Abstract
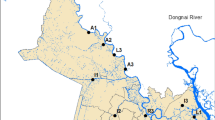
Polluted river water systems can contribute to the acquisition and propagation of antibiotic resistance in the ecosystem. This study focused on describing the occurrence of antibiotic-resistant bacteria and genes in a river system in relation to the concentration of organic pollutants. Water samples were collected in triplicate from 13 different sampling points of the Bagmati River and analyzed for physicochemical characteristics and antibiotic-resistant bacteria. The presence of antibiotic resistance genes (ARGs) in the isolated organisms was studied using conventional polymerase chain reaction (PCR). It was observed that ARGs and antibiotic-resistance bacteria (ARB) showed a statistically significant positive connection with organic contaminants. The mean ratio of doxycycline, oxytetracycline, and sulphamethoxazole-resistant bacterial population to the total bacterial population was found to be 0.44 ± 0.068, 0.42 ± 0.067, and 0.46 ± 0.087, respectively. Here, 40 bacterial isolates were identified, of which 70% were Escherichia coli, 16% were Klebsiella spp., 5% were Staphylococcus aureus, 3% were Salmonella Typhi, 3% were Pseudomonas aeruginosa, and 3% were Alcaligenes faecalis. Antimicrobial susceptibility testing showed 80% of the bacterial isolates were multi-drug resistant. Frequent detection of selected genes (tetA and sul1) through conventional PCR in the isolates (90.6%) indicated that the river water was heavily contaminated with antibiotic resistance genes. Among the isolates harboring the genes, most of them were found to be phenotypically resistant to the respective antibiotics. The presence of therapeutically pertinent ARGs in the river system shows the necessity of keeping track of environmental antibiotic resistance and corrective measures to overcome challenges.









Similar content being viewed by others
Data Availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Alderton, I., Palmer, B. R., Heinemann, J. A., Pattis, I., Weaver, L., Gutiérrez Ginés, M. J., Horswell, J., & Tremblay, L. A. (2021). The role of emerging organic contaminants in the development of antimicrobial resistance. Emerging Contaminants, 7, 160–171. https://doi.org/10.1016/j.emcon.2021.07.001
Amarasiri, M., Takezawa, T., Malla, B., Furukawa, T., Sherchand, J. B., Haramoto, E., & Sei, K. (2022). Prevalence of antibiotic resistance genes in drinking and environmental water sources of the Kathmandu Valley, Nepal. Frontiers in Microbiology, 13, 894014. https://doi.org/10.3389/fmicb.2022.894014
Andersson, D. I., & Hughes, D. (2010). Antibiotic resistance and its cost: Is it possible to reverse resistance? Nature Reviews Microbiology, 8(4), 260–271. https://doi.org/10.1038/nrmicro2319
APHA. (2017). Standard Methods for the Examination of Water and Wastewater (23rd ed.). American Public Health Association.
Bajracharya, R., & Tamrakar, N. K. (1970). Environmental status of Manahara River, Kathmandu, Nepal. Bulletin of the Department of Geology, 10, 21–32. https://doi.org/10.3126/bdg.v10i0.1417
Clinical and Laboratory Standards Institute. (2018). M100 Performance Standards for Antimicrobial Susceptibility Testing, 8(3). http://www.emeraldinsight.com/doi/10.1108/08876049410065598
Frank, T., Gautier, V., Talarmin, A., Bercion, R., & Arlet, G. (2007). Characterization of sulphonamide resistance genes and class 1 integron gene cassettes in Enterobacteriaceae, Central African Republic (CAR). Journal of Antimicrobial Chemotherapy, 59(4), 742–745. https://doi.org/10.1093/jac/dkl538
Gao, J., Yu, F. Q., Luo, L. P., He, J. Z., Hou, R. G., Zhang, H. Q., Li, S. M., Su, J. L., & Han, B. (2012). Antibiotic resistance of Streptococcus agalactiae from cows with mastitis. Veterinary Journal, 194(3), 423–424. https://doi.org/10.1016/j.tvjl.2012.04.020
Guo, X., Pang, W., Dou, C., & Yin, D. (2017). Sulfamethoxazole and COD increase abundance of sulfonamide resistance genes and change bacterial community structures within sequencing batch reactors. Chemosphere, 175, 21–27. https://doi.org/10.1016/j.chemosphere.2017.01.134
Jiang, L., Hu, X., Xu, T., Zhang, H., Sheng, D., & Yin, D. (2013). Prevalence of antibiotic resistance genes and their relationship with antibiotics in the Huangpu River and the drinking water sources, Shanghai, China. Science of the Total Environment, 458, 267–272. https://doi.org/10.1016/j.scitotenv.2013.04.038
Joshi, D. R. (2017). The wastewater resistome: Lurking antibiotic resistance in the environment. Tribhuvan University Journal of Microbiology, 4, 79–83. https://doi.org/10.3126/tujm.v4i0.21681
Kaczorek, E., Małaczewska, J., Wójcik, R., Rękawek, W., & Siwicki, A. K. (2017). Phenotypic and genotypic antimicrobial susceptibility pattern of Streptococcus spp. isolated from cases of clinical mastitis in dairy cattle in Poland. Journal of Dairy Science, 100(8), 6442–6453. https://doi.org/10.3168/jds.2017-12660
Klein, E. Y., Van Boeckel, T. P., Martinez, E. M., Pant, S., Gandra, S., Levin, S. A., Goossens, H., & Laxminarayan, R. (2018). Global increase and geographic convergence in antibiotic consumption between 2000 and 2015. Proceedings of the National Academy of Sciences of the United States of America, 115(15), E3463–E3470. https://doi.org/10.1073/pnas.1717295115
Li, W. C. (2014). Occurrence, sources, and fate of pharmaceuticals in aquatic environment and soil. Environmental Pollution, 187, 193–201. https://doi.org/10.1016/j.envpol.2014.01.015
Lin, X., Ruan, J., Huang, L., Zhao, J., & Xu, Y. (2021). Comparison of the elimination effectiveness of tetracycline and AmpC β-lactamase resistance genes in a municipal wastewater treatment plant using four parallel processes. Ecotoxicology, 30(8), 1586–1597. https://doi.org/10.1007/s10646-020-02306-0
Magiorakos, A. P., Srinivasan, A., Carey, R. B., Carmeli, Y., Falagas, M. E., Giske, C. G., Harbarth, S., Hindler, J. F., Kahlmeter, G., Olsson-Liljequist, B., Paterson, D. L., Rice, L. B., Stelling, J., Struelens, M. J., Vatopoulos, A., Weber, J. T., & M. D. (2012). Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clinical Microbiology and Infection, 18(3), 268–281.
Mishra, B. K., Regmi, R. K., Masago, Y., Fukushi, K., Kumar, P., & Saraswat, C. (2017). Assessment of Bagmati River pollution in Kathmandu Valley: Scenario-based modeling and analysis for sustainable urban development. Sustainability of Water Quality and Ecology, 9, 67–77. https://doi.org/10.1016/j.swaqe.2017.06.001
Mohameda, H. S. A., & U. M. and R. H. (2018). Correlation between antibiotic concentrations and antibiotic resistance genes contamination at Mafisa wastewater treatment plant in Morogoro Municipality, Tanzania. Global Environment, Health and Safety, 2, 1–7.
Pantha, K., Acharya, K., Mohapatra, S., Khanal, S., Amatya, N., Ospina-Betancourth, C., Butte, G., Shrestha, S. D., Rajbhandari, P., & Werner, D. (2021). Faecal pollution source tracking in the holy Bagmati River by portable 16S rRNA gene sequencing. Npj Clean. Water, 4(1). https://doi.org/10.1038/s41545-021-00099-1
Shallcross, L. J., Howard, S. J., Fowler, T., & Davies, S. C. (2015). Tackling the threat of antimicrobial resistance: From policy to sustainable action. Philosophical Transactions of the Royal Society B: Biological Sciences, 370(1670). https://doi.org/10.1098/rstb.2014.0082
Shenoy, A. K., Jyothi, E. K., & Ravikumar, R. (2014). Phenotypic identification & molecular detection of blandm-1 gene in multidrug resistant gram-negative Bacilli in a tertiary care center. The Indian Journal of Medical Research, 139(APR), 625–631.
Shrestha, R. G., Tandukar, S., Bhandari, D., Sherchan, S. P., Tanaka, Y., Sherchand, J. B., & Haramoto, E. (2019). Prevalence of Arcobacter and other pathogenic bacteria in River water in Nepal. Water, 11(7), 13–16. https://doi.org/10.3390/w11071416
Shrestha, S., Aihara, Y., Bhattarai, A. P., Bista, N., Rajbhandari, S., Kondo, N., Kazama, F., Nishida, K., & Shindo, J. (2017). Dynamics of domestic water consumption in the urban area of the Kathmandu valley: Situation analysis pre and post 2015 Gorkha Earthquake. Water, 9(3), 1–17. https://doi.org/10.3390/w9030222
Shrestha, S., Shrestha, S., Shindo, J., Sherchand, J. B., & Haramoto, E. (2018). Virological quality of irrigation water sources and pepper mild mottle virus and tobacco mosaic virus as index of pathogenic virus contamination level. Food and Environmental Virology, 10(1), 107–120. https://doi.org/10.1007/s12560-017-9324-2
Thakali, O., Tandukar, S., Brooks, J. P., Sherchan, S. P., Sherchand, J. B., & Haramoto, E. (2020). The occurrence of antibiotic resistance genes in an urban river in Nepal. Water, 12(2), 1–7. https://doi.org/10.3390/w12020450
Yuan, Q. B., Guo, M. T., & Yang, J. (2014). Monitoring and assessing the impact of wastewater treatment on release of both antibiotic-resistant bacteria and their typical genes in a Chinese municipal wastewater treatment plant. Environmental Sciences: Processes and Impacts, 16(8), 1930–1937. https://doi.org/10.1039/c4em00208c
Zar, J. H. (1998). Biostatistical Analysis (4th ed.). Prentice Hall.
Zhang, X., Wu, B., Zhang, Y., Zhang, T., Yang, L., Fang, H. H. P., Ford, T., & Cheng, S. (2009). Class 1 integronase gene and tetracycline resistance genes tetA and tetC in different water environments of Jiangsu Province, China. Ecotoxicology, 18(6), 652–660. https://doi.org/10.1007/s10646-009-0332-3
Acknowledgements
We are grateful to Nepal Academy of Science and Technology, Lalitpur, Nepal for providing laboratory facilities during this research. This study was partially supported with a Master’s thesis grant by National Youth Council (NYC). This research work was partially supported with the aid of a grant from UNESCO and the International Development Research Center (IDRC), Ottawa, Canada. The views expressed herein do not necessarily represent those of UNESCO, IDRC, or its Board of Governors.
Author information
Authors and Affiliations
Contributions
Conceptualization of the study by P. S, D. R. J, and T. P.J; Lab work, P. S., S. N. K. S.; Data analysis, P. S, T. P. J; writing-original draft preparation, P. S, D. R. J T. P.J and P. S review and editing; D. R. J, T.P.J, J. M, and R. R; supervision: D. R. J and T. P. J. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Institutional Review Board Statement
Not applicable
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shrestha, P., Prasai Joshi, T., Nhemhaphuki, S. et al. Occurrence of Antibiotic-Resistant Bacteria and Their Genes in Bagmati River, Nepal. Water Air Soil Pollut 234, 475 (2023). https://doi.org/10.1007/s11270-023-06499-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11270-023-06499-y