Abstract
For decades, diverse plasmid vectors have been continuously developed for molecular cloning of DNA fragment in the bacterial host cell Escherichia coli. Even with deliberate performances in vector preparation, the cloning approaches still face inevitable background colonies, or false positive clones, that may be arisen from intact or self-ligated plasmid molecules. To assist in such problem, two plasmids, pBS2ndd and pBS3ndd, which resistant to ampicillin and kanamycin respectively, were developed in this study as more advantageous cloning vector. The plasmids carry ndd, a lethal gene from bacteriophage T4 coding for nucleoid disruption protein that binds to the host chromosome and progressively kill the cell. The deadly toxicity of Ndd inhibits host cells that obtain intact or ndd-religated vector from growing, which results in low background and dramatically reduces the effort for selection of recombinants. Moreover, their identical multiple cloning site was designed to support various cloning strategies. Digestion of plasmids with XcmI allows for in vitro T/A ligation, while with EcoRV permits blunt-end ligation, with capability of blue-white colony screening. In vivo homologous recombination cloning is also utilizable by amplification of insert fragments using primers containing homology arms and transformation into capable E. coli strains. To demonstrate their advantages, the plasmids were used to clone PCR product samples for DNA sequencing with low-background and versatile cloning strategies. Such rapid and cost-effective cloning procedures are also proposed here. Finally, the cloning for protein expression with blue-white selection was also possible using egfp as a model regulated by lac and T7 promoters on the plasmid or other build-in promoters with the insert.
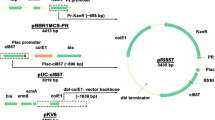
Graphical Abstract









Similar content being viewed by others
References
Aslanidis C, De Jong PJ (1990) Ligation-independent cloning of PCR products (LIC-PCR). Nucleic Acids Res 18:6069–6074
Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K (2003) Current protocols in molecular biology. Wiley, New York
Bernard P, Gabant P, Bahassi EM, Couturier M (1994) Positive-selection vectors using the F plasmid ccdB killer gene. Gene 148:71–74. https://doi.org/10.1016/0378-1119(94)90235-6
Bervoets I, Van Brempt M, Van Nerom K, Van Hove B, Maertens J, De Mey M, Charlier D (2018) A sigma factor toolbox for orthogonal gene expression in Escherichia coli. Nucleic Acids Res 46:2133–2144. https://doi.org/10.1093/nar/gky010
Bolivar F, Rodriguez RL, Greene PJ, Betlach MC, Heyneker HL, Boyer HW, Crosa JH, Falkow S (1977) Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene 2:95–113. https://doi.org/10.1016/0378-1119(77)90000-2
Bouet JY, Woszczyk J, Repoila F, François V, Louarn JM, Krisch HM (1994) Direct PCR sequencing of the ndd gene of bacteriophage T4: identification of a product involved in bacterial nucleoid disruption. Gene 141:9–16. https://doi.org/10.1016/0378-1119(94)90121-X
Bouet JY, Campo NJ, Krisch HM, Louarn JM (1996) The effects on Escherichia coli of expression of the cloned bacteriophage T4 nucleoid disruption (ndd) gene. Mol Microbiol 20:519–528. https://doi.org/10.1046/j.1365-2958.1996.5411067.x
Bouet JY, Krisch HM, Louarn JM (1998) Ndd, the bacteriophage T4 protein that disrupts the Escherichia coli nucleoid, has a DNA binding activity. J Bacteriol 180:5227–5230
Bubeck P, Winkler M, Bautsch W (1993) Rapid cloning by homologous recombination in vivo. Nucleic Acids Res 21:3601–3602
Casali N (2003) Escherichia coli host strains. Methods Mol Biol 235:27–48. https://doi.org/10.1385/1-59259-409-3:27
Chang ACY, Cohen SN (1978) Construction and characterization of amplifiable multicopy DNA cloning vehicles derived from the P15A cryptic miniplasmid. J Bacteriol 134:1141–1156
Clark JM (1988) Novel non-templated nucleotide addition reactions catalyzed by procaryotic and eucaryotic DNA polymerases. Nucleic Acids Res 16:9677–9686
Cunningham DS, Koepsel RR, Ataai MM, Domach MM (2009) Factors affecting plasmid production in Escherichia coli from a resource allocation standpoint. Microb Cell Fact 8:27. https://doi.org/10.1186/1475-2859-8-27
De Bellis D, Schwartz I (1990) Regulated expression of foreign genes fused to lac: control by glucose levels in growth medium. Nucleic Acids Res 18:1311
Dennison C, Lovrien R (1997) Three phase partitioning: concentration and purification of proteins. Protein Expr Purif 11:149–161. https://doi.org/10.1006/prep.1997.0779
Glascock CB, Weickert MJ (1998) Using chromosomal lacI Q1 to control expression of genes on high-copy-number plasmids in Escherichia coli. Gene 223:221–231. https://doi.org/10.1016/S0378-1119(98)00240-6
Görke B, Stülke J (2008) Carbon catabolite repression in bacteria: many ways to make the most out of nutrients. Nat Rev Microbiol 6:613–624. https://doi.org/10.1038/nrmicro1932
Henrich B, Plapp R (1986) Use of the lysis gene of bacteriophage ΦX174 for the construction of a positive selection vector. Gene 42:345–349. https://doi.org/10.1016/0378-1119(86)90239-8
Holton TA, Graham MW (1991) A simple and efficient method for direct cloning of PCR products using ddT-tailed vectors. Nucleic Acids Res 19:1156
Ichihara Y, Kurosawa Y (1993) Construction of new T vectors for direct cloning of PCR products. Gene 130:153–154. https://doi.org/10.1016/0378-1119(93)90361-6
Jacobus AP, Gross J (2015) Optimal cloning of PCR fragments by homologous recombination in Escherichia coli. PLoS ONE 10:e0119221. https://doi.org/10.1371/journal.pone.0119221
Jechlinger W, Szostak MP, Witte A, Lubitz W (1999) Altered temperature induction sensitivity of the lambda pR/cI857 system for controlled gene E expression in Escherichia coli. FEMS Microbiol Lett 137:347–352. https://doi.org/10.1111/j.1574-6968.1999.tb13524.x
Jo C, Jo SA (2001) A simple method to construct T-vectors using XcmI cassettes amplified by nonspecific PCR. Plasmid 45:37–40. https://doi.org/10.1006/plas.2000.1500
Kaiser K, Murray NE (1980) On the nature of sbcA mutations in E. coli K12. Mol Gen Genet 179:555–563. https://doi.org/10.1007/BF00271745
Karcher SJ (1995) Molecular biology: a project approach. Academic Press, San Diego
Katana A, Kwiatowski J, Spalik K, Zakryś B, Szalacha E, Szymańska H (2001) Phylogenetic position of Koliella (Chlorophyta) as inferred from nuclear and chloroplast small subunit rDNA. J Phycol 37:443–451. https://doi.org/10.1046/j.1529-8817.2001.037003443.x
Kim HG, Kim HS, Hwang HJ, Chung SK, Lee JM, Chung DK (2004) Construction of a pTOC-T vector using GST-ParE toxin for direct cloning and selection of PCR products. Biotechnol Lett 26:1659–1663. https://doi.org/10.1007/s10529-004-3518-z
Kirschman JA, Cramer JH (1988) Two new tools: multi-purpose cloning vectors that carry kanamycin or spectinomycin/streptomycin resistance markers. Gene 68:163–165. https://doi.org/10.1016/0378-1119(88)90609-9
Koerner JF, Thies SK, Snustad DP (1979) Protein induced by bacteriophage T4 which is absent in Escherichia coli infected with nuclear disruption-deficient phage mutants. J Virol 31:506–513
Kolodner R, Hall SD, Luisi-DeLuca C (1994) Homologous pairing proteins encoded by the Escherichia coli recE and recT genes. Mol Microbiol 11:23–30. https://doi.org/10.1111/j.1365-2958.1994.tb00286.x
Kovalic D, Kwak JH, Weisblum B (1991) General method for direct cloning of DNA fragments generated by the polymerase chain reaction. Nucleic Acids Res 19:4560. https://doi.org/10.1093/nar/19.16.4560
Li MZ, Elledge SJ (2007) Harnessing homologous recombination in vitro to generate recombinant DNA via SLIC. Nat Methods 4:251–256. https://doi.org/10.1038/nmeth1010
Lovett ST, Hurley RL, Sutera VA Jr, Aubuchon RH, Lebedeva MA (2002) Crossing over between regions of limited homology in Escherichia coli: RecA-dependent and RecA-independent pathways. Genetics 160:851–859
Luria SE, Human ML (1950) Chromatin staining of bacteria during bacteriophage infection. J Bacteriol 59:551–560
Malys N (2012) Shine-Dalgarno sequence of bacteriophage T4: GAGG prevails in early genes. Mol Biol Rep 39:33–39. https://doi.org/10.1007/s11033-011-0707-4
Marchuk D, Drumm M, Saulino A, Collins FS (1991) Construction of T-vectors, a rapid and general system for direct cloning of unmodified PCR products. Nucleic Acids Res 19:1154. https://doi.org/10.1093/nar/19.5.1154
Maunders MJ (1993) DNA polymerases (EC 2.7.7.7). Enzym Mol Biol. https://doi.org/10.1385/0-89603-234-5:17
McManus HA, Lewis LA (2011) Molecular phylogenetic relationships in the freshwater family Hydrodictyaceae (Sphaeropleales, Chlorophyceae), with an emphasis on Pediastrum duplex. J Phycol 47:152–163. https://doi.org/10.1111/j.1529-8817.2010.00940.x
Mead DA, Pey NK, Herrnstadt C, Marcil RA, Smith LM (1991) A universal method for the direct cloning of PCR amplified nucleic acid. Nat Biotechnol 9:657–663. https://doi.org/10.1038/nbt0791-657
Miller WG, Lindow SE (1997) An improved GFP cloning cassette designed for prokaryotic transcriptional fusions. Gene 191:149–153. https://doi.org/10.1016/S0378-1119(97)00051-6
Miller ES, Kutter E, Mosig G, Arisaka F, Kunisawa T, Rüger W (2003) Bacteriophage T4 genome. Microbiol Mol Biol Rev 67:86–156. https://doi.org/10.1128/MMBR.67.1.86-156.2003
Müller-Hill B, Crapo L, Gilbert W (1968) Mutants that make more lac repressor. Proc Natl Acad Sci USA 59:1259–1264
Newton GL, Koledin T, Gorovitz B, Rawat M, Fahey RC, Av-Gay Y (2003) The glycosyltransferase gene encoding the enzyme catalyzing the first step of mycothiol biosynthesis (mshA). J Bacteriol 185:3476–3479. https://doi.org/10.1128/JB.185.11.3476-3479.2003
Oliner JD, Kinzler KW, Vogelstein B (1993) In vivo cloning of PCR products in E. coli. Nucleic Acids Res 21:5192–5197
Ozaki LS, Maeda S, Shimada K, Takagi Y (1980) A novel ColE1::Tn3 plasmid vector that allows direct selection of hybrid clones in E. coli. Gene 8:301–314. https://doi.org/10.1016/0378-1119(80)90006-2
Parrish JR, Limjindaporn T, Hines JA, Liu J, Liu G, Finley RL Jr (2004) High-throughput cloning of Campylobacter jejuni ORFs by in vivo recombination in Escherichia coli. J Proteome Res 3:582–586. https://doi.org/10.1021/pr0341134
Preston A (2003) Choosing a cloning vector. Methods Mol Biol 235:19–26. https://doi.org/10.1385/1-59259-409-3:19
Rosano GL, Ceccarelli EA (2014) Recombinant protein expression in Escherichia coli: advances and challenges. Front Microbiol 5:172. https://doi.org/10.3389/fmicb.2014.00172
Rotchanapreeda T, Wongniam S, Swangpol SC, Chareonsap PP, Sukkaewmanee N, Somana J (2016) Development of SSR markers from Musa balbisiana for genetic diversity analysis among Thai bananas. Plant Syst Evol 302:739–761. https://doi.org/10.1007/s00606-015-1274-2
Saeed IA, Ashraf SS (2009) Denaturation studies reveal significant differences between GFP and blue fluorescent protein. Int J Biol Macromol 45:236–241. https://doi.org/10.1016/j.ijbiomac.2009.05.010
Sambrook J, Russell DW (2006) Preparation of plasmid DNA by alkaline lysis with SDS: minipreparation. CSH Protoc. https://doi.org/10.1101/pdb.prot4084
Siebenlist U, Gilbert W (1980) Contacts between Escherichia coli RNA polymerase and an early promoter of phage T7. Proc Natl Acad Sci USA 77:122–126
Siebenlist U, Simpson RB, Gilbert W (1980) E. coli RNA polymerase interacts homologously with two different promoters. Cell 20:269–281. https://doi.org/10.1016/0092-8674(80)90613-3
Snustad DP (1974) Mutants of bacteriophage T4 deficient in the ability to induce nuclear disruption: II. Physiological state of the host nucleoid in infected cells. J Mol Biol 89:675–687
Snustad DP, Bursch CJH (1977) Shutoff of host RNA synthesis in bacteriophage T4-infected Escherichia coli in the absence of host DNA degradation and nuclear disruption. J Virol 21:1240–1242
Snustad DP, Conroy LM (1974) Mutants of bacteriophage T4 deficient in the ability to induce nuclear disruption: I. Isolation and genetic characterization. J Mol Biol 89:663–673
Snustad DP, Warner HR, Parson KA, Anderson DL (1972) Nuclear disruption after infection of Escherichia coli with a bacteriophage T4 mutant unable to induce endonuclease II. J Virol 10:124–133
Snustad DP, Bursch CJH, Parson KA, Hefeneider SH (1976) Mutants of bacteriophage T4 deficient in the ability to induce nuclear disruption: shutoff of host DNA and protein synthesis, gene dosage experiments, identification of a restrictive host, and possible biological significance. J Virol 18:268–288
Swangpol S, Volkaert H, Sotto RC, Seelanan T (2007) Utility of selected non-coding chloroplast DNA sequences for lineage assessment of Musa interspecific hybrids. J Biochem Mol Biol 40:577–587. https://doi.org/10.5483/BMBRep.2007.40.4.577
Swangpol CS, Triperm P, Somana J, Sukkaewmanee N, Srisanga P, Suksathan P (2015) Musa nanensis, a new banana (Musaceae) species from northern Thailand. Syst Bot 40(2):426–432. https://doi.org/10.1600/036364415X688790
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17:1105–1109. https://doi.org/10.1007/BF00037152
Techaprasan J, Ngamriabsakul C, Klinbunga S, Chusacultanachai S, Jenjittikul T (2006) Genetic variation and species identification of Thai Boesenbergia (Zingiberaceae) analyzed by chloroplast DNA polymorphism. J Biochem Mol Biol 39:361–370. https://doi.org/10.5483/BMBRep.2006.39.4.361
Thomson CM, Ward WW (2002) Three-phase partitioning (TPP): a rapid and preparative purification tool for GFP. Biolumin Chemilumin. https://doi.org/10.1142/9789812776624_0025
Vernet T, Lau PCK, Narang SA, Visentin LP (1985) A direct-selection vector derived from pColE3-CA38 and adapted for foreign gene expression. Gene 34:87–93. https://doi.org/10.1016/0378-1119(85)90298-7
von Gabain A, Bujard H (1979) Interaction of Escherichia coli RNA polymerase with promoters of several coliphage and plasmid DNAs. Proc Natl Acad Sci USA 76:189–193
Walhout AJ, Temple GF, Brasch MA, Hartley JL, Lorson MA, van den Heuvel S, Vidal M (2000) GATEWAY recombinational cloning: application to the cloning of large numbers of open reading frames or ORFeomes. Methods Enzymol 328:575–592. https://doi.org/10.1016/S0076-6879(00)28419-X
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, New York, pp 315–322
Yazynin SA, Deyev SM, Jucovič M, Hartley RW (1996) A plasmid vector with positive selection and directional cloning based on a conditionally lethal gene. Gene 169:131–132. https://doi.org/10.1016/0378-1119(95)00814-4
Yazynin S, Lange H, Mokros T, Deyev S, Lemke H (1999) A new phagemid vector for positive selection of recombinants based on a conditionally lethal barnase gene. FEBS Lett 452:351–354. https://doi.org/10.1016/S0014-5793(99)00661-4
Zhang Y, Werling U, Edelmann W (2012) SLiCE: a novel bacterial cell extract-based DNA cloning method. Nucleic Acids Res 40:e55. https://doi.org/10.1093/nar/gkr1288
Acknowledgements
This research was financially supported by Agricultural Research Development Agency (ARDA), Thailand, with grant number CRP5605011990 and Science Achievement Scholarship of Thailand from Office of the Higher Education Commission for Manadsaree Klomtun. The authors are much obliged to Prof. Dr. Chayan Picheansoonthon and Dr. Boonmee Phokham for their distribution of the plant materials of C. limiana, C. saxicola, and C. yunnanensis. Our gratitude is also expressed to Dr. Thakorn Sornwatana for the advices about the protein experiment.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rotchanapreeda, T., Ngonsawan, W., Klomtun, M. et al. The plasmid vectors, pBS2ndd and pBS3ndd, for versatile cloning with low background in Escherichia coli. World J Microbiol Biotechnol 34, 85 (2018). https://doi.org/10.1007/s11274-018-2466-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-018-2466-z