016).
As for the construction of a naïve library, a pool of blood from multiple non-immunized animals is required. This method has the advantage of being rapid and the ability to recover a VHH repertoire that should at least acquire the size of 109–1011 clones, with the added benefit of being more diversified (Muyldermans 2021b; Sabir et al. 2014). On the other hand, taking up to 10 L of blood to build a diversified naïve Nb library with around 1010 different VHH clones is tedious (Muyldermans 2021b). However, the procedure has been found to yield over-adequate Nb libraries of a size of 107 with as little as 23 mL, from which high-affinity Nbs can be extracted (Sabir et al. 2014).
Synthetic libraries are the third source for Nbs and they can provide access to bigger repertoires without the benefits of target immunization and affinity maturation (Ingram et al. 2018). A stable and well-expressed Nb scaffold, preferably with a crystal structure, is usually chosen for the construction of a synthetic library without the need to draw blood from animals (Muyldermans 2021b). Synthetic libraries have a diverse clone size of 109–1015 and often exhibit a single shape and are randomized in only one region of their surface (Muyldermans 2021b; Zimmermann et al. 2018). A single or a few Nbs with desirable biochemical features are randomly selected and their sequences are amplified by PCR (Muyldermans 2021b). Following that, the PCR products are ligated into phage or ribosome display, or both and three synthetic Nbs selection platforms tailored to membrane protein targets are then engineered with varying CDR3 loop lengths and configurations. The Nb library is displayed using both phage and ribosome systems and created by analyzing many deposited camelids VHHs structures (Zimmermann et al. 2018).
It is also worth noting that Nb libraries can be developed from human origins through phage-display technology under the hypothesis that certain VH framework sites can compensate for the loss of the light chain, resulting in soluble human Nbs (Wu et al. 2020). A previous study reported cloning of 17 human germline immunoglobulin heavy chain variable region (IGHV) alleles, and expressing them in E. coli, and then characterizing their properties, along with a camelid Nb as a control. Another previously experimented approach is that fully human single-domain antibodies were obtained by grafting the complementarity determining regions (CDR1, CDR2, and CDR3) from naïve libraries into the FR regions of a human germline immunoglobulin VH variable region allele (Wu et al. 2020). This technology promises antibodies derived entirely from human sequences which exhibit less immunogenicity compared to camelid or humanized Nbs, leading to improved safety and efficacy for human use.
Nbs can be expressed in both prokaryotic and eukaryotic systems, such as E. coli, S. cerevisiae, and Pichia pastoris. The most common approach for generating Nbs is to promote their secretion in the P. pastoris or the E. coli periplasm (Chen et al. 2019; de Marco 2020). The periplasm's oxidizing conditions promote the formation of disulfide bonds, which help in stabilizing the Nb structure. After an osmotic shock step to permeabilize the bacterial outer membrane, the folded binders are normally recovered in the supernatant and affinity purification is used to recover the Nbs (de Marco 2020). It is also noted that upon precipitation of the highly temperature-sensitive E. coli proteins, heat incubation of the supernatant has been successfully used to purify the comparatively thermal-resistant VHHs (Olichon et al. 2007). Although periplasmic extraction has its benefits in terms of protein folding, it also has limitations, such as aggregation and low yields of proteins. The latter could be due to a number of reasons, including the secretion system saturation, the absence of adequate chaperone machinery that can inhibit improper folding at high expression rates, high proteolytic activity, and a lengthy-expression technique (Pleiner et al. 2015).
Advantages of nanobodies
In terms of size, the single variable segment of the heavy chain antibodies is the smallest functional antigen-binding domain natively created by the adaptive immune system (Muyldermans 2013). The myriad uses of Nbs can be attributed to their exceptionally small and structurally convenient nature that in turn accounts for their fast tissue penetration and short half-life. In terms of antigen-binding capabilities, the diversity in the VHHs and VNAR loop structures dramatically expands the repertoire of the antigen-binding sites. This diversity also significantly affect their access to and interaction with more antigen’s clefts and buried epitopes, known as cryptic antigenic regions which are not usually accessible by conventional antibodies (Desmyter et al. 1996; Stanfield et al. 2004; Stijlemans et al. 2004). Another remarkable feature of Nbs’ antigen-binding paratopes is their ability to adopt flat, concave and convex configurations which easily favors their use against folded proteins and recessed epitopes (Chaikuad et al. 2014; Custódio et al. 2020; De Genst et al. 2006; Henry and MacKenzie 2018; Muyldermans and Smider 2016). Furthermore, it is assumed that sdAbs can access holed sites on membrane proteins including ion channels and G protein-coupled receptors (Henry and MacKenzie 2018; Wei et al. 2011).
With regard to their autonomous behavior, Nbs serve as effective building blocks for multi-domain compositions, such as bivalent or multivalent to improve affinity, or bispecific to cross-link independent antigens (Muyldermans 2021a). Since VHHs are monomeric in nature, they do not cluster in multimers like scFv molecules. Furthermore, considering their high solubility and stability, Nbs can be easily fused to each other without the mispairing and solubility challenges that face the scFv dimers and multimers (Bannas et al. 2017). Moreover, varying the valency of the Nb domains that target tumors can strengthen the cell-killing and downregulation effect on certain tumor cells (Bannas et al. 2017; Oliveira et al. 2010; Sadeghnezhad et al. 2019). To achieve this, linkers can be used to create multivalent or multispecific configurations of Nbs. Also, fusion with albumin or short peptide tags can be used to extend the half-life or facilitate their purification and detection (Bannas et al. 2017; Beirnaert et al. 2017; Zupancic et al. 2021a, b). Moreover, Nbs have been successfully fused with larger proteins called megabodies. The subsequent binding of these megabodies to smaller proteins, guided by the Nb specificity, could convert them into larger protein complexes. This allows their structural analysis by cryo-electron microscopy, which is otherwise not the best strategy for solving the structure of low-molecular weight proteins (Masiulis et al. 2019).
From the crystallization ability perspective, Nbs are easy to crystallize due to their small size. They also have several properties that aid in the crystallization of harsh proteins including;(i) the ability to block domain movement, (ii) the ability to hide mobile polysaccharides bounded proteins, and (iii) the ability to insert in clefts or between interfaces. Those properties stabilize loops or large complexes and assist in the solubilization of proteins with limited solubility or even provide beneficial crystal contacts for membrane proteins (Desmyter et al. 2015). Practically, Nbs significantly helped in the stabilization of G protein-coupled receptors in their active-state conformations (Steyaert and Kobilka 2011).
Finally, an attractive advantage of Nbs is their ability to cross the blood–brain barrier (BBB), unlike regular antibodies. This makes them unique potential diagnostic and therapeutic tools for the central nervous system (Li et al. 2012). Nbs are also showing potential as screening tools via genetic modifications that links them to fluorescent proteins and thus could be used as biosensors or to trace target antigens intracellularly in living cells (Rothbauer et al. 2006). They also present a detailed depiction of immune specificity in-display libraries and are easily adaptable to high-throughput screening (Gonzalez-Sapienza et al. 2017; Rahbarizadeh et al. 2011). Being an efficient diagnostic tool, Nbs became among the best tracers for non-invasive imaging for either positron emission tomography/computerized tomography (PET/CT) or single-photon emission computerized tomography (SPECT) imaging.
Since the original patent on Nbs expired in 2013 for Europe and 2017 for the US (Arbabi-Ghahroudi 2017), the biotechnological, academic, industrial, and therapeutics communities have been pushing to commercialize Nbs. Currently, the number of studies on unique and inventive compositions and applications of Nbs is rapidly increasing.
Applications of Nbs in infectious diseases
Among the broad spectrum of applications in which Nbs have been successfully involved, the infectious disease domain comes with a big and impactful share. The developed Nbs for this purpose could be categorized into three main groups: therapeutic/prophylactic, diagnostic, and functional and structural elucidation tools.
Nbs as therapeutic and/or prophylactic tools in infectious diseases.
This is the largest category in which multiple Nbs have been tested and evaluated against different types of pathogens. For example, many Nbs have been tested to act as neutralizing agents for viral infections including foot & mouth disease virus (FMDV) (Harmsen et al. 2008), human immunodeficiency virus (HIV-1) (Forsman et al. 2008; Lutje Hulsik et al. 2013; McCoy et al. 2012), influenza A virus (Wei et al. 2011), the Middle-East respiratory syndrome coronavirus (MERS-CoV) (Wrapp et al. 2020), poliovirus (Strauss et al. 2016), rabies virus (Terryn et al. 2016), respiratory syncytial virus (RSV) (Rossey et al. 2017), rotavirus (Maffey et al. 2016), and lately the Covid-19 causing virus (SARS-CoV-2) (Chen et al. 2021; Schoof et al. 2020; Yang et al. 2023).
Among the strategies through which Nbs were used to treat infections is to target key moieties within the pathogen to block its pathogenesis. For example, to interfere with the ability of Campylobacter to colonize the host, Nbs were tested by targeting an outer membrane protein and the flagella (Vanmarsenille et al. 2018). Other Nbs targeted the F4 fimbriae of E. coli (Harmsen et al. 2005), the Salmonella enterica FilC flagellin (Huen et al. 2019), and the Streptococcus mutans adhesin (Krüger et al. 2006). Another group of Nbs were developed to target the toxins produced by some pathogens so that they block their toxic effects on the host’s cells. This category included Nbs against the Bacillus anthracis toxin (Shali et al. 2018), the Clostridium botulinum neurotoxin (Dong et al. 2010; Mukherjee et al. 2012), the C. difficile toxin (Hussack et al. 2018), the E. coli heat-labile toxin (Harmsen et al. 2009a, b) and the Staphylococcus aureus Toxic-Shock Syndrome Toxin (TSST-1) (Adams et al. 2009). Moreover, Nbs were generated to target other virulence factors such as the type III secretion system of Pseudomonas aeruginosa accordingly blocking the transfer of toxins to the host’s cell (De Tavernier et al. 2016), the urease enzyme of Helicobacter pylori inhibiting this key enzyme for the survival of the pathogen within the host (Fouladi et al. 2019), and the internalin B (InlB) of Listeria monocytogenes blocking bacterial invasion (King et al. 2018).
In viral pathogens, Nbs targeted surface structures to block entry to the host cell such as the Ebola envelope glycoprotein (Liu et al. 2017), the hepatitis B virus envelope protein S (Serruys et al. 2009), the hepatitis C E2 glycoprotein (Tarr et al. 2013), and others. Also for the viral pathogens, Nbs targeted viral replication as in the case of the Ebola nucleoprotein (Darling et al. 2017), the HCV RNA-dependent RNA polymerase (NS5B) (Thueng-in et al. 2012), and the nucleoproteins of the influenza A (Hanke et al. 2016) and the Marburg virus (Darling et al. 2017).
Among the therapeutic applications of Nbs in infectious diseases is their use for targeted drug delivery as has been demonstrated against Herpes simplex virus 2, where Nbs against glycoprotein D conjugated to the cytotoxic domain of the P. aeruginosa exotoxin acted as immunotoxins and were very effective in killing the virus-infected cells (Geoghegan et al. 2015). Also, Nbs directed against β-lactamases such as TEM-1 and BclI successfully inhibited the enzymatic activity of these enzymes and rendered the resistant pathogen susceptible to β-lactam antibiotics (Conrath et al. 2001).
Nbs as diagnostic tools in infectious diseases.
Another area in which Nbs are used actively is in the diagnosis of infectious diseases. Many Nbs targeted against moieties in the pathogens have been considered for diagnostics purposes. For example, the type 2 NS1 protein of the Dengue virus (Fatima et al. 2014), ORF2 of the hepatitis E virus (Arce et al. 2023), HIV capsid proteins (Helma et al. 2012), and other viral targets. In addition, Nbs are also used for the diagnosis of bacterial pathogens including Acinetobacter baumannii (Rasoulinejad and Gargari 2016), Brucella spp. (Abbady et al. 2011), E. coli (Salhi et al. 2020), S. aureus (Hu et al. 2021), and Vibrio cholerae (Goldman et al. 2006).
Nbs as structural and functional elucidation tools in infectious diseases.
Another application of Nbs in infectious diseases is the use as tools to elucidate the crystal structure of a pathogen-related protein or investigate its function. To this end, multiple Nbs have been developed. For instance, Nbs against the gp120 of HIV-1 were used to elucidate both its function and structure (Chen et al. 2010), while the function of the Nef protein of the same virus was studied using another Nb (Bouchet et al. 2011). In the case of bacterial pathogens, Nbs were used for structural biology studies of the MazEF toxin/antitoxin of E. coli (Lah et al. 2003), and that of the EpsJ pseudopillin of V. vulnificus (Lam et al. 2009).
An updated comprehensive list of the diverse applications of Nbs against viral and bacterial infectious diseases is presented in Tables 1 and 2, respectively.
The majority of the reported Nbs that are listed in Table 1 are directed towards viral targets, which could be attributed to the Nbs advantages discussed earlier, especially their high accessibility and penetration capabilities. With the global concerns associated with the SARS-CoV-2 pandemic in the previous three years, there was a plethora of attempts to face this threat using Nbs platforms. Over fifteen studies, targeted engineered Nbs showed promising results in neutralizing the SARS-CoV-2 virus and suppressing mutational escape in different pre-clinical animal models. In addition, several Nbs studies have contributed to the ongoing efforts to find a cure for HIV infections using multiple approaches.
Compared to viral antigens, the application of Nbs in dealing with bacterial pathogens is still limited (Table 2). Up to date, the applications of using multiple Nbs have been directed to the neutralization of the botulinum neurotoxin. It is an attractive target for the development of monospecific antibodies owing to its extreme lethality and having the least LD50 value among known toxins. It is worth mentioning that the only currently FDA-approved treatment for botulism is an equine-driven polyclonal antibody cocktail shot (Tomic et al. 2021). Additionally, E. coli with its diverse pathogenic potentials attracted attention for the development of therapeutic Nbs either for blocking attachment (Harmsen et al. 2005) or toxin neutralization (Harmsen et al. 2009a, b).
On another front, very few attempts have been implemented in the production of anti-fungal Nbs. Most of the studies aimed at detecting food product contamination with mycotoxins, specifically the aflatoxin B1 (He et al. 2022; Salvador et al. 2022). Recently, Liu et al (2023) designated Nb-natamycin conjugates that were specific to the Aspergillus fumigatus β-glucan. A. fumigatus is known to be a common causative agent of fungal keratitis, an inflammatory eye disease affecting the cornea. These conjugates successfully attenuated the virulence of A. fumigatus, and favorably modulated the inflammatory responses in fungal keratitis (Liu et al. 2023). Earlier, the same group described another Nb that is specific to the mammalian pattern-recognition receptor for fungi dectin 1. The anti-dectin 1 Nb alleviated the clinical symptoms of fungal keratitis in a mouse model, and this was attributed to the reduced expression of inflammatory cytokines IL-1β and IL-6 (Liu et al. 2022).
Disadvantages of nanobodies
The Nbs technology has become successively incorporated in a lot of therapeutic and diagnostic applications due to its small size. However, Nb's small size accounts for its short half-life by being rapidly eliminated by kidneys. This is attributed to their low molecular weight (~ 15 kDa) which is below the renal threshold for glomerular filtration (~ 50 kDa) (Ruggiero et al. 2010). Hence, their diminutive size and thereby their short half-life accounts for some challenges or limitations for using Nbs in different therapeutic fields such as screening and in vivo diagnosis applications. One of these challenges is the high uptake and accumulation of Nbs in the kidneys while being eliminated, which in turn limits their use as in vivo imaging probes for kidney screening along with some vicinity organs like the pancreas (Schoonooghe et al. 2012). In addition, the binding capacity of some Nbs is altered after being conjugated with either fluorophore or radioactive probes, for example, the Nbs conjugated with chelators having gallium-68 (68Ga) or zirconium-89 (89Zr) for immuno-positron emission tomography (immunoPET) imaging. These radiolabeled nanobodies may exhibit different features, including affinity, size, structure, and pharmacokinetics. However, site-directed conjugation and nanobody-engineering strategies have been recently applied to demonstrate the effectiveness, reliability, and safety of their use as molecular imaging probes (Yang et al. 2022). Another challenge is the low persistence of Nbs within the bloodstream due to their rapid clearance which in turn hampers their uptake. As a result, only a negligible fraction of the administered nanobody reaches the target sites, thereby hindering their efficacy. This may account for the frequent administration of Nbs along with using higher doses to maintain their therapeutic level, however, this is not recommended for an efficient therapeutic application. Further approaches seek different strategies to prolong Nbs half-life by enhancing their accumulation and pharmacokinetics either by Nbs multimerization approach or by Nb-serum albumin conjugation approach (Jovčevska and Muyldermans 2020).
Lastly, due to the high homology between the camelid germline IgV gene repertoire and its human counterparts, with up to 95% in the case of the camelid IGHV family III and its human FR counterpart, Nbs inherently pose a low immunogenic profile, which allows for prolonged and repeated administrations of Nbs in patients (Klarenbeek et al. 2015). However, the generation of antibodies against administered Nbs is possible and can be problematic, as demonstrated in the aborted clinical trial with an anti-DR5 receptor nanobody (Papadopoulos et al. 2015). This suggests that moderate humanization of Nbs sequences may be beneficial in some cases.
Despite of these challenges and limitations, the Nb technology still shows significant advantages over the conventional antibody as an effective immunotherapy.
Conclusions
Nbs represent a very promising tool for a vast array of biomedical applications owing to their superiority in terms of small molecular size, modulative specificity, and their physico-chemical properties that allow for easier downstream processing. The majority of current Nb applications are focused on the fields of diagnostics, and structural biology, being used as structural aids for troublesome proteins. The recent surge in anti-viral development accelerated the expansion in therapeutic Nb research, with many promising candidates designed to target viral infections. Anti-bacterial and anti-fungal Nb candidates are still limited in numbers and targets, which calls for future investigation of their potential applications for this purpose. This could be especially warranted in the post-antibiotic era, where available antibiotics are failing to suppress extremely resistant microbes.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Abbady AQ, Al-Mariri A, Zarkawi M, Al-Assad A, Muyldermans S (2011) Evaluation of a nanobody phage display library constructed from a Brucella-immunised camel. Vet Immunol Immunopathol 142(1):49–56. https://doi.org/10.1016/j.vetimm.2011.04.004
Adams H, Brummelhuis W, Maassen B, van Egmond N, Khattabi ME, Detmers F, Hermans P, Braam B, Stam J, Verrips T (2009) Specific immuno capturing of the Staphylococcal superantigen toxic-shock syndrome toxin-1 in plasma. Biotechnol Bioeng 104(1):143–151. https://doi.org/10.1002/bit.22365
Alfadhli A, Romanaggi C, Barklis RL, Merutka I, Bates TA, Tafesse FG, Barklis E (2021) Capsid-specific nanobody effects on HIV-1 assembly and infectivity. Virology 562:19–28. https://doi.org/10.1016/j.virol.2021.07.001
Andersen KK, Strokappe NM, Hultberg A, Truusalu K, Smidt I, Mikelsaar R-H, Mikelsaar M, Verrips T, Hammarström L, Marcotte H (2016) Neutralization of Clostridium difficile toxin B mediated by engineered lactobacilli that produce single-domain antibodies. Infect Immun 84(2):395–406. https://doi.org/10.1128/IAI.00870-15
Arbabi-Ghahroudi M (2017) Camelid single-domain antibodies: historical perspective and future outlook. Front Immunol 8:1589–1589. https://doi.org/10.3389/fimmu.2017.01589
Arbabi Ghahroudi M, Desmyter A, Wyns L, Hamers R, Muyldermans S (1997) Selection and identification of single domain antibody fragments from camel heavy-chain antibodies. FEBS Lett 414(3):521–526. https://doi.org/10.1016/s0014-5793(97)01062-4
Arce LP, Pavan MF, Bok M, Gutiérrez SE, Estein SM, Santos AT, Condorí WE, Uhart MM, Parreño V, Vizoso-Pinto MG (2023) A multispecies competitive nanobody-based ELISA for the detection of antibodies against hepatitis E virus. Sci Rep 13(1):15448. https://doi.org/10.1038/s41598-023-41955-z
Asaadi Y, Jouneghani FF, Janani S, Rahbarizadeh F (2021) A comprehensive comparison between camelid nanobodies and single chain variable fragments. Biomark Res 9:1–20. https://doi.org/10.1186/s40364-021-00332-6
Ashour J, Schmidt FI, Hanke L, Cragnolini J, Cavallari M, Altenburg A, Brewer R, Ingram J, Shoemaker C, Ploegh HL (2015) Intracellular expression of camelid single-domain antibodies specific for influenza virus nucleoprotein uncovers distinct features of its nuclear localization. J Virol 89(5):2792–2800. https://doi.org/10.1128/JVI.02693-14
Bannas P, Hambach J, Koch-Nolte F (2017) Nanobodies and nanobody-based human heavy chain antibodies as antitumor therapeutics. Front Immunol. https://doi.org/10.3389/fimmu.2017.01603
Bates A, Power CA (2019) David vs. Goliath: the structure, function, and clinical prospects of antibody fragments. Antibodies (Basel, Switzerland) 8(2):28. https://doi.org/10.3390/antib8020028
Beekwilder J, Houwelingen Av, Beckhoven JRCMV, Speksnijder AGCL (2007) Stable recombinant alpaca antibodies for detection of Tulip virus X. Eur J Plant Pathol 121:477–485
Beirnaert E, Desmyter A, Spinelli S, Lauwereys M, Aarden L, Dreier T, Loris R, Silence K, Pollet C, Cambillau C, de Haard H (2017) Bivalent llama single-domain antibody fragments against tumor necrosis factor have picomolar potencies due to intramolecular interactions. Front Immunol. https://doi.org/10.3389/fimmu.2017.00867
Boons E, Li G, Vanstreels E, Vercruysse T, Pannecouque C, Vandamme AM, Daelemans D (2014) A stably expressed llama single-domain intrabody targeting Rev displays broad-spectrum anti-HIV activity. Antiviral Res 112:91–102. https://doi.org/10.1016/j.antiviral.2014.10.007
Bouchet J, Basmaciogullari SE, Chrobak P, Stolp B, Bouchard N, Fackler OT, Chames P, Jolicoeur P, Benichou S, Baty D (2011) Inhibition of the Nef regulatory protein of HIV-1 by a single-domain antibody. Blood 117(13):3559–3568. https://doi.org/10.1182/blood-2010-07-296749
Buss NA, Henderson SJ, McFarlane M, Shenton JM, De Haan L (2012) Monoclonal antibody therapeutics: history and future. Curr Opin Pharmacol 12(5):615–622
Cao Z, Yin D, Zhang L, Ma S, Zhang K, Yang R, Shan H, Qin Z (2023) A novel blocking enzyme-linked immunosorbent assay based on a biotinylated nanobody for the rapid and sensitive clinical detection of classical swine fever virus antibodies. Microbiol Spectrum 11(1):e02996-e2922. https://doi.org/10.1128/spectrum.02996-22
Cardoso FM, Ibañez LI, Van den Hoecke S, De Baets S, Smet A, Roose K, Schepens B, Descamps FJ, Fiers W, Muyldermans S, Depicker A, Saelens X (2014) Single-domain antibodies targeting neuraminidase protect against an H5N1 influenza virus challenge. J Virol 88(15):8278–8296. https://doi.org/10.1128/JVI.03178-13
Chaikuad A, Keates T, Vincke C, Kaufholz M, Zenn M, Zimmermann B, Gutiérrez C, Zhang R-g, Hatzos-Skintges C, Joachimiak A, Muyldermans S, Herberg FW, Knapp S, Müller S (2014) Structure of cyclin G-associated kinase (GAK) trapped in different conformations using nanobodies. Biochem J 459(1):59–69. https://doi.org/10.1042/bj20131399
Chen Q, Zhou Y, Yu J, Liu W, Li F, Xian M, Nian R, Song H, Feng D (2019) An efficient constitutive expression system for Anti-CEACAM5 nanobody production in the yeast Pichia pastoris. Protein Expr Purif 155:43–47. https://doi.org/10.1016/j.pep.2018.11.001
Chen W, Xiao X, Wang Y, Zhu Z, Dimitrov DS (2010) Bifunctional fusion proteins of the human engineered antibody domain m36 with human soluble CD4 are potent inhibitors of diverse HIV-1 isolates. Antiviral Res 88(1):107–115. https://doi.org/10.1016/j.antiviral.2010.08.004
Chen W, Zhu Z, Feng Y, Dimitrov DS (2008) Human domain antibodies to conserved sterically restricted regions on gp120 as exceptionally potent cross-reactive HIV-1 neutralizers. Proc Natl Acad Sci USA 105(44):17121–17126. https://doi.org/10.1073/pnas.0805297105
Chen X, Gentili M, Hacohen N, Regev A (2021) A cell-free nanobody engineering platform rapidly generates SARS-CoV-2 neutralizing nanobodies. Nat Commun 12(1):5506–5506. https://doi.org/10.1038/s41467-021-25777-z
Conrath KE, Lauwereys M, Galleni M, Matagne A, Frère JM, Kinne J, Wyns L, Muyldermans S (2001) Beta-lactamase inhibitors derived from single-domain antibody fragments elicited in the camelidae. Antimicrob Agents Chemother 45(10):2807–2812. https://doi.org/10.1128/AAC.45.10.2807-2812.2001
Conway JO, Sherwood LJ, Collazo MT, Garza JA, Hayhurst A (2010) Llama single domain antibodies specific for the 7 botulinum neurotoxin serotypes as heptaplex immunoreagents. PLoS ONE 5(1):e8818. https://doi.org/10.1371/journal.pone.0008818
Cortez-Retamozo V, Backmann N, Senter PD, Wernery U, De Baetselier P, Muyldermans S, Revets H (2004) Efficient cancer therapy with a nanobody-based conjugate. Cancer Res 64(8):2853–2857. https://doi.org/10.1158/0008-5472.can-03-3935
Custódio TF, Das H, Sheward DJ, Hanke L, Pazicky S, Pieprzyk J, Sorgenfrei M, Schroer MA, Gruzinov AY, Jeffries CM, Graewert MA (2020) Selection, biophysical and structural analysis of synthetic nanobodies that effectively neutralize SARS-CoV-2. Nat Commun 11(1):5588
Darling TL, Sherwood LJ, Hayhurst A (2017) Intracellular crosslinking of filoviral nucleoproteins with xintrabodies restricts viral packaging. Front Immunol 8:1197–1197. https://doi.org/10.3389/fimmu.2017.01197
De Genst E, Silence K, Decanniere K, Conrath K, Loris R, Kinne J, Muyldermans S, Wyns L (2006) Molecular basis for the preferential cleft recognition by dromedary heavy-chain antibodies. Proc Natl Acad Sci U S A 103(12):4586–4591. https://doi.org/10.1073/pnas.0505379103
de Marco A (2020) Recombinant expression of nanobodies and nanobody-derived immunoreagents. Protein Expr Purif 172:105645–105645. https://doi.org/10.1016/j.pep.2020.105645
De Tavernier E, Detalle L, Morizzo E, Roobrouck A, De Taeye S, Rieger M, Verhaeghe T, Correia A, Van Hegelsom R, Figueirido R (2016) High throughput combinatorial formatting of PcrV nanobodies for efficient potency improvement. J Biol Chem 291(29):15243–15255. https://doi.org/10.1074/jbc.M115.684241
De Vos J, Devoogdt N, Lahoutte T, Muyldermans S (2013) Camelid single-domain antibody-fragment engineering for (pre) clinical in vivo molecular imaging applications: adjusting the bullet to its target. Expert Opin Biol Ther 13(8):1149–1160. https://doi.org/10.1517/14712598.2013.800478
Deffar K, Shi H, Li L, Wang X, Zhu A (2009) Nanobodies—the new concept in antibody engineering. Afr J Biotechnol 8:2645–2652
Dekker S, Toussaint W, Panayotou G, de Wit T, Visser P, Grosveld F, Drabek D (2003) Intracellularly expressed single-domain antibody against p15 matrix protein prevents the production of porcine retroviruses. J Virol 77(22):12132–12139. https://doi.org/10.1128/jvi.77.22.12132-12139.2003
Desmyter A, Spinelli S, Payan F, Lauwereys M, Wyns L, Muyldermans S, Cambillau C (2002) Three Camelid VHH domains in complex with porcine pancreatic α-amylase: inhibition and versatility of binding topology. J Biol Chem 277(26):23645–23650. https://doi.org/10.1074/jbc.M202327200
Desmyter A, Spinelli S, Roussel A, Cambillau C (2015) Camelid nanobodies: killing two birds with one stone. Curr Opin Struct Biol 32:1–8. https://doi.org/10.1016/j.sbi.2015.01.001
Desmyter A, Transue TR, Ghahroudi MA, Thi MH, Poortmans F, Hamers R, Muyldermans S, Wyns L (1996) Crystal structure of a camel single-domain VH antibody fragment in complex with lysozyme. Nat Struct Biol 3(9):803–811. https://doi.org/10.1038/nsb0996-803
Detalle L, Stohr T, Palomo C, Piedra PA, Gilbert BE, Mas V, Millar A, Power UF, Stortelers C, Allosery K, Melero JA, Depla E (2015) Generation and characterization of ALX-0171, a potent novel therapeutic nanobody for the treatment of respiratory syncytial virus infection. Antimicrob Agents Chemother 60(1):6–13. https://doi.org/10.1128/AAC.01802-15
Dolk E, van der Vaart M, Lutje Hulsik D, Vriend G, de Haard H, Spinelli S, Cambillau C, Frenken L, Verrips T (2005) Isolation of llama antibody fragments for prevention of dandruff by phage display in shampoo. Appl Environ Microbiol 71(1):442–450. https://doi.org/10.1128/AEM.71.1.442-450.2005
Dong J, Thompson AA, Fan Y, Lou J, Conrad F, Ho M, Pires-Alves M, Wilson BA, Stevens RC, Marks JD (2010) A single-domain llama antibody potently inhibits the enzymatic activity of botulinum neurotoxin by binding to the non-catalytic alpha-exosite binding region. J Mol Biol 397(4):1106–1118. https://doi.org/10.1016/j.jmb.2010.01.070
Duan H, Chen X, Zhang Z, Zhang Z, Li Z, Wang X, Zhao J, Nan Y, Liu B, Zhang A (2023) A nanobody inhibiting porcine reproductive and respiratory syndrome virus replication via blocking self-interaction of viral nucleocapsid protein. J Virol. https://doi.org/10.1128/jvi.01319-23
Duggan S (2018) Caplacizumab: first global approval. Drugs 78:1639–1642
El Khattabi M, Adams H, Heezius E, Hermans P, Detmers F, Maassen B, van der Ley P, Tommassen J, Verrips T, Stam J (2006) Llama single-chain antibody that blocks lipopolysaccharide binding and signaling: prospects for therapeutic applications. Clin Vaccine Immunol 13(10):1079–1086. https://doi.org/10.1128/CVI.00107-06
English H, Hong J, Ho M (2020) Ancient species offers contemporary therapeutics: an update on shark V(NAR) single domain antibody sequences, phage libraries and potential clinical applications. Antibody Ther 3(1):1–9. https://doi.org/10.1093/abt/tbaa001
Fatima A, Wang H, Kang K, Xia L, Wang Y, Ye W, Wang J, Wang X (2014) Development of VHH antibodies against dengue virus type 2 NS1 and comparison with monoclonal antibodies for use in immunological diagnosis. PLoS ONE 9:e95263
Feige MJ, Gräwert MA, Marcinowski M, Hennig J, Behnke J, Ausländer D, Herold EM, Peschek J, Castro CD, Flajnik M, Hendershot LM, Sattler M, Groll M, Buchner J (2014) The structural analysis of shark IgNAR antibodies reveals evolutionary principles of immunoglobulins. Proc Natl Acad Sci USA 111(22):8155–8160. https://doi.org/10.1073/pnas.1321502111
Feng M, Bian H, Wu X, Fu T, Fu Y, Hong J, Fleming BD, Flajnik MF, Ho M (2019) Construction and next-generation sequencing analysis of a large phage-displayed V(NAR) single-domain antibody library from six naïve nurse sharks. Antibody Therapeutics 2(1):1–11. https://doi.org/10.1093/abt/tby011
Forsman A, Beirnaert E, Aasa-Chapman MM, Hoorelbeke B, Hijazi K, Koh W, Tack V, Szynol A, Kelly C, McKnight A, Verrips T, de Haard H, Weiss RA (2008) Llama antibody fragments with cross-subtype human immunodeficiency virus type 1 (HIV-1)-neutralizing properties and high affinity for HIV-1 gp120. J Virol 82(24):12069–12081. https://doi.org/10.1128/jvi.01379-08
Fouladi M, Sarhadi S, Tohidkia M, Fahimi F, Samadi N, Sadeghi J, Barar J, Omidi Y (2019) Selection of a fully human single domain antibody specific to Helicobacter pylori urease. Appl Microbiol Biotechnol 103:3407–3420
Gai J, Ma L, Li G, Zhu M, Qiao P, Li X, Zhang H, Zhang Y, Chen Y, Ji W, Zhang H, Cao H, Li X, Gong R, Wan Y (2021) A potent neutralizing nanobody against SARS-CoV-2 with inhaled delivery potential. MedComm 2(1):101–113. https://doi.org/10.1002/mco2.60
Garaicoechea L, Olichon A, Marcoppido G, Wigdorovitz A, Mozgovoj M, Saif L, Surrey T, Parreño V (2008) Llama-derived single-chain antibody fragments directed to rotavirus VP6 protein possess broad neutralizing activity in vitro and confer protection against diarrhea in mice. J Virol 82(19):9753–9764. https://doi.org/10.1128/jvi.00436-08
Geoghegan EM, Zhang H, Desai PJ, Biragyn A, Markham RB (2015) Antiviral activity of a single-domain antibody immunotoxin binding to glycoprotein D of herpes simplex virus 2. Antimicrob Agents Chemother 59(1):527–535. https://doi.org/10.1128/AAC.03818-14
Goldman ER, Anderson GP, Conway J, Sherwood LJ, Fech M, Vo B, Liu JL, Hayhurst A (2008) Thermostable llama single domain antibodies for detection of botulinum a neurotoxin complex. Anal Chem 80(22):8583–8591. https://doi.org/10.1021/ac8014774
Goldman ER, Anderson GP, Liu JL, Delehanty JB, Sherwood LJ, Osborn LE, Cummins LB, Hayhurst A (2006) Facile generation of heat-stable antiviral and antitoxin single domain antibodies from a semisynthetic llama library. Anal Chem 78(24):8245–8255. https://doi.org/10.1021/ac0610053
Gómez-Sebastián S, Nuñez MC, Garaicoechea L, Alvarado C, Mozgovoj M, Lasa R, Kahl A, Wigdorovitz A, Parreño V, Escribano JM (2012) Rotavirus A-specific single-domain antibodies produced in baculovirus-infected insect larvae are protective in vivo. BMC Biotechnol 12:59–59. https://doi.org/10.1186/1472-6750-12-59
Gonzalez-Sapienza G, Rossotti MA, Tabares-da Rosa S (2017) Single-domain antibodies as versatile affinity reagents for analytical and diagnostic applications. Front Immunol 8:977. https://doi.org/10.3389/fimmu.2017.00977
Gray AC, Sidhu SS, Chandrasekera PC, Hendriksen CFM, Borrebaeck CAK (2016) Animal-friendly affinity reagents: replacing the needless in the haystack. Trends Biotechnol 34(12):960–969. https://doi.org/10.1016/j.tibtech.2016.05.017
Gray ER, Brookes JC, Caillat C, Turbé V, Webb BLJ, Granger LA, Miller BS, McCoy LE, El Khattabi M, Verrips CT, Weiss RA, Duffy DM, Weissenhorn W, McKendry RA (2017) Unravelling the molecular basis of high affinity nanobodies against HIV p24: in vitro functional, structural, and in silico insights. ACS Infect Dis 3(7):479–491. https://doi.org/10.1021/acsinfecdis.6b00189
Greenberg AS, Avila D, Hughes M, Hughes A, McKinney EC, Flajnik MF (1995) A new antigen receptor gene family that undergoes rearrangement and extensive somatic diversification in sharks. Nature 374(6518):168–173
Hamers-Casterman C, Atarhouch T, Muyldermans S, Robinson G, Hamers C, Songa EB, Bendahman N, Hamers R (1993) Naturally occurring antibodies devoid of light chains. Nature 363(6428):446–448. https://doi.org/10.1038/363446a0
Hanke L, Knockenhauer KE, Brewer RC, van Diest E, Schmidt FI, Schwartz TU, Ploegh HL (2016) The antiviral mechanism of an influenza A virus nucleoprotein-specific single-domain antibody fragment. mBio 7(6):e01569–e01516. https://doi.org/10.1128/mBio.01569-16
Hanke L, Vidakovics Perez L, Sheward DJ, Das H, Schulte T, Moliner-Morro A, Corcoran M, Achour A, Karlsson Hedestam GB, Hällberg BM, Murrell B, McInerney GM (2020) An alpaca nanobody neutralizes SARS-CoV-2 by blocking receptor interaction. Nat Commun 11(1):4420–4420. https://doi.org/10.1038/s41467-020-18174-5
Harmsen MM, Blokker JC, Pritz-Verschuren SB, Bartelink W, Van der Burg H, Koch G (2013) isolation of panels of llama single-domain antibody fragments binding all nine neuraminidase subtypes of influenza A virus. Antibodies 2(2):168–192
Harmsen MM, Fijten HP, Dekker A, Eblé PL (2008) Passive immunization of pigs with bispecific llama single-domain antibody fragments against foot-and-mouth disease and porcine immunoglobulin. Vet Microbiol 132(1–2):56–64. https://doi.org/10.1016/j.vetmic.2008.04.030
Harmsen MM, Fijten HP, Engel B, Dekker A, Eblé PL (2009a) Passive immunization with llama single-domain antibody fragments reduces foot-and-mouth disease transmission between pigs. Vaccine 27(13):1904–1911. https://doi.org/10.1016/j.vaccine.2009.01.110
Harmsen MM, van Solt CB, Fijten HPD (2009b) Enhancement of toxin- and virus-neutralizing capacity of single-domain antibody fragments by N-glycosylation. Appl Microbiol Biotechnol 84(6):1087–1094. https://doi.org/10.1007/s00253-009-2029-1
Harmsen MM, van Solt CB, Fijten HPD, van Keulen L, Rosalia RA, Weerdmeester K, Cornelissen AHM, De Bruin MGM, Eblé PL, Dekker A (2007) Passive immunization of guinea pigs with llama single-domain antibody fragments against foot-and-mouth disease. Vet Microbiol 120(3):193–206. https://doi.org/10.1016/j.vetmic.2006.10.029
Harmsen MM, van Solt CB, Hoogendoorn A, van Zijderveld FG, Niewold TA, van der Meulen J (2005) Escherichia coli F4 fimbriae specific llama single-domain antibody fragments effectively inhibit bacterial adhesion in vitro but poorly protect against diarrhoea. Vet Microbiol 111(1–2):89–98. https://doi.org/10.1016/j.vetmic.2005.09.005
He T, Nie Y, Yan T, Zhu J, He X, Li Y, Zhang Q, Tang X, Hu R, Yang Y (2022) Enhancing the detection sensitivity of nanobody against aflatoxin B1 through structure-guided modification. Int J Biol Macromol 194:188–197. https://doi.org/10.1016/j.ijbiomac.2021.11.182
Helma J, Schmidthals K, Lux V, Nüske S, Scholz AM, Kräusslich H-G, Rothbauer U, Leonhardt H (2012) Direct and dynamic detection of HIV-1 in living cells. PLoS ONE 7(11):e50026. https://doi.org/10.1371/journal.pone.0050026
Henry KA, MacKenzie CR (2018) Antigen recognition by single-domain antibodies: structural latitudes and constraints. Mabs 10(6):815–826. https://doi.org/10.1080/19420862.2018.1489633
Hinz A, Lutje Hulsik D, Forsman A, Koh WW-L, Belrhali H, Gorlani A, de Haard H, Weiss RA, Verrips T, Weissenhorn W (2010) Crystal structure of the neutralizing Llama V(HH) D7 and its mode of HIV-1 gp120 interaction. PLoS ONE 5(5):e10482–e10482. https://doi.org/10.1371/journal.pone.0010482
Hu Y, Sun Y, Gu J, Yang F, Wu S, Zhang C, Ji X, Lv H, Muyldermans S, Wang S (2021) Selection of specific nanobodies to develop an immuno-assay detecting Staphylococcus aureus in milk. Food Chem 353:129481
Huen J, Yan Z, Iwashkiw J, Dubey S, Gimenez MC, Ortiz ME, Patel SV, Jones MD, Riazi A, Terebiznik M, Babaei S, Shahinas D (2019) A novel single domain antibody targeting FliC flagellin of Salmonella enterica for effective inhibition of host cell invasion. Front Microbiol. https://doi.org/10.3389/fmicb.2019.02665
Hultberg A, Temperton NJ, Rosseels V, Koenders M, Gonzalez-Pajuelo M, Schepens B, Ibañez LI, Vanlandschoot P, Schillemans J, Saunders M, Weiss RA, Saelens X, Melero JA, Verrips CT, Van Gucht S, de Haard HJ (2011) Llama-derived single domain antibodies to build multivalent, superpotent and broadened neutralizing anti-viral molecules. PLoS ONE 6(4):e17665. https://doi.org/10.1371/journal.pone.0017665
Huo J, Mikolajek H, Le Bas A, Clark JJ, Sharma P, Kipar A, Dormon J, Norman C, Weckener M, Clare DK, Harrison PJ, Tree JA, Buttigieg KR, Salguero FJ, Watson R, Knott D, Carnell O, Ngabo D, Elmore MJ, Fotheringham S, Harding A, Moynié L, Ward PN, Dumoux M, Prince T, Hall Y, Hiscox JA, Owen A, James W, Carroll MW, Stewart JP, Naismith JH, Owens RJ (2021) A potent SARS-CoV-2 neutralising nanobody shows therapeutic efficacy in the Syrian golden hamster model of COVID-19. Nat Commun 12(1):5469–5469. https://doi.org/10.1038/s41467-021-25480-z
Hussack G, Ryan S, Van Faassen H, Rossotti M, MacKenzie CR, Tanha J (2018) Neutralization of Clostridium difficile toxin B with VHH-Fc fusions targeting the delivery and CROPs domains. PLoS ONE 13(12):e0208978. https://doi.org/10.1371/journal.pone.0208978
Ibañez LI, De Filette M, Hultberg A, Verrips T, Temperton N, Weiss RA, Vandevelde W, Schepens B, Vanlandschoot P, Saelens X (2011) Nanobodies with in vitro neutralizing activity protect mice against H5N1 influenza virus infection. J Infect Dis 203(8):1063–1072. https://doi.org/10.1093/infdis/jiq168
Ingram JR, Schmidt FI, Ploegh HL (2018) Exploiting nanobodies’ singular traits. Annu Rev Immunol 36:695–715. https://doi.org/10.1146/annurev-immunol-042617-053327
Jähnichen S, Blanchetot C, Maussang D, Gonzalez-Pajuelo M, Chow KY, Bosch L, De Vrieze S, Serruys B, Ulrichts H, Vandevelde W, Saunders M, De Haard HJ, Schols D, Leurs R, Vanlandschoot P, Verrips T, Smit MJ (2010) CXCR4 nanobodies (VHH-based single variable domains) potently inhibit chemotaxis and HIV-1 replication and mobilize stem cells. Proc Natl Acad Sci U S A 107(47):20565–20570. https://doi.org/10.1073/pnas.1012865107
Jin H, Tang X, Li L, Chen Y, Zhu Y, Chong H, He Y (2021) Generation of HIV-resistant cells with a single-domain antibody: implications for HIV-1 gene therapy. Cell Mol Immunol 18(3):660–674. https://doi.org/10.1038/s41423-020-00627-y
Jittavisutthikul S, Thanongsaksrikul J, Thueng-In K, Chulanetra M, Srimanote P, Seesuay W, Malik, AAChaicumpa, W. (2015) Humanized-VHH transbodies that inhibit HCV protease and replication. Viruses 7(4):2030–2056. https://doi.org/10.3390/v7042030
Jovčevska I, Muyldermans S (2020) The Therapeutic potential of nanobodies. BioDrugs 34(1):11–26. https://doi.org/10.1007/s40259-019-00392-z
Keyaerts M, Xavier C, Heemskerk J, Devoogdt N, Everaert H, Ackaert C, Vanhoeij M, Duhoux FP, Gevaert T, Simon P (2016) Phase I study of 68Ga-HER2-nanobody for PET/CT assessment of HER2 expression in breast carcinoma. J Nucl Med 57(1):27–33
Kijanka MM, van Brussel AS, van der Wall E, Mali WP, van Diest PJ, van Bergen en Henegouwen PM, Oliveira S (2016) Optical imaging of pre-invasive breast cancer with a combination of VHHs targeting CAIX and HER2 increases contrast and facilitates tumour characterization. Ejnmmi Res 6:1–13
King MT, Huh I, Shenai A, Brooks TM, Brooks CL (2018) Structural basis of VHH-mediated neutralization of the food-borne pathogen Listeria monocytogenes. J Biol Chem 293(35):13626–13635. https://doi.org/10.1074/jbc.RA118.003888
Klarenbeek A, Mazouari KE, Desmyter A, Blanchetot C, Hultberg A, De Jonge N, Roovers RC, Cambillau C, Spinelli S, Del-Favero J (2015) Camelid Ig V genes reveal significant human homology not seen in therapeutic target genes, providing for a powerful therapeutic antibody platform. Mabs 7(4):693–706. https://doi.org/10.1080/19420862.2015.1046648
Koenig P-A, Das H, Liu H, Kümmerer BM, Gohr FN, Jenster L-M, Schiffelers LDJ, Tesfamariam YM, Uchima M, Wuerth JD, Gatterdam K, Ruetalo N, Christensen MH, Fandrey CI, Normann S, Tödtmann JMP, Pritzl S, Hanke L, Boos J, Yuan M, Zhu X, Schmid-Burgk JL, Kato H, Schindler M, Wilson IA, Geyer M, Ludwig KU, Hällberg BM, Wu NC, Schmidt FI (2021) Structure-guided multivalent nanobodies block SARS-CoV-2 infection and suppress mutational escape. Science (New York, NY) 371(6530):eabe6230
Koh WWL, Steffensen S, Gonzalez-Pajuelo M, Hoorelbeke B, Gorlani A, Szynol A, Forsman A, Aasa-Chapman MMI, de Haard H, Verrips T, Weiss RA (2010) Generation of a family-specific phage library of llama single chain antibody fragments that neutralize HIV-1. J Biol Chem 285(25):19116–19124. https://doi.org/10.1074/jbc.M110.116699
Könning D, Rhiel L, Empting M (2017) Semi-synthetic vNAR libraries screened against therapeutic antibodies primarily deliver anti-idiotypic binders. Sci Rep 7(1):9676. https://doi.org/10.1038/s41598-017-10513-9
Koromyslova AD, Hansman GS (2015) Nanobody binding to a conserved epitope promotes norovirus particle disassembly. J Virol 89(5):2718–2730. https://doi.org/10.1128/JVI.03176-14
Koromyslova AD, Hansman GS (2017) Nanobodies targeting norovirus capsid reveal functional epitopes and potential mechanisms of neutralization. PLoS Pathog 13(11):e1006636–e1006636. https://doi.org/10.1371/journal.ppat.1006636
Krüger C, Hultberg A, Marcotte H, Hermans P, Bezemer S, Frenken LGJ, Hammarström L (2006) Therapeutic effect of llama derived VHH fragments against Streptococcus mutans on the development of dental caries. Appl Microbiol Biotechnol 72(4):732–737. https://doi.org/10.1007/s00253-006-0347-0
Lah J, Marianovsky I, Glaser G, Engelberg-Kulka H, Kinne J, Wyns L, Loris R (2003) Recognition of the intrinsically flexible addiction antidote MazE by a dromedary single domain antibody fragment: STRUCTURE, THERMODYNAMICS OF BINDING, STABILITY, AND INFLUENCE ON INTERACTIONS WITH DNA *. J Biol Chem 278(16):14101–14111. https://doi.org/10.1074/jbc.M209855200
Lam AY, Pardon E, Korotkov KV, Hol WGJ, Steyaert J (2009) Nanobody-aided structure determination of the EpsI:EpsJ pseudopilin heterodimer from Vibrio vulnificus. J Struct Biol 166(1):8–15. https://doi.org/10.1016/j.jsb.2008.11.008
Li T, Bourgeois J-P, Celli S, Glacial F, Le Sourd A-M, Mecheri S, Weksler B, Romero I, Couraud P-O, Rougeon F, Lafaye P (2012) Cell-penetrating anti-GFAP VHH and corresponding fluorescent fusion protein VHH-GFP spontaneously cross the blood-brain barrier and specifically recognize astrocytes: application to brain imaging. FASEB J 26(10):3969–3979. https://doi.org/10.1096/fj.11-201384
Li T, Cai H, Yao H, Zhou B, Zhang N, Fentener van Vlissingen M, Kuiken T, Han W, Geurtsvankessel C, Gong Y, Zhao Y, Shen Q, Qin W, Tian X-X, Peng C, Lai Y, Wang Y, Hutter C, Kuo S-M, Li D (2021) A synthetic nanobody targeting RBD protects hamsters from SARS-CoV-2 infection. Nat Commun. https://doi.org/10.1038/s41467-021-24905-z
Liu H, Liang C, Duan H, Zhang X, Wang X, Xiao S, Zhou EM (2016) Intracellularly expressed nanobodies against non-structural protein 4 of porcine reproductive and respiratory syndrome virus inhibit virus replication. Biotechnol Lett 38(7):1081–1088. https://doi.org/10.1007/s10529-016-2086-3
Liu H, Wang Y, Duan H, Zhang A, Liang C, Gao J, Zhang C, Huang B, Li Q, Li N, Xiao S, Zhou E-M (2015) An intracellularly expressed Nsp9-specific nanobody in MARC-145 cells inhibits porcine reproductive and respiratory syndrome virus replication. Vet Microbiol 181(3):252–260. https://doi.org/10.1016/j.vetmic.2015.10.021
Liu JL, Shriver-Lake LC, Anderson GP, Zabetakis D, Goldman ER (2017) Selection, characterization, and thermal stabilization of llama single domain antibodies towards Ebola virus glycoprotein. Microb Cell Fact 16(1):223–223. https://doi.org/10.1186/s12934-017-0837-z
Liu X, Sui J, Li C, Peng X, Wang Q, Jiang N, Xu Q, Wang L, Lin J, Zhao G (2022) Preparation of a nanobody specific to dectin 1 and its anti-inflammatory effects on fungal keratitis. Int J Nanomed. https://doi.org/10.2147/IJN.S338974
Liu X, Sui J, Li C, Wang Q, Peng X, Meng F, Xu Q, Jiang N, Zhao G, Lin J (2023) The preparation and therapeutic effects of β-glucan-specific nanobodies and nanobody-natamycin conjugates in fungal keratitis. Acta Biomater 169:398–409. https://doi.org/10.1016/j.actbio.2023.08.019
Liu Z-H, Lei K-X, Han G-W, Xu H-L, He F (2020) Novel lentivirus-based method for rapid selection of inhibitory nanobody against PRRSV. Viruses 12(2):229. https://doi.org/10.3390/v12020229
Lu Q, Li X, Zhao J, Zhu J, Luo Y, Duan H, Ji P, Wang K, Liu B, Wang X, Fan W, Sun Y, Zhou E-M, Zhao Q (2020) Nanobody-horseradish peroxidase and -EGFP fusions as reagents to detect porcine parvovirus in the immunoassays. J Nanobiotechnol 18(1):7–7. https://doi.org/10.1186/s12951-019-0568-x
Lutje Hulsik D, Liu Y-y, Strokappe NM, Battella S, El Khattabi M, McCoy LE, Sabin C, Hinz A, Hock M, Macheboeuf P, Bonvin AMJJ, Langedijk JPM, Davis D, Forsman Quigley A, Aasa-Chapman MMI, Seaman MS, Ramos A, Poignard P, Favier A, Simorre J-P, Weiss RA, Verrips CT, Weissenhorn W, Rutten L (2013) A gp41 MPER-specific llama VHH requires a hydrophobic CDR3 for neutralization but not for antigen recognition. PLoS Pathog 9(3):e1003202–e1003202. https://doi.org/10.1371/journal.ppat.1003202
Maffey L, Vega CG, Miño S, Garaicoechea L, Parreño V (2016) Anti-VP6 VHH: an experimental treatment for rotavirus A-associated disease. PLoS ONE 11(9):e0162351–e0162351. https://doi.org/10.1371/journal.pone.0162351
Martin F, Volpari C, Steinkuhler C, Dimasi N, Brunetti M, Biasiol G, Altamura S, Cortese R, De Francesco R, Sollazzo M (1997) Affinity selection of a camelized V(H) domain antibody inhibitor of hepatitis C virus NS3 protease. Protein Eng 10(5):607–614. https://doi.org/10.1093/protein/10.5.607
Martín MC, Pant N, Ladero V, Günaydin G, Andersen KK, Alvarez B, Martínez N, Alvarez MA, Hammarström L, Marcotte H (2011) Integrative expression system for delivery of antibody fragments by lactobacilli. Appl Environ Microbiol 77(6):2174–2179. https://doi.org/10.1128/AEM.02690-10
Masiulis S, Desai R, Uchański T, Serna Martin I, Laverty D, Karia D, Malinauskas T, Zivanov J, Pardon E, Kotecha A, Steyaert J, Miller KW, Aricescu AR (2019) GABA(A) receptor signalling mechanisms revealed by structural pharmacology. Nature 565(7740):454–459. https://doi.org/10.1038/s41586-018-0832-5
Matz J, Hérate C, Bouchet J, Dusetti N, Gayet O, Baty D, Benichou S, Chames P (2014) Selection of intracellular single-domain antibodies targeting the HIV-1 Vpr protein by cytoplasmic yeast two-hybrid system. PLoS ONE 9(12):e113729–e113729. https://doi.org/10.1371/journal.pone.0113729
Matz J, Kessler P, Bouchet J, Combes O, Ramos OHP, Barin F, Baty D, Martin L, Benichou S, Chames P (2013) Straightforward selection of broadly neutralizing single-domain antibodies targeting the conserved CD4 and coreceptor binding sites of HIV-1 gp120. J Virol 87(2):1137–1149. https://doi.org/10.1128/JVI.00461-12
McCoy LE, Quigley AF, Strokappe NM, Bulmer-Thomas B, Seaman MS, Mortier D, Rutten L, Chander N, Edwards CJ, Ketteler R, Davis D, Verrips T, Weiss RA (2012) Potent and broad neutralization of HIV-1 by a llama antibody elicited by immunization. J Exp Med 209(6):1091–1103. https://doi.org/10.1084/jem.20112655
Mitchell LS, Colwell LJ (2018) Comparative analysis of nanobody sequence and structure data. Proteins 86(7):697–706. https://doi.org/10.1002/prot.25497
Mukherjee J, Tremblay JM, Leysath CE, Ofori K, Baldwin K, Feng X, Bedenice D, Webb RP, Wright PM, Smith LA, Tzipori S, Shoemaker CB (2012) A novel strategy for development of recombinant antitoxin therapeutics tested in a mouse botulism model. PLoS ONE 7(1):e29941–e29941. https://doi.org/10.1371/journal.pone.0029941
Müller MR, Saunders K, Grace C, Jin M, Piche-Nicholas N, Steven J, O’Dwyer R, Wu L, Khetemenee L, Vugmeyster Y, Hickling TP, Tchistiakova L, Olland S, Gill D, Jensen A, Barelle CJ (2012) Improving the pharmacokinetic properties of biologics by fusion to an anti-HSA shark VNAR domain. Mabs 4(6):673–685. https://doi.org/10.4161/mabs.22242
Muyldermans S (2013) Nanobodies: natural single-domain antibodies. Annu Rev Biochem 82:775–797. https://doi.org/10.1146/annurev-biochem-063011-092449
Muyldermans S (2021a) Applications of nanobodies. Annu Rev Anim Biosci 9(1):401–421. https://doi.org/10.1146/annurev-animal-021419-083831
Muyldermans S (2021b) A guide to: generation and design of nanobodies. FEBS J 288(7):2084–2102. https://doi.org/10.1111/febs.15515
Muyldermans S, Smider VV (2016) Distinct antibody species: structural differences creating therapeutic opportunities. Curr Opin Immunol 40:7–13. https://doi.org/10.1016/j.coi.2016.02.003
Nambulli S, Xiang Y, Tilston-Lunel NL, Rennick LJ, Sang Z, Klimstra WB, Reed DS, Crossland NA, Shi Y, Duprex WP (2021) Inhalable Nanobody (PiN-21) prevents and treats SARS-CoV-2 infections in Syrian hamsters at ultra-low doses. Sci Adv 7(22):eabh0319. https://doi.org/10.1126/sciadv.abh0319
Nguyen VK, Hamers R, Wyns L, Muyldermans S (2000) Camel heavy-chain antibodies: diverse germline VHH and specific mechanisms enlarge the antigen-binding repertoire. EMBO J 19(5):921–930
Olichon A, Schweizer D, Muyldermans S, de Marco A (2007) Heating as a rapid purification method for recovering correctly-folded thermotolerant VH and VHH domains. BMC Biotechnol 7:7–7. https://doi.org/10.1186/1472-6750-7-7
Oliveira S, Schiffelers RM, van der Veeken J, van der Meel R, Vongpromek R, van Bergen en Henegouwen PMP, Storm G, Roovers RC (2010) Downregulation of EGFR by a novel multivalent nanobody-liposome platform. J Controlled Release 145(2):165–175. https://doi.org/10.1016/j.jconrel.2010.03.020
Pant N, Hultberg A, Zhao Y, Svensson L, Pan-Hammarström Q, Johansen K, Pouwels PH, Ruggeri FM, Hermans P, Frenken L, Borén T, Marcotte H, Hammarström L (2006) Lactobacilli expressing variable domain of llama heavy-chain antibody fragments (Lactobodies) confer protection against rotavirus-induced diarrhea. J Infect Dis 194(11):1580–1588. https://doi.org/10.1086/508747
Papadopoulos KP, Isaacs R, Bilic S, Kentsch K, Huet HA, Hofmann M, Rasco D, Kundamal N, Tang Z, Cooksey J, Mahipal A (2015) Unexpected hepatotoxicity in a phase I study of TAS266, a novel tetravalent agonistic Nanobody® targeting the DR5 receptor. Cancer Chemother Pharmacol 75(5):887–895. https://doi.org/10.1007/s00280-015-2712-0
Pezeshkian N, Groves NS, van Engelenburg SB (2019) Single-molecule imaging of HIV-1 envelope glycoprotein dynamics and Gag lattice association exposes determinants responsible for virus incorporation. Proc Natl Acad Sci USA 116(50):25269–25277. https://doi.org/10.1073/pnas.1910008116
Phalaphol A, Thueng-in K, Thanongsaksrikul J, Poungpair O, Bangphoomi K, Sookrung N, Srimanote P, Chaicumpa W (2013) Humanized-VH/VHH that inhibit HCV replication by interfering with the virus helicase activity. J Virol Methods 194(1):289–299. https://doi.org/10.1016/j.jviromet.2013.08.032
Pillay TS, Muyldermans S (2021) Application of single-domain antibodies (“nanobodies”) to laboratory diagnosis. Ann Lab Med 41(6):549–558
Pleiner T, Bates M, Trakhanov S, Lee C-T, Schliep JE, Chug H, Böhning M, Stark H, Urlaub H, Görlich D (2015) Nanobodies: site-specific labeling for super-resolution imaging, rapid epitope-mapping and native protein complex isolation. eLife 4:e11349. https://doi.org/10.7554/eLife.11349
Pymm P, Adair A, Chan L-J, Cooney JP, Mordant FL, Allison CC, Lopez E, Haycroft ER, O’Neill MT, Tan LL, Dietrich MH, Drew D, Doerflinger M, Dengler MA, Scott NE, Wheatley AK, Gherardin NA, Venugopal H, Cromer D, Davenport MP, Pickering R, Godfrey DI, Purcell DFJ, Kent SJ, Chung AW, Subbarao K, Pellegrini M, Glukhova A, Tham W-H (2021) Nanobody cocktails potently neutralize SARS-CoV-2 D614G N501Y variant and protect mice. Proc Natl Acad Sci USA 118(19):e2101918118. https://doi.org/10.1073/pnas.2101918118
Rahbarizadeh F, Ahmadvand D, Sharifzadeh Z (2011) Nanobody; an Old Concept and New Vehicle for Immunotargeting. Immunol Invest 40(3):299–338. https://doi.org/10.3109/08820139.2010.542228
Rasoulinejad S, Gargari SLM (2016) Aptamer-nanobody based ELASA for specific detection of Acinetobacter baumannii isolates. J Biotechnol 231:46–54. https://doi.org/10.1016/j.jbiotec.2016.05.024
Riazi A, Strong P, Coleman R, Chen W, Hirama T, Faassen H, Henry M, Logan S, Szymanski C, Mackenzie R, Arbabi Ghahroudi M (2013) Pentavalent single-domain antibodies reduce Campylobacter jejuni motility and colonization in chickens. PLoS ONE 8:e83928. https://doi.org/10.1371/journal.pone.0083928
Rossey I, Gilman MSA, Kabeche SC, Sedeyn K, Wrapp D, Kanekiyo M, Chen M, Mas V, Spitaels J, Melero JA, Graham BS, Schepens B, McLellan JS, Saelens X (2017) Potent single-domain antibodies that arrest respiratory syncytial virus fusion protein in its prefusion state. Nat Commun 8:14158–14158. https://doi.org/10.1038/ncomms14158
Rothbauer U, Zolghadr K, Tillib S, Nowak D, Schermelleh L, Gahl A, Backmann N, Conrath K, Muyldermans S, Cardoso MC, Leonhardt H (2006) Targeting and tracing antigens in live cells with fluorescent nanobodies. Nat Methods 3(11):887–889. https://doi.org/10.1038/nmeth953
Ruggiero A, Villa CH, Bander E, Rey DA, Bergkvist M, Batt CA, Manova-Todorova K, Deen WM, Scheinberg DA, McDevitt MR (2010) Paradoxical glomerular filtration of carbon nanotubes. Proc Natl Acad Sci 107(27):12369–12374
Sabir JS, Atef A, El-Domyati FM, Edris S, Hajrah N, Alzohairy AM, Bahieldin A (2014) Construction of naïve camelids VHH repertoire in phage display-based library. C R Biol 337(4):244–249. https://doi.org/10.1016/j.crvi.2014.02.004
Sadeghnezhad G, Romão E, Bernedo-Navarro R, Massa S, Khajeh K, Muyldermans S, Hassania S (2019) Identification of new DR5 agonistic nanobodies and generation of multivalent nanobody constructs for cancer treatment. Int J Mol Sci 20(19):4818. https://doi.org/10.3390/ijms20194818
Salhi I, Bessalah S, Snoun D, Khorchani T, Hammadi M (2020) Construction of a nanobodies phage display library from an Escherichia coli immunized dromedary. Iran J Biotechnol 18(1):e2247–e2247. https://doi.org/10.30498/IJB.2020.127753.2247
Salmen W, Hu L, Bok M, Chaimongkol N, Ettayebi K, Sosnovtsev SV, Soni K, Ayyar BV, Shanker S, Neill FH (2023) A single nanobody neutralizes multiple epochally evolving human noroviruses by modulating capsid plasticity. Nat Commun 14(1):6516
Salvador J-P, Vasylieva N, Gonzalez-Garcia I, Jin M, Caster R, Siegel JB, Hammock BD (2022) Nanobody-based lateral flow immunoassay for the rapid detection of aflatoxin B1 in almond milk. ACS Food Sci Technol 2(8):1276–1282. https://doi.org/10.1021/acsfoodscitech.2c00118
Sánchez-García L, Voltà-Durán E, Parladé E, Mazzega E, Sánchez-Chardi A, Serna N, López-Laguna H, Mitstorfer M, Unzueta U, Vázquez E, Villaverde A, de Marco A (2021) Self-assembled nanobodies as selectively targeted, nanostructured, and multivalent materials. ACS Appl Mater Interfaces 13(25):29406–29415. https://doi.org/10.1021/acsami.1c08092
Schmidt FI, Hanke L, Morin B, Brewer R, Brusic V, Whelan SPJ, Ploegh HL (2016) Phenotypic lentivirus screens to identify functional single domain antibodies. Nat Microbiol 1(8):16080–16080. https://doi.org/10.1038/nmicrobiol.2016.80
Schoof M, Faust B, Saunders RA, Sangwan S, Rezelj V, Hoppe N, Boone M, Billesbølle CB, Puchades C, Azumaya CM, Kratochvil HT, Zimanyi M, Deshpande I, Liang J, Dickinson S, Nguyen HC, Chio CM, Merz GE, Thompson MC, Diwanji D, Schaefer K, Anand AA, Dobzinski N, Zha BS, Simoneau CR, Leon K, White KM, Chio US, Gupta M, Jin M, Li F, Liu Y, Zhang K, Bulkley D, Sun M, Smith AM, Rizo AN, Moss F, Brilot AF, Pourmal S, Trenker R, Pospiech T, Gupta S, Barsi-Rhyne B, Belyy V, Barile-Hill AW, Nock S, Liu Y, Krogan NJ, Ralston CY, Swaney DL, García-Sastre A, Ott M, Vignuzzi M, Consortium QSB, Walter P, Manglik A (2020) An ultrapotent synthetic nanobody neutralizes SARS-CoV-2 by stabilizing inactive Spike. Science (New York, NY) 370(6523):1473–1479. https://doi.org/10.1126/science.abe3255
Schoonooghe S, Laoui D, Van Ginderachter JA, Devoogdt N, Lahoutte T, De Baetselier P, Raes G (2012) Novel applications of nanobodies for in vivo bio-imaging of inflamed tissues in inflammatory diseases and cancer. Immunobiology 217(12):1266–1272. https://doi.org/10.1016/j.imbio.2012.07.009
Schrankel, CS, Gökirmak, T, Lee, C-W, Chang, G, Hamdoun, A. (2019). Generation, expression and utilization of single-domain antibodies for in vivo protein localization and manipulation in sea urchin embryos. In A. Hamdoun & K. R. Foltz (Eds.), Methods cell biol (Vol. 151, pp. 353–376). Academic Press. doi: https://doi.org/10.1016/bs.mcb.2018.11.005
Sela-Culang I, Kunik V, Ofran Y (2013) The structural basis of antibody-antigen recognition. Front Immunol 4:302–302. https://doi.org/10.3389/fimmu.2013.00302
Serruys B, Van Houtte F, Farhoudi-Moghadam A, Leroux-Roels G, Vanlandschoot P (2010) Production, characterization and in vitro testing of HBcAg-specific VHH intrabodies. J Gen Virol 91(Pt 3):643–652. https://doi.org/10.1099/vir.0.016063-0
Serruys B, Van Houtte F, Verbrugghe P, Leroux-Roels G, Vanlandschoot P (2009) Llama-derived single-domain intrabodies inhibit secretion of hepatitis B virions in mice. Hepatology 49(1):39–49. https://doi.org/10.1002/hep.22609
Shali A, Hasannia S, Gashtasbi F, Abdous M, Shahangian SS, Jalili S (2018) Generation and screening of efficient neutralizing single domain antibodies (VHHs) against the critical functional domain of anthrax protective antigen (PA). Int J Biol Macromol 114:1267–1278. https://doi.org/10.1016/j.ijbiomac.2018.03.034
Sherwood LJ, Hayhurst A (2013) Ebolavirus nucleoprotein C-termini potently attract single domain antibodies enabling monoclonal affinity reagent sandwich assay (MARSA) formulation. PLoS ONE 8(4):e61232–e61232. https://doi.org/10.1371/journal.pone.0061232
Sherwood LJ, Osborn RC Jr, Patterson JL, Hayhurst A (2007) Rapid assembly of sensitive antigen-capture assays for Marburg, virus, using in vitro selection of llama single-domain antibodies, at biosafety level. J Infect Dis 196(Supplement_2):S213–S219. https://doi.org/10.1086/520586
Stanfield RL, Dooley H, Flajnik MF, Wilson IA (2004) Crystal structure of a shark single-domain antibody V region in complex with lysozyme. Science 305(5691):1770–1773. https://doi.org/10.1126/science.1101148
Starzl T, Fung J (1986) Orthoclone OKT3 in treatment of allografts rejected under cyclosporine-steroid therapy. Transplant Proc
Steyaert J, Kobilka BK (2011) Nanobody stabilization of G protein-coupled receptor conformational states. Curr Opin Struct Biol 21(4):567–572. https://doi.org/10.1016/j.sbi.2011.06.011
Stijlemans B, Conrath K, Cortez-Retamozo V, Van Xong H, Wyns L, Senter P, Revets H, De Baetselier P, Muyldermans S, Magez S (2004) Efficient targeting of conserved cryptic epitopes of infectious agents by single domain antibodies: AFRICAN TRYPANOSOMES AS PARADIGM *. J Biol Chem 279(2):1256–1261. https://doi.org/10.1074/jbc.M307341200
Strauss M, Schotte L, Thys B, Filman DJ, Hogle JM (2016) Five of five VHHs neutralizing poliovirus bind the receptor-binding site. J Virol 90(7):3496–3505. https://doi.org/10.1128/JVI.03017-15
Sun T, Zhao Z, Liu W, Xu Z, He H, Ning B, Jiang Y, Gao Z (2020) Development of sandwich chemiluminescent immunoassay based on an anti-staphylococcal enterotoxin B Nanobody-Alkaline phosphatase fusion protein for detection of staphylococcal enterotoxin B. Anal Chim Acta 1108:28–36. https://doi.org/10.1016/j.aca.2020.01.032
Tang JC, Drokhlyansky E, Etemad B, Rudolph S, Guo B, Wang S, Ellis EG, Li JZ, Cepko CL (2016) Detection and manipulation of live antigen-expressing cells using conditionally stable nanobodies. eLife 5:e15312. https://doi.org/10.7554/eLife.15312
Tarr AW, Lafaye P, Meredith L, Damier-Piolle L, Urbanowicz RA, Meola A, Jestin J-L, Brown RJP, McKeating JA, Rey FA, Ball JK, Krey T (2013) An alpaca nanobody inhibits hepatitis C virus entry and cell-to-cell transmission. Hepatology 58(3):932–939. https://doi.org/10.1002/hep.26430
Terryn S, Francart A, Rommelaere H, Stortelers C, Van Gucht S (2016) Post-exposure treatment with anti-rabies VHH and vaccine significantly improves protection of mice from lethal rabies infection. PLoS Negl Trop Dis 10(8):e0004902. https://doi.org/10.1371/journal.pntd.0004902
Thanongsaksrikul J, Srimanote P, Maneewatch S, Choowongkomon K, Tapchaisri P, Makino S-I, Kurazono H, Chaicumpa W (2010) A V H H that neutralizes the zinc metalloproteinase activity of botulinum neurotoxin type A. J Biol Chem 285(13):9657–9666. https://doi.org/10.1074/jbc.M109.073163
Thueng-in K, Thanongsaksrikul J, Srimanote P, Bangphoomi K, Poungpair O, Maneewatchararangsri S, Choowongkomon K, Chaicumpa W (2012) Cell penetrable humanized-VH/VHHs that inhibit RNA dependent RNA polymerase (NS5B) of HCV. PLoS ONE 7:e49254. https://doi.org/10.1371/journal.pone.0049254
Thys B, Schotte L, Muyldermans S, Wernery U, Hassanzadeh-Ghassabeh G, Rombaut B (2010) In vitro antiviral activity of single domain antibody fragments against poliovirus. Antiviral Res 87:257–264. https://doi.org/10.1016/j.antiviral.2010.05.012
Tillib SV, Ivanova TI, Vasilev LA, Rutovskaya MV, Saakyan SA, Gribova IY, Tutykhina IL, Sedova ES, Lysenko AA, Shmarov MM, Logunov DY, Naroditsky BS, Gintsburg AL (2013) Formatted single-domain antibodies can protect mice against infection with influenza virus (H5N2). Antiviral Res 97(3):245–254. https://doi.org/10.1016/j.antiviral.2012.12.014
Tomic MT, Farr-Jones S, Syar ES, Niemuth N, Kobs D, Hackett MJ, Espinoza Y, Martinez Z, Pham K, Snow DM (2021) Neutralizing concentrations of anti-botulinum toxin antibodies positively correlate with mouse neutralization assay results in a guinea pig model. Toxins 13(9):671. https://doi.org/10.3390/toxins13090671
Tremblay JM, Kuo CL, Abeijon C, Sepulveda J, Oyler G, Hu X, Jin MM, Shoemaker CB (2010) Camelid single domain antibodies (VHHs) as neuronal cell intrabody binding agents and inhibitors of Clostridium botulinum neurotoxin (BoNT) proteases. Toxicon 56(6):990–998. https://doi.org/10.1016/j.toxicon.2010.07.003
van der Vaart JM, Pant N, Wolvers D, Bezemer S, Hermans PW, Bellamy K, Sarker SA, van der Logt CP, Svensson L, Verrips CT, Hammarstrom L, van Klinken BJ (2006) Reduction in morbidity of rotavirus induced diarrhoea in mice by yeast produced monovalent llama-derived antibody fragments. Vaccine 24(19):4130–4137. https://doi.org/10.1016/j.vaccine.2006.02.045
Van Hout A, Klarenbeek A, Bobkov V, Doijen J, Arimont M, Zhao C, Heukers R, Rimkunas R, de Graaf C, Verrips T, van der Woning B, de Haard H, Rucker JB, Vermeire K, Handel T, Van Loy T, Smit MJ, Schols D (2018) CXCR4-targeting nanobodies differentially inhibit CXCR4 function and HIV entry. Biochem Pharmacol 158:402–412. https://doi.org/10.1016/j.bcp.2018.10.015
Vanmarsenille C, Díaz Del Olmo I, Elseviers J, Hassanzadeh Ghassabeh G, Moonens K, Vertommen D, Martel A, Haesebrouck F, Pasmans F, Hernalsteens J-P, De Greve H (2017) Nanobodies targeting conserved epitopes on the major outer membrane protein of Campylobacter as potential tools for control of Campylobacter colonization. Vet Res 48(1):86–86. https://doi.org/10.1186/s13567-017-0491-9
Vanmarsenille C, Elseviers J, Yvanoff C, Hassanzadeh-Ghassabeh G, Garcia G, Martens E, Depicker A, Martel A, Haesebrouck F, Pasmans F, Hernalsteens J-P, De Greve H (2018) In planta expression of nanobody-based designer chicken antibodies targeting Campylobacter. PLoS ONE 13:e0204222. https://doi.org/10.1371/journal.pone.0204222
Vega CG, Bok M, Vlasova AN, Chattha KS, Gómez-Sebastián S, Nuñez C, Alvarado C, Lasa R, Escribano JM, Garaicoechea LL, Fernandez F, Bok K, Wigdorovitz A, Saif LJ, Parreño V (2013) Recombinant monovalent llama-derived antibody fragments (VHH) to rotavirus VP6 protect neonatal gnotobiotic piglets against human rotavirus-induced diarrhea. PLoS Pathog 9(5):e1003334–e1003334. https://doi.org/10.1371/journal.ppat.1003334
Vercruysse T, Pardon E, Vanstreels E, Steyaert J, Daelemans D (2010) An intrabody based on a llama single-domain antibody targeting the N-terminal alpha-helical multimerization domain of HIV-1 rev prevents viral production. J Biol Chem 285(28):21768–21780. https://doi.org/10.1074/jbc.M110.112490
Vincke C, Muyldermans S (2012) Introduction to heavy chain antibodies and derived nanobodies. In: Saerens D, Muyldermans S (eds) Single domain antibodies: methods and protocols, pp 15–26. Humana Press. https://doi.org/10.1007/978-1-61779-968-6_2
Walter JD, Hutter CAJ, Zimmermann I, Wyss M, Egloff P, Sorgenfrei M, Hürlimann LM, Gonda I, Meier G, Remm S, Thavarasah S, Plattet P, Seeger MA (2020) Sybodies targeting the SARS-CoV-2 receptor-binding domain. bioRxiv. 2020.2004.2016.045419. https://doi.org/10.1101/2020.04.16.045419
Wang L, Zhang L, Huang B, Li K, Hou G, Zhao Q, Wu C, Nan Y, Du T, Mu Y, Lan J, Chen H, Zhou E-M (2019) A nanobody targeting viral nonstructural protein 9 inhibits porcine reproductive and respiratory syndrome virus replication. J Virol 93(4):e01888-e1818. https://doi.org/10.1128/JVI.01888-18
Wanner N, Eden T, Liaukouskaya N, Koch-Nolte F (2021) Nanobodies: new avenue to treat kidney disease. Cell Tissue Res 385(2):445–456. https://doi.org/10.1007/s00441-021-03479-8
Wei G, Meng W, Guo H, Pan W, Liu J, Peng T, Chen L, Chen C-Y (2011) Potent neutralization of influenza A virus by a single-domain antibody blocking M2 ion channel protein. PLoS ONE 6(12):e28309–e28309. https://doi.org/10.1371/journal.pone.0028309
Wrapp, D, De Vlieger, D, Corbett, KS, Torres, GM, Wang, N, Van Breedam, W, Roose, K, van Schie, L, Team, V-CC-R, Hoffmann, M, Pöhlmann, S, Graham, BS, Callewaert, N, Schepens, B, Saelens, X, McLellan, JS (2020) Structural basis for potent neutralization of betacoronaviruses by single-domain camelid antibodies. Cell 181(5):1004–1015. https://doi.org/10.1016/j.cell.2020.04.031
Wu Y, Li C, Xia S, Tian X, Kong Y, Wang Z, Gu C, Zhang R, Tu C, Xie Y, Yang Z, Lu L, Jiang S, Ying T (2020) Identification of human single-domain antibodies against SARS-CoV-2. Cell Host Microbe 27(6):891–898. https://doi.org/10.1016/j.chom.2020.04.023
Xiang Y, Nambulli S, Xiao Z, Liu H, Sang Z, Duprex WP, Schneidman-Duhovny D, Zhang C, Shi Y (2020) Versatile and multivalent nanobodies efficiently neutralize SARS-CoV-2. Science (New York, NY) 370(6523):1479–1484. https://doi.org/10.1126/science.abe4747
Yang E, Liu Q, Huang G, Liu J, Wei W (2022) Engineering nanobodies for next-generation molecular imaging. Drug Discov Today 27(6):1622–1638. https://doi.org/10.1016/j.drudis.2022.03.013
Yang J, Lin S, Chen Z, Yang F, Guo L, Wang L, Duan Y, Zhang X, Dai Y, Yin K (2023) Development of a bispecific nanobody conjugate broadly neutralizes diverse SARS-CoV-2 variants and structural basis for its broad neutralization. PLoS Pathog 19(11):e1011804. https://doi.org/10.1371/journal.ppat.1011804
Yang Z, Schmidt D, Liu W, Li S, Shi L, Sheng J, Chen K, Yu H, Tremblay JM, Chen X (2014) A novel multivalent, single-domain antibody targeting TcdA and TcdB prevents fulminant Clostridium difficile infection in mice. J Infect Dis 210(6):964–972. https://doi.org/10.1093/infdis/jiu196
Ye G, Gallant JP, Massey C, Shi K, Tai W, Zheng J, Odle AE, Vickers MA, Shang J, Wan Y, Drelich A, Kempaiah KR, Tat V, Perlman S, Du L, Tseng C-T, Aihara H, LeBeau AM, Li F (2020) The development of a novel nanobody therapeutic for SARS-CoV-2. bioRxiv, 2020.2011.2017.386532. https://doi.org/10.1101/2020.11.17.386532
Zhao Y, Yang J, Niu Q, Wang J, Jing M, Guan G, Liu M, Luo J, Yin H, Liu Z (2023) Identification and characterization of nanobodies from a phage display library and their application in an immunoassay for the sensitive detection of African swine fever virus. J Clin Microbiol. https://doi.org/10.1128/jcm.01197-22
Zielonka S, Empting M, Grzeschik J, Könning D, Barelle CJ, Kolmar H (2015) Structural insights and biomedical potential of IgNAR scaffolds from sharks. Mabs 7(1):15–25. https://doi.org/10.4161/19420862.2015.989032
Zimmermann I, Egloff P, Hutter CAJ, Arnold FM, Stohler P, Bocquet N, Hug MN, Huber S, Siegrist M, Hetemann L, Gera J, Gmür S, Spies P, Gygax D, Geertsma ER, Dawson RJP, Seeger MA (2018) Synthetic single domain antibodies for the conformational trapping of membrane proteins. eLife 7:e34317. https://doi.org/10.7554/eLife.34317
Zupancic JM, Desai AA, Schardt JS, Pornnoppadol G, Makowski EK, Smith MD, Kennedy AA, de Mattos G, Barbosa M, Cascalho M, Lanigan TM, Tai AW, Tessier PM (2021a) Directed evolution of potent neutralizing nanobodies against SARS-CoV-2 using CDR-swapping mutagenesis. Cell Chem Biol 28(9):1379–1388. https://doi.org/10.1016/j.chembiol.2021.05.019
Zupancic JM, Schardt JS, Desai AA, Makowski EK, Smith MD, Pornnoppadol G, de Mattos G, Barbosa M, Cascalho M, Lanigan TM, Tessier PM (2021b) Engineered multivalent nanobodies potently and broadly neutralize SARS-CoV-2 variants. Advanced Therapeutics 4(8):2100099. https://doi.org/10.1002/adtp.202100099
Acknowledgements
This work was supported by a grant from the Academy of Scientific Research and Technology (ASRT), Science and Technology Center (STC) grant #D62/2020 to Ahmed S. Attia.
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB). This work was supported by a grant from the Academy of Scientific Research and Technology (ASRT), Science and Technology Center (STC) Grant #D62/2020 to Ahmed S. Attia.
Author information
Authors and Affiliations
Contributions
All authors have participated in collecting and analyzing the data, drafting, revising, and approving the submitted manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rizk, S.S., Moustafa, D.M., ElBanna, S.A. et al. Nanobodies in the fight against infectious diseases: repurposing nature's tiny weapons. World J Microbiol Biotechnol 40, 209 (2024). https://doi.org/10.1007/s11274-024-03990-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-024-03990-4