Abstract
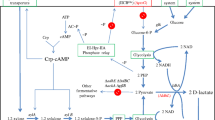
High-quality environmentally-friendly bioplastics can be produced by mixing poly-L-lactate with poly-D-lactate. On an industrial scale, this process simultaneously consumes large amounts of both optically pure lactate stereoisomers. However, because optimal growth conditions of L-lactate producers often differ from those of D-lactate producers, each stereoisomer is produced in a specialised facility, which raises cost and lowers sustainability. To address this challenge, we metabolically engineered Lactobacillus gasseri JCM 1131T, a bioprocess-friendly and genetically malleable strain of homofermentative lactic acid bacterium, to efficiently produce either pure L- or pure D-lactate under the same bioprocess conditions. Transformation of L. gasseri with plasmids carrying additional genes for L- or D-lactate dehydrogenases failed to affect the ratio of produced stereoisomers, but inactivation of the endogenous genes created strains which yielded 0.96 g of either L- or D-lactate per gram of glucose. In this study, the plasmid pHBintE, routinely used for gene disruption in Bacillus megaterium, was used for the first time to inactivate genes in lactobacilli. Strains with inactivated genes for endogenous lactate dehydrogenases efficiently fermented sugars released by enzymatic hydrolysis of alkali pre-treated wheat straw, an abundant lignocellulose-containing raw material, producing 0.37–0.42 g of lactate per gram of solid part of alkali-treated wheat straw. Thus, the constructed strains are primed to serve as producers of both optically pure L-lactate and D-lactate in the next-generation biorefineries.
Graphic abstract








Similar content being viewed by others
Data availability
Reported nucleotide sequence data are available in the Third Party Annotation Section of the DDBJ/ENA/GenBank databases under the accession numbers TPA: BK010903-BK010905. L. gasseri JCM 1131T is available from Japan Collection of Microorganisms (JCM).
References
Abdel-Rahman MA, Tashiro Y, Sonomoto K (2013) Recent advances in lactic acid production by microbial fermentation processes. Biotechnol Adv 31:877–902. https://doi.org/10.1016/j.biotechadv.2013.04.002
Alexandri M, Neu A-K, Schneider R, López-Gómez JP, Venus J (2019) Evaluation of various Bacillus coagulans isolates for the production of high purity L-lactic acid using defatted rice bran hydrolysates. Int J Food Sci Technol 54:1321–1329. https://doi.org/10.1111/ijfs.14086
Alting-Mees MA, Short JM (1989) pBluescript II: gene mapping vectors. Nucleic Acids Res 17:9494–9494. https://doi.org/10.1093/nar/17.22.9494
Azcarate-Peril MA et al (2008) Analysis of the genome sequence of Lactobacillus gasseri ATCC 33323 reveals the molecular basis of an autochthonous intestinal organism. Appl Environ Microbiol 74:4610. https://doi.org/10.1128/aem.00054-08
Barg H, Malten M, Jahn M, Jahn D (2005) Protein and vitamin production in Bacillus megaterium. In: Barredo J-L (ed) Microbial processes and products. Humana Press, Totowa, pp 205–223
Biswas I, Gruss A, Ehrlich SD, Maguin E (1993) High-efficiency gene inactivation and replacement system for gram-positive bacteria. J Bacteriol 175:3628. https://doi.org/10.1128/jb.175.11.3628-3635.1993
Bosma EF, Forster J, Nielsen AT (2017) Lactobacilli and pediococci as versatile cell factories: evaluation of strain properties and genetic tools. Biotechnol Adv 35:419–442. https://doi.org/10.1016/j.biotechadv.2017.04.002
Bruno-Bárcena JM, Azcárate-Peril MA, Klaenhammer TR, Hassan HM (2005) Marker-free chromosomal integration of the manganese superoxide dismutase gene (sodA) from Streptococcus thermophilus into Lactobacillus gasseri. FEMS Microbiol Lett 246:91–101. https://doi.org/10.1016/j.femsle.2005.03.044
Castillo Martinez FA, Balciunas EM, Salgado JM, Domínguez González JM, Converti A, Oliveira RPdS (2013) Lactic acid properties, applications and production: a review. Trends Food Sci Technol 30:70–83. https://doi.org/10.1016/j.tifs.2012.11.007
Chapot-Chartier M-P, Kulakauskas S (2014) Cell wall structure and function in lactic acid bacteria. Microb Cell Fact 13:S9. https://doi.org/10.1186/1475-2859-13-S1-S9
Chang DE, Jung HC, Rhee JS, Pan JG (1999) Homofermentative production of D- or L-Lactate in metabolically engineered Escherichia coli RR1. Appl Environ Microbiol 65:1384–1389. https://doi.org/10.1128/AEM.65.4.1384-1389.1999
Crawley AB, Henriksen ED, Stout E, Brandt K, Barrangou R (2018) Characterizing the activity of abundant, diverse and active CRISPR-Cas systems in lactobacilli. Sci Rep 8:11544. https://doi.org/10.1038/s41598-018-29746-3
Cselovszky J, Wolf G, Hammes WP (1992) Production of formate, acetate, and succinate by anaerobic fermentation of Lactobacillus pentosus in the presence of citrate. Appl Microbiol Biotechnol 37:94–97. https://doi.org/10.1007/BF00174210
Cubas-Cano E, González-Fernández C, Ballesteros M, Tomás-Pejó E (2018) Biotechnological advances in lactic acid production by lactic acid bacteria: lignocellulose as novel substrate. Biofuels Bioprod Biorefin 12:290–303. https://doi.org/10.1002/bbb.1852
Cubas-Cano E, González-Fernández C, Ballesteros M, Tomás-Pejó E (4023T) Lactobacillus pentosus CECT 4023T co-utilizes glucose and xylose to produce lactic acid from wheat straw hydrolysate: anaerobiosis as a key factor. Biotechnol Prog 35:e2739. https://doi.org/10.1002/btpr.2739
Desguin B, Soumillion P, Hausinger RP, Hols P (2017) Unexpected complexity in the lactate racemization system of lactic acid bacteria. Fems Microbiol Rev 41:S71–S83. https://doi.org/10.1093/femsre/fux021
Feldman-Salit A et al (2013) Regulation of the activity of lactate dehydrogenases from four lactic acid bacteria. J Biol Chem 288:21295–21306. https://doi.org/10.1074/jbc.M113.458265
Ferain T, Garmyn D, Bernard N, Hols P, Delcour J (1994) Lactobacillus plantarum ldhL gene: overexpression and deletion. J Bacteriol 176:596. https://doi.org/10.1128/jb.176.3.596-601.1994
Food and Agriculture Organization of the United Nations (2019) Crop Prospects and Food Situation: Quaterly Global Report, December 2019
Forde A, Fitzgerald GF (1999) Bacteriophage defence systems in lactic acid bacteria. In: Konings WN, Kuipers OP, Veld JHJH (eds) Lactic acid bacteria: genetics, metabolism and applications: Proceedings of the Sixth Symposium on lactic acid bacteria: genetics, metabolism and applications, 19–23 September 1999, Veldhoven, The Netherlands. Springer Netherlands, Dordrecht, pp 89–113. doi: 10.1007/978-94017-2027-4_4
Francl AL, Thongaram T, Miller MJ (2010) The PTS transporters of Lactobacillus gasseri ATCC 33323. BMC Microbiol 10:77. https://doi.org/10.1186/1471-2180-10-77
Garde A, Jonsson G, Schmidt AS, Ahring BK (2002) Lactic acid production from wheat straw hemicellulose hydrolysate by Lactobacillus pentosus and Lactobacillus brevis. Bioresour Technol 81:217–223. https://doi.org/10.1016/S0960-8524(01)00135-3
Glaasker E, Tjan FSB, Steeg PFT, Konings WN, Poolman B (1998) Physiological response of Lactobacillus plantarum to salt and nonelectrolyte stress. J Bacteriol 180(17):4718
Gottschalk G (2012) Bacterial metabolism. Springer, New York, Heidlberg, Berlin
Hama S, Mizuno S, Kihara M, Tanaka T, Ogino C, Noda H, Kondo A (2015) Production of D-lactic acid from hardwood pulp by mechanical milling followed by simultaneous saccharification and fermentation using metabolically engineered Lactobacillus plantarum. Bioresour Technol 187:167–172. https://doi.org/10.1016/j.biortech.2015.03.106
Hanahan D, Jessee J, Bloom FR (1991) Plasmid transformation of Escherichia coli and other bacteria. Methods in enzymology, vol 204. Academic Press, Cambdrige, pp 63–113
Hassan SS, Williams GA, Jaiswal AK (2019) Lignocellulosic biorefineries in Europe: current state and prospects. Trends Biotechnol 37:231–234. https://doi.org/10.1016/j.tibtech.2018.07.002
Hofvendahl K, Hahn-Hägerdal B (2000) Factors affecting the fermentative lactic acid production from renewable resources. Enzyme Microb Technol 26:87–107. https://doi.org/10.1016/S0141-0229(99)00155-6
Ivančić Šantek M, Lisičar J, Mušak L, Vrana Špoljarić I, Beluhan S, Šantek B (2018) Lipid production by yeast Trichosporon oleaginosus on the enzymatic hydrolysate of alkaline pretreated corn cobs for biodiesel production. Energy Fuels 32:12501–12513. https://doi.org/10.1021/acs.energyfuels.8b02231
Karlskås IL, Maudal K, Axelsson L, Rud I, Eijsink VGH, Mathiesen G (2014) Heterologous protein secretion in lactobacilli with modified pSIP vectors. PLoS ONE 9:e91125. https://doi.org/10.1371/journal.pone.0091125
Kleerebezem M, Beerthuyzen MM, Vaughan EE, de Vos WM, Kuipers OP (1997) Controlled gene expression systems for lactic acid bacteria: transferable nisin-inducible expression cassettes for Lactococcus, Leuconostoc, and Lactobacillus spp. Appl Environ Microbiol 63:4581
Kuo Y-C, Yuan S-F, Wang C-A, Huang Y-J, Guo G-L, Hwang W-S (2015) Production of optically pure L-lactic acid from lignocellulosic hydrolysate by using a newly isolated and D-lactate dehydrogenase gene-deficient Lactobacillus paracasei strain. Bioresour Technol 198:651–657. https://doi.org/10.1016/j.biortech.2015.09.071
Kylä-Nikkilä K, Hujanen M, Leisola M, Palva A (2000) Metabolic engineering of Lactobacillus helveticus CNRZ32 for production of pure L-(+)-lactic acid. Appl Environ Microbiol 66:3835. https://doi.org/10.1128/aem.66.9.3835-3841.2000
Lauer E, Kandler O (1980) Lactobacillus gasseri sp. nov., a new species of the subgenus Thermobacterium. Zentralblatt für Bakteriologie: I Abt Originale C: Allgemeine, angewandte und ökologische Mikrobiologie 1:75–78. https://doi.org/10.1016/S0172-5564(80)80019-4
Leenhouts K et al (1996) A general system for generating unlabelled gene replacements in bacterial chromosomes. Mol Gen Genet 253:217–224. https://doi.org/10.1007/s004380050315
Liang S, Gao D, Liu H, Wang C, Wen J (2018) Metabolomic and proteomic analysis of D-lactate-producing Lactobacillus delbrueckii under various fermentation conditions. J Ind Microbiol Biotechnol 45:681–696. https://doi.org/10.1007/s10295-018-2048-y
Liu SQ (2003) Practical implications of lactate and pyruvate metabolism by lactic acid bacteria in food and beverage fermentations. Int J Food Microbiol 83:115–131. https://doi.org/10.1016/S0168-1605(02)00366-5
Makarova K et al (2006) Comparative genomics of the lactic acid bacteria. Proc Natl Acad Sci USA 103:15611. https://doi.org/10.1073/pnas.0607117103
Marđetko N, Novak M, Trontel A, Grubišić M, Galić M, Šantek B (2018) Bioethanol production from dilute-acid pre-treated wheat straw liquor hydrolysate by genetically engineered Saccharomyces cerevisiae. Chem Biochem Eng Q 32:483–499. https://doi.org/10.15255/CABEQ.2018.1409
Marshall OJ (2004) PerlPrimer: cross-platform, graphical primer design for standard, bisulphite and real-time PCR. Bioinformatics 20:2471–2472. https://doi.org/10.1093/bioinformatics/bth254
Martín-Platero AM, Valdivia E, Maqueda M, Martínez-Bueno M (2007) Fast, convenient, and economical method for isolating genomic DNA from lactic acid bacteria using a modification of the protein “salting-out” procedure. Anal Biochem 366:102–104. https://doi.org/10.1016/j.ab.2007.03.010
Mazzoli R (2019) Metabolic engineering strategies for consolidated production of lactic acid from lignocellulosic biomass. Biotechnol Appl Biochem. https://doi.org/10.1002/bab.1869
Miller EM, Nickoloff JA (1995) Escherichia coli electrotransformation. In: Nickoloff JA (ed) Electroporation protocols for microorganisms. Humana Press, Totowa, pp 105–113
Moncada JB, Aristizábal VM, Cardona CA (2016) Design strategies for sustainable biorefineries. Biochem Eng J 116:122–134. https://doi.org/10.1016/j.bej.2016.06.009
Montane D, Farriol X, Salvadó J, Jollez P, Chornet E (1998) Application of steam explosion to the fractionation and rapid vapor-phase alkaline pulping of wheat straw. Biomass Bioenergy 14:261–276. https://doi.org/10.1016/S0961-9534(97)10045-9
Okano K, Hama S, Kihara M, Noda H, Tanaka T, Kondo A (2017) Production of optically pure D-lactic acid from brown rice using metabolically engineered Lactobacillus plantarum. Appl Microbiol Biotechnol 101:1869–1875. https://doi.org/10.1007/s00253-016-7976-8
Okano K, Uematsu G, Hama S, Tanaka T, Noda H, Kondo A, Honda K (2018) Metabolic engineering of Lactobacillus plantarum for direct L-lactic acid production from raw corn starch. Biotechnol J 13:1700517. https://doi.org/10.1002/biot.201700517
Okano K, Zhang Q, Shinkawa S, Yoshida S, Tanaka T, Fukuda H, Kondo A (2009) Efficient production of optically pure D-lactic acid from raw corn starch by using a genetically modified L-lactate dehydrogenase gene-deficient and α-amylase-secreting Lactobacillus plantarum strain. Appl Environ Microbiol 75:462. https://doi.org/10.1128/aem.01514-08
Oliveira RAd, Komesu A, Rossell CEV, Filho RM (2018) Challenges and opportunities in lactic acid bioprocess design-From economic to production aspects. Biochem Eng J 133:219–239. https://doi.org/10.1016/j.bej.2018.03.003
Overbeek R et al (2014) The SEED and the rapid annotation of microbial genomes using subsystems technology (RAST). Nucleic Acids Res 42:D206–D214. https://doi.org/10.1093/nar/gkt1226
Passoth V, Sandgren M (2019) Biofuel production from straw hydrolysates: current achievements and perspectives. Appl Microbiol Biotechnol. https://doi.org/10.1007/s00253-019-09863-3
Posno M, Heuvelmans PT, van Giezen MJ, Lokman BC, Leer RJ, Pouwels PH (1991) Complementation of the inability of Lactobacillus strains to utilize D-xylose with D-xylose catabolism-encoding genes of Lactobacillus pentosus. Appl Environ Microbiol 57:2764
Puah SM, Huynh HV, Wu JC (2013) Novel two-in-one bioreactor greatly improves lactic acid production from xylose by Lactobacillus pentosus. J Chem Technol Biotechnol 88:594–598. https://doi.org/10.1002/jctb.3867
Pusch O, Kalyanaraman R, Tucker LD, Wells JM, Ramratnam B, Boden D (2006) An anti-HIV microbicide engineered in commensal bacteria: secretion of HIV-1 fusion inhibitors by lactobacilli. AIDS 20:1917–1922. https://doi.org/10.1097/01.aids.0000247112.36091.f8
Russell WM, Klaenhammer TR (2001) Efficient system for directed integration into the Lactobacillus acidophilus and Lactobacillus gasseri chromosomes via homologous recombination. Appl Environ Microbiol 67:4361. https://doi.org/10.1128/aem.67.9.4361-4364.2001
Saha BC, Iten LB, Cotta MA, Wu YV (2005) Dilute acid pretreatment, enzymatic saccharification and fermentation of wheat straw to ethanol. Process Biochem 40:3693–3700. https://doi.org/10.1016/j.procbio.2005.04.006
Sambrook J (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York
Sanozky-Dawes R, Selle K, Flaherty S, Klaenhammer T, Barrangou R (2015) Occurrence and activity of a type II CRISPR-Cas system in Lactobacillus gasseri. Microbiology 161:1752–1761. https://doi.org/10.1099/mic.0.000129
Sauer M, Russmayer H, Grabherr R, Peterbauer CK, Marx H (2017) The efficient clade: lactic acid bacteria for industrial chemical production. Trends Biotechnol 35:756–769. https://doi.org/10.1016/j.tibtech.2017.05.002
Sørvig E, Mathiesen G, Naterstad K, Eijsink VGH, Axelsson L (2005) High-level, inducible gene expression in Lactobacillus sakei and Lactobacillus plantarum using versatile expression vectors. Microbiology 151:2439–2449. https://doi.org/10.1099/mic.0.28084-0
Sun Z et al (2015) Expanding the biotechnology potential of lactobacilli through comparative genomics of 213 strains and associated genera. Nat Commun 6:8322. https://doi.org/10.1038/ncomms9322
Štafa A, Miklenić MS, Zandona A, Žunar B, Čadež N, Petković H, Svetec IK (2017) In Saccharomyces cerevisiae gene targeting fidelity depends on a transformation method and proportion of the overall length of the transforming and targeted DNA. FEMS Yeast Res 17:fox041. https://doi.org/10.1093/femsyr/fox041
Talebnia F, Karakashev D, Angelidaki I (2010) Production of bioethanol from wheat straw: an overview on pretreatment, hydrolysis and fermentation. Bioresour Technol 101:4744–4753. https://doi.org/10.1016/j.biortech.2009.11.080
The R Core Team (2019) R: a language and environment for statistical computing 3.5.2 edn. R Foundation for Statistical Computing, Vienna
Townsend TJ, Sparkes DL, Wilson P (2017) Food and bioenergy: reviewing the potential of dual-purpose wheat crops. GCB Bioenergy 9:525–540. https://doi.org/10.1111/gcbb.12302
Tsuge Y, Kato N, Yamamoto S, Suda M, Jojima T, Inui M (2019) Metabolic engineering of Corynebacterium glutamicum for hyperproduction of polymer-grade L- and D-lactic acid. Appl Microbiol Biotechnol 103:3381–3391. https://doi.org/10.1007/s00253-019-09737-8
Tsuji H (2005) Poly(lactide) Stereocomplexes: formation, structure, properties, degradation, and applications. Macromol Biosci 5:569–597. https://doi.org/10.1002/mabi.200500062
Wegkamp A, Starrenburg M, de Vos WM, Hugenholtz J, Sybesma W (2004) Transformation of folate-consuming Lactobacillus gasseri into a folate producer. Appl Environ Microbiol 70:3146. https://doi.org/10.1128/aem.70.5.3146-3148.2004
Yang B, Dai Z, Ding SY, Wyman CE (2011) Enzymatic hydrolysis of cellulosic biomass. Biofuels 2:421–449. https://doi.org/10.4155/BFS.11.116
Yi X, Zhang P, Sun J, Tu Y, Gao Q, Zhang J, Bao J (2016) Engineering wild-type robust Pediococcus acidilactici strain for high titer L- and D-lactic acid production from corn stover feedstock. J Biotechnol 217:112–121. https://doi.org/10.1016/j.jbiotec.2015.11.014
Yuan S-F et al (2018) Production of optically pure L(+)-lactic acid from waste plywood chips using an isolated thermotolerant Enterococcus faecalis SI at a pilot scale. J Ind Microbiol Biotechnol 45:961–970. https://doi.org/10.1007/s10295-018-2078-5
Zhang C, Zhou C, Assavasirijinda N, Yu B, Wang L, Ma Y (2017) Non-sterilized fermentation of high optically pure D-lactic acid by a genetically modified thermophilic Bacillus coagulans strain. Microb Cell Fact 16:213. https://doi.org/10.1186/s12934-017-0827-1
Zhang Y, Chen X, Luo J, Qi B, Wan Y (2014) An efficient process for lactic acid production from wheat straw by a newly isolated Bacillus coagulans strain IPE22. Bioresour Technol 158:396–399. https://doi.org/10.1016/j.biortech.2014.02.128
Zhang Y, Kumar A, Hardwidge PR, Tanaka T, Kondo A, Vadlani PV (2016) D-lactic acid production from renewable lignocellulosic biomass via genetically modified Lactobacillus plantarum. Biotechnol Prog 32:271–278. https://doi.org/10.1002/btpr.2212
Zhao Y, Wang Y, Zhu JY, Ragauskas A, Deng Y (2008) Enhanced enzymatic hydrolysis of spruce by alkaline pretreatment at low temperature. Biotechnol Bioeng 99:1320–1328. https://doi.org/10.1002/bit.21712
Funding
This work was supported by the Croatian Science Foundation (Grant Numbers IP-11-2013-9158-SPECH-LRM and IP-01-2018-9717-SPB-LCF, awarded to Prof Božidar Šantek).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study design. Material preparation, data collection and analysis were performed by BŽ, AT, JLP, MSM, AS, NM and MN. BŠ and IKS initiated and coordinated the study. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Žunar, B., Trontel, A., Svetec Miklenić, M. et al. Metabolically engineered Lactobacillus gasseri JCM 1131 as a novel producer of optically pure L- and D-lactate. World J Microbiol Biotechnol 36, 111 (2020). https://doi.org/10.1007/s11274-020-02887-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-020-02887-2