Abstract
Post-translational modifications (PTMs), as epigenetic modifications, are significant in the interaction between virus and its host. However, it is unclear whether rotavirus (RV) causes changes in both the host cell epigenetic protein modification and the regulatory mechanism of viral replication. Here, we analyzed the proteome of Caco-2 cells to determine if acetylation modification occurred within the cells after RV infection. We found that glyceraldehyde-3-phosphate dehydrogenase (GAPDH), a protein involved in glycolysis, was deacetylated at lysine 219 via histone deacetylase 9 (HDAC9) in 50 h after the RV infection. Remarkably, the deacetylation of GAPDH promoted RV replication. Finally, we found that glycolysis was alterable in Caco-2 cells by RV or the deacetylation of GAPDH lysine 219, using the Seahorse XF Glycolysis Stress Test. In conclusion, our results demonstrate for the first time that RV infection promoted deacetylation of GAPDH at lysine 219 in order to increase its own viral replication in Caco-2 cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Post-translational modifications (PTMs), as epigenetic modifications, are central to increasing the functional diversity of the cellular proteome, which plays a crucial role in the function of proteins in health and disease processes [1]. One of the most common PTMs in eukaryotes is protein acetylation. This transfer of an acetyl group neutralizes the positive electrostatic charge on the protein, thereby inhibiting electrostatic interactions with other amino acids and changing the biological function of the protein in the cell [2].
In some cases, viruses are an obligate parasite, and their life cycle depends on the successful facilitation of host cell acetylation in order to promote viral replication and restrain the abilities of the host immune system [3]. Both the hepatitis C virus and vesicular stomatitis virus have been shown to induce rapid deacetylation of RIG-I, modifying the acetylation status of RIG-I at lysine 909 (K909), and in turn, altering the immune response [4,5,6]. Furthermore, human immunodeficiency virus protein Tat increases the acetylation level of host histones and causes changes in chromatin structure from transcription inactivity to transcription activity, which is the region required for the viral genetic material to insert into the host genome for replication. Nonetheless, the underlying mechanisms of many known epigenetic protein modifications caused by viruses hijacking the host machinery for their own benefit are still unclear. This is especially true for the rotavirus (RV)-induced viroplasms, which rely on acetylated stable microtubule networks to fuse to the perinuclear replication point for replication [7].
Interestingly, the cellular metabolic pathways under acetylation regulation are always associated with viral survival. Dengue virus infection has been shown to relocate and aggregate the NS1 protein at the perinuclear area, which causes increased glyceraldehyde-3-phosphate dehydrogenase (GAPDH) activity and regulation of cellular energy metabolism [8]. Additionally, the upregulation of histone deacetylase 9 (HDAC9) has been reported to be utilized by the hepatitis C virus to promote the expression of gluconeogenesis genes to enhance gluconeogenesis [9]. There are limited studies on the ability of RV to facilitate host cell acetylation regulation for its own survival. Our lab previously demonstrated that RV could hinder the glucose consumption of host cells to promote gluconeogenesis [10]. Thus, we were interested in determining the epigenetic profile of the host cell after RV infection and identifying the biological functions, which were affected by RV infection, of the host cell.
Here, we determined the characteristics and effects of acetylated proteins in Caco-2 cells infected with RV via acetylation modification proteome quantitative research. We found that the host cells were deacetylated at lysine 219 of GAPDH (GAPDH K219) by HDAC9 in 50 h after the RV infection. Furthermore, by altering the acetylation status of GAPDH at K219, thereby promoting its own replication. RV also simultaneously altered glycolysis. This discovery revealed the hitherto unknown protein acetylation as a trigger mechanism for RV replication, which highlights the importance of acetylation for the replication of RV.
Results
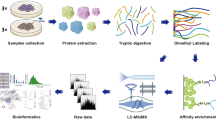
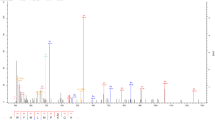
RVs induced the deacetylation of glycolytic proteins in Caco-2 cells. To confirm that deacetylation of GAPDH at lysine 219 occurs universally in different RV-strains-infected cells,we repeated these experiments using two rotavirus strains (Wa strain, SA-11 strain). We found that the SA-11-RV and Wa-RV groups both downregulated acetylation in Caco-2 cells compared to the control group, but the acetylation was somewhat more pronounced in the Wa-RV group (Fig. 1A). In order to determine the effect of RV infection on host cell protein modification, we first carried out acetylome profiling on RVs-infected Caco-2 cells to discover the proteins and their sites with altered acetylation, using non-standard quantitative, acetylation modification enrichment technology and high-resolution liquid chromatography-mass spectrometry (LC–MS) quantitative proteomics research strategy. We identified 1533 acetylation modification sites located on 821 proteins, which could be used in future experiments for normalization. The above data were screened by using a 1.5-fold change threshold at the modification site as the screening criterion. We found 220 sites with increased acetylation levels and 141 sites with decreased acetylation levels after RV infection. The majority of the proteins were distributed within the cytoplasm, which accounted for 42.08% of the proteins in the cell. Interestingly, the functional enrichment analysis of the sites at which the acetylation level was downregulated revealed that a total of 17 signaling pathways related to glycometabolism, including glycolysis/gluconeogenesis (hsa00010), were statistically significant. To further explore how the acetylation modification induced by RV influenced glycometabolism, we found that the hsa00010 glycolysis/gluconeogenesis pathway was associated with six proteins: glyceraldehyde-3-phosphate dehydrogenase (GAPDH, Fig. 1B), dihydrolipoamide dehydrogenase (DLD, Fig. 1C), phosphoglycerate mutase 1 (PGAM1, Fig. 1D), enolase 1 (ENO1, Fig. 1E), dihydrolipoamide S-acetyltransferase (DLAT, Fig. 1F), and pyruvate dehydrogenase E1 subunit beta (PDHB, Fig. 1G). Importantly, GAPDH is considered a housekeeping gene and plays an essential role in the physiological activities of the cell, thus explaining the close relationship between viruses, glycolysis, and GAPDH. Given this, we decided to further investigate the effect of RV-decreased acetylation on GAPDH during RV infection in Caco-2 cells.
The acetylation level of glycolysis-related proteins in RV-infected Caco-2 cells was downregulated. A. Reduced acetylation levels in Wa-RV and SA-RV-infected Caco-2 cells. B-G. In RV-infected Caco-2 cells: the MS/MS spectrum of modified ‘‘AVGK(Acetyl)VIPELNGK” at GAPDH, ‘‘ALLNNSHYYHMAHGK(Acetyl)DFASR” at DLD, ‘‘ETAAK(Acetyl)HGEAQVK” at PGAM1, ‘‘SCNCLLLK(Acetyl)VNQIGSVTESLQACK” at ENO1, ‘‘K(Acetyl)YLEKPITMLL” at DLAT, ‘‘VFLLGEEVAQYDGAYK(Acetyl)VS” at PDHB. (X-axis title is ‘‘Intensity” and y-axis title is ‘‘Abundance”)
RVs decreased acetylation of GAPDH at K219
We immunoprecipitated endogenous GAPDH from RV-infected Caco-2 cells and showed by western blotting with an anti-acetyl lysine antibody that RV strongly induced GAPDH decreased acetylation (Fig. 2A), which was consistent with our mass spectrometry results. In addition, the only modified residue identified on GAPDH from RV-infected Caco-2 cells was acetylated K219, via determination of modification sites by collision-induced dissociation (CID) analysis and the MS/MS spectrum of modified ‘‘AVGK(Acetyl)VIPELNGK” (Fig. 1A). We next aimed to address the potential association between K219 downregulated acetylation and RV infection using an anti-acetyl lysine-GAPDH K219 antibody. It was deemed preferable to permit Caco-2 cells to generate the GAPDH-K219 mutation (K219R) by means of the knockout of the endogenous GAPDH and the subsequent transfection of the K219R plasmid, which may result in the replacement of the original GAPDH by K219R GAPDH. As shown in Fig. 2B, C, we manipulated GAPDH levels in Caco-2 cells by si-GAPDH and K219R plasmid transfection, obtaining approximately 40% knockdown of the original GAPDH, as measured by qPCR and western blot after si-GAPDH transfection. However, under these conditions, the K219R GAPDH was only overexpressed approximately 50% as compared with the si-NC1 group after K219R plasmid transfection. Furthermore, we surprisingly found that while the GAPDH protein level in the K219R group was not statistically different from the si-NC1 group, the acetylation level of GAPDH in the K219R group was decreased, which confirmed that the GAPDH-K219 mutation model was feasible. Importantly, RV-infected Caco-2 cells maintained strong GAPDH downregulated acetylation, as detected with an anti-acetyl lysine-GAPDH K219 antibody, indicating that RV caused GAPDH downregulated acetylation at K219 (Fig. 2D). We next sought to determine the underlying mechanism of GAPDH decreased acetylation at K219.
The acetylation level of GAPDH at lysine 219 (GAPDH K219) was reduced in RV-infected Caco-2 cells. A The glycolysis-related protein, GAPDH, was deacetylated in RV-infected Caco-2 cells (48 h after RV infection). B Protein expression of GAPDH after transfection of si-GAPDH and plasmid K219R (without RV infection). C RNA expression of GAPDH after transfection of GAPDH siRNA (si-GAPDH) and plasmid K219R (K219 mutant GAPDH) (without RV infection). D The level of acetylation of GAPDH at K219 appears to be reduced following transfection with a K219 mutant or RV infection. (After the parallel operation, the RV group that treated like Control before, was infected with RV for 48 h, and the other groups were only treated with DMEM). (The transfection of si-GAPDH and plasmid K219 caused the host GAPDH to be replaced by GAPDH with a K219 mutation, which resulted in a lower acetylation level; *P < 0.05, **P < 0.01, and ***P < 0.001; n = 3)
HDAC9 catalyzed the deacetylation of GAPDH K219 and promoted RV replication
The main cause of protein deacetylation is that either the catalyst is a deacetylase or there is a lack of the raw material, acetyl-CoA. During RV infection, we investigated the function of enzymes in the HDAC and Sirt family, which can deacetylate GAPDH. We found that the RNA expression of HDAC9 in RV-infected Caco-2 cells was increased 1.5 times as compared with the control group (P < 0.05) (Fig. 3A). Additionally, as the duration of RV infection increased, the protein expression of HDAC9 also increased, with the levels of HDAC9 increasing up to almost 4 times that in the control group within 48 h (Fig. 3B). We then immunoprecipitated endogenous GAPDH from RV-infected Caco-2 cells and showed via western blotting that GAPDH interacted with HDAC9 (Fig. 3C). Therefore, HDAC9 is most likely an important enzyme that regulates the acetylation status of GAPDH at K219 during RV infection. In order to determine the ability of HDAC9 to modify GAPDH at K219, we knocked down the expression of HDAC9 in Caco-2 cells in order to test its impact on GAPDH K219 deacetylation. As shown in Fig. 3D, the acetylation level of GAPDH at K219 in the si-HDAC9 group was increased by about 75% during RV infection as compared with the si-NC2 group that was transfected with empty siRNAs, indicating that HDAC9 is responsible for the deacetylation of GAPDH at K219 during RV infection. Consistent with the si-NC2 group, RV-infected Caco-2 cells transfected with si-HDAC9 exhibited a decrease (approximately 40%) in the levels of HDAC9 RNA. We also explored the effect of HDAC9 on RV replication. We showed that the RNA expression of a constitutively expressed protein from RV, VP6, was reduced by approximately 60% in RV-infected Caco-2 cells transfected with si-HDAC9 (Fig. 3E). We confirmed via western blotting that VP6 was also downregulated at the protein level in RV-infected Caco-2 cells transfected with si-HDAC9, further indicating that inhibition of HDAC9 expression limits the replication of RV replication (Fig. 3F).
Upregulated expression of histone deacetylase 9 (HDAC9) not only catalyzed the deacetylation of GAPDH at K219 but also promoted RV replication in RV-infected Caco-2 cells. A HDAC9 RNA expression was upregulated among several deacetylases (HDAC 1–9 and Sirt 1–4,6) in RV-infected Caco-2 cells (48 h after RV infection). B HDAC9 protein expression increased as the duration of RV infection increased. C The interaction between HDAC9 and GAPDH in RV-infected Caco-2 cells was detected by co-immunoprecipitation (48 h after RV infection). D Transfection with HDAC9 siRNA (si-HDAC9) relieved GAPDH K219 deacetylation, which indicated that HDAC9 was responsible for the deacetylation of GAPDH K219 in RV-infected Caco-2 cells (After parallel operation, RV, si-NC2 and si-HDAC9 groups were all infected with RV for 48 h). E Transfection with si-HDAC9 downregulated VP6 RNA expression. F Transfection with si-HDAC9 downregulated VP6 protein expression (After parallel operation, RV, si-NC2 and si-HDAC9 groups were all infected with RV for 48 h). (Figures E, F show that HDAC9 promoted the replication of RVs; *P < 0.05, **P < 0.01, and ***P < 0.001; n = 3)
The deacetylation of GAPDH at K219 promoted RV replication and altered glycolysis
Having demonstrated that K219 was deacetylated by HDAC9, we further studied the impact of GAPDH deacetylation at K219 on RV replication. We first assessed the expression of VP6 in RV-infected Caco-2 cells that had a mutated form of GAPDH K219R and found that the RNA expression of VP6 in the K219R group was increased by almost 2.5 times as compared with the si-NC1 group. Intriguingly, GAPDH in the si-GAPDH group without mutation but with the knock down and overall expression level downregulated may leads to the downregulation of acetylation level at the K219 site in total. Because we found that the RNA expression of VP6 increased, as well as the protein level of VP6. (Fig. 4B). Together, these results demonstrated that the deacetylation of GAPDH at K219 promotes the replication of RV (Fig. 4A).
RV promoted the deacetylation of GAPDH at K219, which resulted in the promotion of RV replication and the alteration of glycolysis. A RNA expression of VP6 after transfection of si-GAPDH and K219 mutant GAPDH (After the parallel operation, the RV group that treated like Control before, si-NC1, si-GAPDH and K219R groups were infected with RV for 48 h). B Protein expression of VP6 after transfection of si-GPADH and plasmid K219R (a). C The GAPDH K219 mutation inhibited the upregulation of glycolysis in RV-infected Caco-2 cells (a). (The transfection of si-GPADH and plasmid K219 caused the host GAPDH to be replaced by GAPDH with a K219 mutation, which resulted in a lower acetylation level; *P < 0.05, **P < 0.01, and ***P < 0.001; n = 3)
GAPDH is a central enzyme within the glycolytic pathway. As such, we were interested in understanding how deacetylation of GAPDH significantly influences RV replication. Moreover, we sought to validate the relationship between glycolysis and GAPDH K219 deacetylation (Fig. 4C). We first measured the glycolysis index extracellular acidification rate (ECAR) via a Seahorse XF analyzer, and we discovered that the ECAR was improved in RV-infected Caco-2 cells saturated with glucose. The maximum ECAR of RV-infected cells after adding oligomycin also increased (glycolysis capability); however, there was no statistical difference in the ability of RV-infected cells to respond to energy requirements and the theoretical maximum of glycolysis (glycolytic reserve). In general, deacetylation of GAPDH at K219 in RV-infected Caco-2 cells enhanced glycolysis. Additionally, the degree of deacetylation of GAPDH at K219 increased, and the increase in glycolysis was halted after mutating basal GAPDH to GAPDH K219R. After GAPDH K219 deacetylation was further strengthened, the amount of RV replication increased. Therefore, we hypothesized that glycolysis in Caco-2 cells was enhanced during RV infection due to the increase in the deacetylation of GAPDH at K219. If the degree of deacetylation of K219 of GAPDH at K219 was further strengthened, the metabolic energy substances from further enhanced glycolysis would also further increase and promote the replication of RV itself, but negative feedback regulation was then produced, which destroyed the balance of glycolysis in host cells and retarded the increase in glycolysis level.
Discussion
We demonstrated here for the first time that RV regulates the acetylation status of host proteins, especially GAPDH at K219, in RV-infected Caco-2 cells. We found no significant difference in the levels of succinylation, crotonylation, 2-hydroxyisobutylation, and malonylation in RV-infected Caco-2 cells. By combining mass spectrometry and bioinformatics analysis, we identified that GAPDH, a glycolytic pathway protein and housekeeping gene, which is stably expressed at a high level in a complex environment [11, 12], was deacetylated after RV infection in Caco-2 cells. However, although there are abundant signaling pathways enriched in acetylated proteins, the raw materials for acetylation were almost originated from metabolic intermediates with different functions and effects, suggesting glycolysis pathways [13, 14]. Enzymes in glycolysis metabolism are in different acetylated states based on the concentrations of metabolites [15]. Interestingly enough, acetylation can affect virus invasion and host defense when cells or viruses face different nutrients and stress states [16].
Previous studies have demonstrated the association of RV with host cell metabolism. For examples, RV has been shown to change the paracellular permeability of Caco-2 cells, causing increased lactic acid production, decreased mitochondrial oxygen consumption, and decreased cellular ATP [17, 18]. Here, we investigated the relationship between acetylation changes, metabolism, and RV infection. We showed that GAPDH was deacetylated at K219 during RV infection, which significantly promoted RV replication. GAPDH is regarded as the “switching station” of metabolism, as its catalytic capacity is necessary for cells to maintain adequate glycolytic flux [19, 20]. Reversible PTMs of GAPDH, especially acetylation, contribute to the mutual regulation between glycolysis and gluconeogenesis. It was reported that the acetylation of GAPDH at lysine 254 can increase the glycolytic activity of GAPDH, thereby providing support for cell proliferation and tumor growth [21, 22]. A previous study demonstrated that acetylation of GAPDH catalyzed glycolysis and strengthened the response of fast memory CD8 (+) T cells for a better immune control function [23]. Thus, we presumed that GAPDH deacetylation at K219 may also play a role in the systemic metabolism of the host, specifically influencing the immune system. Nonetheless, there are limited studies on the relationship between GAPDH and viral pathogenesis. Our study highlights the importance of GAPDH in host cell metabolism and RV pathogenesis.
The deacetylation of GAPDH at K219 during RV infection was dependent upon HDAC9. It has been reported that HDAC9 interacts with metalloproteinase 3, HDAC6, and NKX3.2 [24,25,26], but our study is the first to show an interaction between HDAC9 and GAPDH. HDAC9 is a negative regulator of adipogenic differentiation, while GAPDH is a key enzyme of glycolysis, both of which have potential link to glucose and lipid metabolism implying that HDAC9 and GAPDH may interact through glycolipid metabolism [27]. Considering that the knockdown of HDAC9 resulted in reduced replication of RV in our study, we established a hypothesis that HDAC9 controls RV replication by regulating glycolysis via deacetylation of GAPDH at K219. Similarly, HDAC9 was shown to deacetylate FoxO1 to regulate hepatic gluconeogenesis during hepatitis C virus infection [28]. In addition, there is a report about the high expression of HDAC9 in the macrophages, which keeps the deacetylation’s condition of the proteins, triggers the production of type I interferon, and strengthens the host against the virus attack [29]. Thus, further research is needed to fully understand the connection between HDAC9 and RV.
In our study, deacetylation of GAPDH at K219 was associated with the glycolysis and, as such, was key to the anti-RV defense of the host and RV replication. Consistent with our study showing RV regulation of host cell glycolysis, recent studies have shown that Zika virus, Semliki forest virus, and avian reovirus control the innate antiviral response and self-replication through virus-induced glycolytic changes [30,31,32]. However, these studies regarding the virus-induced glycolytic changes did not further explore in the direction of PTMs that are very significant to adjust the protein activity and function of the cellular system. Our study shows that RV modifies host protein deacetylation to upregulate glycolysis during the latency period in order to select the best time and space for replication and propagation. We demonstrated that RV infection enhanced host cell glycolysis, which allowed RV to affect glucose metabolism in order to provide energy for its own replication. However, we also encountered limitations. In the case of RV infection, further GAPDH K219 deacetylation does not promote, but rather limits, the continued increased in glycolysis, which remains to be elucidated. It has been reported that excessive glycolysis and tricarboxylic acid cycle substrates, such as acetyl-CoA or citrate, and high ATP/ADP ratios can regulate the activity of phosphofructokinase-1 by inhibiting the activity of phosphofructokinase-1, thereby initiating a sugar glycolysis negative feedback regulation loop [33]. In addition, the Warburg effect in cancer cells and the Crabtree effect in yeast are also examples for the study of this negative feedback regulation [34]. In addition, viruses have been observed to interfere with the expression of cellular molecules such as insulin, hypoxia-inducible factor-1, p53, nitric oxide, and acetylated histones. Furthermore, the function of GAPDH can be affected by these substances at high levels of constant expression. Alterations in acetylation levels in response to these influences have been observed to affect not only glycolysis but also apoptosis itself. It is therefore of interest to explore whether this has a side effect on the host cell or the virus. It would be beneficial to investigate whether there are any indirect effects on host cells or viruses [35].It might be worth exploring incidence and biological significance of GAPDH deacetylation during RV infection remain to be tested in humans and animal models.
In conclusion, we showed for the first time that increased HDAC9 expression in RV-infected Caco-2 cells results in increased deacetylation of GAPDH at K219, thereby enhancing RV replication and altering host cell glycolysis (Fig. 5).The limitation of this study is that it was conducted at a single time after infection, that is, 50 h after infection, when Caco2 showed cytopathic effect, severe shedding, and morphological changes of cell boundary paste, which could probe into the protein expression of Caco-2 cells by RV to the maximum extent. However, this experiment did not explore the process changes at other time points. Thus the effect of RV infection in incubation period or the time and space of replication cannot be inferred. The results of this study provide novel insights into the relationship between RV replication and host cell PTMs, specifically highlighting how RV hijacks the cell’s metabolism for its own benefit. Furthermore, this study sheds light onto host proteins that have the potential to be therapeutically targeted to limit RV infection.
A schematic diagram illustrating how RV induced glycolysis in Caco-2 cells to promote self-replication. RV infection upregulated HDAC9, resulting in increased deacetylation of GAPDH at K219, which resulted in the alteration of host cell glycolysis and increased RV replication. AVGKVIP, which represents alanine, valine, glycine, lysine, valine, isoleucine, and proline, correspond to amino acid residues 216–222 of GAPDH
Methods
Reagents
RV Wa strain and RV SA strain were obtained from the Institute of Immunology, Third Military Medical University (Chongqing, China). The Caco-2 cell line was obtained from Wuhan University Cell Bank (Wuhan, China). Protein A/G PLUS-agarose was purchased from Santa Cruz Biotechnology (Shanghai, China). The SDS-PAGE gel preparation kit and BCA kit were obtained from Solarbio Inc (Beijing, China). Lipofectamine 3000 transfection reagent and TRIzol reagent were obtained from Invitrogen (Shanghai, China). RiboFECT CP Transfection Kit was purchased from Guangzhou RiboBio (Guangzhou, China). Evo M-MLV RT Kit with gDNA Clean for qPCR II and SYBR Green Premix Pro Taq HS qPCR Kit were obtained from Aikerui Bio (Guangzhou, China). Anti-GAPDH (60,004-1-Ig), anti- VP6 (NB110-37243), and anti-VP6 (sc-101363) antibodies were obtained from Proteintech Group (Wuhan, China), Novus Biologicals (Shanghai, China), and Santa Cruz Biotechnology (Shanghai, China), respectively. Anti-β-actin (AF0003), anti-HDAC9 (AF7074), human IgG (A7001), HRP-labeled goat anti-mouse IgG (A0216), and HRP-labeled goat anti-rabbit IgG (A0208) were obtained from Beyotime Bio (Shanghai, China). Anti-acetyl lysine (PTM-101) and anti-acetyl lysine-GAPDH K219 (CK70601) were obtained from Jingjie Biotechnology Co (Hangzhou, China). All other reagents, unless otherwise specified, were obtained from Sigma (Shanghai, China).
Cell culture, virus amplification, and titer determination
Caco-2 cells or MA104 cells were cultured in DMEM with 10% fetal bovine serum and 1% penicillin/streptomycin at 37 °C, 5% CO2. The cell viability was assayed by 3-(4,5)-dimethylthiahiazol (-z-y1)-3,5-di-phenytetrazoliumromide (MTT) or was assessed by 0.1% crystal violet solution after fixing in 4% paraformaldehyde (Solarbio, Beijing, China). The medium was replaced with 3 mL of DMEM and 500 μL of RV Wa strain, which was incubated with 500 μL of 10-μg/mL trypsin for 30 min prior to being added to the media, until the confluence of the MA104 cells reached 70%. Once the cells exhibited cytopathic effect (CPE; + + +), they were frozen and thawed three times. The cells were then centrifuged for 30 min at 4 °C at 12,000 × g, and the supernatant was collected. The supernatant was then incubated in the same volume with 10-μg/mL trypsin for 30 min, diluted with a tenfold gradient, and added to a 96-well plate (with 6000 MA104 cells/condition) for virus titer determination. Once the cells with the highest RV dilution ratio no longer produced CPE, MTT was used to assess cell survival, and the TCID50 of the RV strain was calculated by the Reed-Muench method (TCID50: median tissue culture infective dose, half of the tissue culture infective dose, also known as 50% tissue cell infection).
Acetylation mass spectrometry
Caco-2 cells were cultured in a 10-cm petri dish at a density of 1 × 106 cells/mL until the confluence reached 60–70%. The medium was replaced with 7 mL of DMEM medium containing 1 μg/mL trypsin. The control group was not infected with the virus, but the RV group was infected with 70 μL of Wa strain RV, which was incubated at a virus titer of 104 TCID50/mL with 10-μg/mL trypsin for 30 min prior to infection. After incubating for 2 h at 37 °C, the medium was replaced with 2 mL of DMEM medium containing 1-μg/mL trypsin and then incubated for another 48 h at 37 °C. The samples were lysed in 8-M urea with 1% protease inhibitors, 3-μM toluenesulfonic acid, and 50-mM niacinamide, and the cell debris were removed by centrifuging at 20,000 × g at 4 °C for 10 min. The protein concentration was measured using a BCA protein assay. The protein concentration was normalized across all samples, and the volume of each sample was then brought up to 1000 μL with 8-M urea. The samples were then incubated with trichloroacetic acid at a final concentration of 20% for 2 h. The precipitate was ultrasonically dispersed with 1 mL of 200-mM triethylammonium bicarbonate, followed by an 18-h trypsin digestion at a ratio of 1:50 (protease: protein, m/m) at 37 °C. Tryptic peptides were then dissolved in IP buffer solution (100-mM NaCl, 1-mM EDTA, 50-mM Tris–HCl, 0.5% NP-40, pH 8.0) and incubated with a pre-washed acetylated resin (PTM-104; Jingjie Biotechnology Co (Hangzhou, China) at 4 °C for 18 h. The resin was washed four times with IP buffer and twice with ddH2O. Bound peptides were eluted from the resin with 0.1% trifluoroacetic acid eluent, collected with C18 ZipTips following the manufacturer’s instructions, and analyzed by liquid chromatography-mass spectrometry/mass spectrometry. Peptides were dissolved in the mobile phase A (aqueous solution containing 0.1% formic acid and 2% acetonitrile) and separated using an EASY-nLC 1200 ultra-high performance liquid system (Thermo Fisher Scientific, Massachusetts, USA) at 400 nL/min in a gradient of 9–25% mobile phase B (aqueous solution containing 0.1% formic acid and 90% acetonitrile) for 36 min, followed by 25% B to 35% B for 18 min, 35% B to 80% B for 3 min, and finishing at 80% B for 4 min. The peptides were separated via an ultra-high performance liquid system (Waters, Massachusetts, USA), injected into the Nano-liquid chromatography (nLC)–nanoelectrospray ion source for ionization, and then analyzed by Q Exactive HF-X mass spectrometry (Thermo Fisher Scientific, Massachusetts, USA). The scanning range of the primary mass spectrum was set to 350–1600 m/z, and the scanning resolution was set to 120,000. The scanning range of the secondary mass spectrum was set to a fixed starting point of 100 m/z, and the secondary scanning resolution was set to 15,000. The resulting secondary mass spectrum data were processed using Maxquant (v1.5.2.8, URL: https://www.maxquant.org/) with the database Homo_sapiens_9606_SP_20191115 (20,380 sequences). Trypsin was specified as the cleavage enzyme, allowing up to four missed cleavages. The main search range was set to 5 ppm and 0.02 Da for fragment ions. Cysteine alkylation was specified as a fixed modification, and oxidation of methionine, acetylation of protein N-terminus, deamidation (NQ), and acetylation of lysine were specified as variable modifications.
The sites where acetylation modification levels were downregulated were subjected to functional enrichment analysis, resulting in the identification of 17 statistically significant signaling pathways, including hsa00010 Glycolysis/Gluconeogenesis, which is related to gluconeogenesis (Fig. 6). To investigate the impact of the altered acetylation modifications induced by the WA strain RV on the gluconeogenesis pathway, a network was constructed of the involved differential sites and signaling pathways. This analysis revealed that the hsa00010 Glycolysis/Gluconeogenesis pathway was associated with glyceraldehyde-3-phosphate dehydrogenase (GAPDH). The following proteins were identified as being associated with the glycolysis/gluconeogenesis pathway: dihydrolipoamide dehydrogenase (DLD), phosphoglycerate mutase 1 (PGAM1), enolase 1 (ENO1), dihydrolipoamide S-acetyltransferase (DLD), and Glycolysis/Gluconeogenesis pathway. The remaining six proteins are acetyltransferase (DLAT) and pyruvate dehydrogenase E1 subunit beta (PDHB).
Protein immunoprecipitation and western blotting
A total of 6 × 105 cells/condition were lysed in 1 mL of radioimmunoprecipitation assay lysis buffer (RIPA) with 1% phenylmethanesulfonyl fluoride solution. The protein concentration was measured using a BCA protein assay. After normalizing the samples, the lysates were precleared with 20 μL of A/G beads for 30 min at 4 °C. For GAPDH immunoprecipitation, 5 μg of antibody was incubated with the lysate for 1 h at 4 °C and then the protein-antibody complex was coupled to 20 μL of A/G beads for another 18 h at 4 °C. The lysates were centrifuged for 15 min at 4 °C, and the beads were washed three times with RIPA buffer. Immune complexes were eluted by adding 40 μL of 1 × Laemmli sample buffer, followed by heating at 95 °C for 5 min. Protein samples were separated by 10% SDS–PAGE and transferred to polyvinylidene fluoride membranes via wet transfer. Membranes were probed with the respective antibodies and visualized using a chemiluminescent HRP substrate. All primary antibodies, β-actin, GAPDH, VP6, acetyl lysine, HDAC9, and acetyl lysine–GAPDH(K219) were used at a 1:1000 dilution. The secondary HRP-conjugated antibodies were also used at a 1:1000 dilution. The protein bands were quantified using Image J software (URL: https://imagej.en.softonic.com/).
RNA extraction, reverse-transcription PCR, and qPCR
RNA was extracted using the TRIzol method. After repeatedly pipetting Caco-2 cells (6 × 105 cells/condition) in 1 mL of TRIzol lysate, 200 μL of chloroform was added. The sample was vortexed for 10 s and then centrifuged for 15 min at 4 °C. Then, 400 μl of the supernatant was mixed with an equal volume of isopropanol and centrifuged for 10 min at 4 °C. The precipitate was washed twice with 1 mL of pre-cooled 75% ethanol and finally dissolved in 15-μL enzyme-free water. Next, the cDNA was synthesized using the Evo M-MLV RT Kit with gDNA Clean for qPCR II. The SYBR Green Premix Pro Taq HS qPCR Kit was used to determine the gene expression of GAPDH, VP6, and HDAC via qPCR. The following primers were used for qPCR (Table 1, purchased from Jierui Company, Shanghai, China):
siRNA and plasmids
GAPDH siRNAs (si-GAPDH) and HDAC9 siRNAs (si-HDAC9) together with negative controls, named si-NC1 and si-NC2, respectively, were obtained from Guangzhou RiboBio Co (Guangzhou, China). Caco-2 cells were seeded at 5 × 104 cells/mL and transfected with the siRNAs using 10-nM riboFECTTM CP Reagent (provided by RiboBio) for 24 h. The following target sequence was used to knock down human GAPDH: TAAAGTACCCTGTGCTCAA, and the following target sequence was used to KD human HDAC9: AGCCACCCTCATGTTACTT. For GAPDH K219R plasmids, the K219R sequence was synthesized into the GV230 vector with the component sequence CMV-MCS-EGFP-SV40-Neomycin (Jikai Gene Chemical Technology Co. Ltd, Shanghai, China), and then the entire plasmid was transformed into Escherichia coli competent cells. Positive clones were selected, and the plasmid was extracted by using the HiPure Plasmid Mini Kit (Magen Bio, Shanghai, China). The K219R plasmid was then transfected into Caco-2 cells seeded at a density of 5 × 104 cells/mL using 3.75 μL of Lipofectamine 3000 reagent and 5 μL of P3000 reagent for 12 h.
Glycolysis assay
Caco-2 cells that were transfected with si-GAPDH, si-HDAC9, or K219 plasmids were inoculated into XF24 microplates at 2.5 × 105 cells/mL, and RV infection was performed. Before sample processing, 1 mL of XF Calibrant was added to each well of the Utility Plate, which was then covered with the Hydro Booster and Sensor Cartridge and incubated in a non-CO2 incubator at 37 °C for 12 h. Next, 200-mM L-glutamine was added to the XF minimal medium at a ratio of 100:1 and adjusted to pH 7.4 with sodium hydroxide. Then, 100-mM glucose (56 μL), 10-µM oligomycin (62 μL), and 500-mM 2-deoxy-D-glucose (69 μL) were added to channels a, b, and c, respectively, of each well of the Sensor Cartridge. The plate was then loaded into the XFe24 analyzer for calibration. During this time, the cells were washed with XF minimal medium twice. Then, 500 μL of XF minimal medium was added into the well, and the plate was placed in a CO2-free incubator at 37 °C for 1 h. After the XFe24 analyzer finished calibrating the Sensor Cartridge, the Utility Plate was replaced with the microplate with the cells, and the protocol settings were entered into the computer for analysis. When the assay was finished, the microplate was removed and 50 μL of RIPA lysis buffer was added to each well. The protein concentration in each well was determined by a BCA kit (Agilent Seahorse, Guangzhou, China).
Statistical analysis
Statistical tests were performed using SPSS 13.0 statistical software (URL: https://www.ibm.com/analytics/spss-statistics-software). One-way ANOVA was used to compare the mean ± standard deviation (x ± s) of multiple samples, and a value of p < 0.05 was considered statistically significant.
Data Availability
The data presented in this study are available on request from the corresponding author.
References
Drazic A, Myklebust LM, Ree R, Arnesen T (2016) The world of protein acetylation. Biochim Biophys Acta 1864:1372–1401. https://doi.org/10.1016/j.bbapap.2016.06.007
Piperno G, Ledizet M, Chang XJ (1987) Microtubules containing acetylated alpha-tubulin in mammalian cells in culture. J Cell Biol 104:289–302. https://doi.org/10.1083/jcb.104.2.289
Murray LA, Sheng X, Cristea IM (2018) Orchestration of protein acetylation as a toggle for cellular defense and virus replication. Nat Commun 9:4967. https://doi.org/10.1038/s41467-018-07179-w
Choi SJ, Lee HC, Kim JH, Park SY, Kim TH, Lee WK, Jang DJ, Yoon JE, Choi YI, Kim S, Ma J, Kim CJ, Yao TP, Jung JU, Lee JY, Lee JS (2016) HDAC6 regulates cellular viral RNA sensing by deacetylation of RIG-I. EMBO J 35:429–442. https://doi.org/10.15252/embj.201592586
Liu HM, Jiang F, Loo YM, Hsu S, Hsiang TY, Marcotrigiano J, Gale MJ (2016) Regulation of retinoic acid inducible gene-I (RIG-I) activation by the histone deacetylase 6. EBioMedicine 9:195–206. https://doi.org/10.1016/j.ebiom.2016.06.015
Dutta D, Dutta S, Veettil MV, Roy A, Ansari MA, Iqbal J, Chikoti L, Kumar B, Johnson KE, Chandran B (2015) BRCA1 regulates IFI16 mediated nuclear innate sensing of herpes viral DNA and subsequent induction of the innate inflammasome and interferon-beta responses. PLoS Pathog 11:e1005030. https://doi.org/10.1371/journal.ppat.1005030
Eichwald C, Arnoldi F, Laimbacher AS, Schraner EM, Fraefel C, Wild P, Burrone OR, Ackermann M (2012) Rotavirus viroplasm fusion and perinuclear localization are dynamic processes requiring stabilized microtubules. PLoS ONE 7:e47947. https://doi.org/10.1371/journal.pone.0047947
Allonso D, Andrade IS, Conde JN, Coelho DR, Rocha DC, Da SM, Ventura GT, Silva EM, Mohana-Borges R (2015) Dengue virus NS1 protein modulates cellular energy metabolism by increasing glyceraldehyde-3-phosphate dehydrogenase activity. J Virol 89:11871–11883. https://doi.org/10.1128/JVI.01342-15
Chen J, Wang N, Dong M, Guo M, Zhao Y, Zhuo Z, Zhang C, Chi X, Pan Y, Jiang J, Tang H, Niu J, Yang D, Li Z, Han X, Wang Q, Chen X (2015) The metabolic regulator histone deacetylase 9 contributes to glucose homeostasis abnormality induced by hepatitis C virus infection. Diabetes. https://doi.org/10.2337/db15-0197
Song L, Zhong P, Zhu X, Zhou R, Gao M, Lan Q, Chen J, Chen Y, Zhao W (2021) The anti-rotavirus effect of baicalin via the gluconeogenesis-related p-JNK-PDK1-AKT-SIK2 signaling pathway. Eur J Pharmacol 897:173927. https://doi.org/10.1016/j.ejphar.2021.173927
Li L, Zhang W, Wang C, Wang H (2016) Mechanisms of glyceraldehyde 3-phosphosphate dehydrogenaseis in bacteria adhesion—a review. Wei Sheng Wu Xue Bao 56:1398–1405. https://doi.org/10.13343/j.cnki.wsxb.20150558
Zhang JY, Zhang F, Hong CQ, Giuliano AE, Cui XJ, Zhou GJ, Zhang GJ, Cui YK (2015) Critical protein GAPDH and its regulatory mechanisms in cancer cells. Cancer Biol Med 12:10–22. https://doi.org/10.7497/j.issn.2095-3941.2014.0019
Yin S, Liu L, Gan W (2021) The roles of post-translational modifications on mTOR signaling. Int J Mol Sci 22:1784. https://doi.org/10.3390/ijms22041784
Sirover MA (2021) The role of posttranslational modification in moonlighting glyceraldehyde 3 phosphate dehydrogenase structure and function. Amino Acids 53:507–515. https://doi.org/10.1007/s00726-021-02959-z
Zhao S, Xu W, Jiang W, Yu W, Lin Y, Zhang T, Yao J, Zhou L, Zeng Y, Li H, Li Y, Shi J, An W, Hancock SM, He F, Qin L, Chin J, Yang P, Chen X, Lei Q, Xiong Y, Guan KL (2010) Regulation of cellular metabolism by protein lysine acetylation. Science 327:1000–1004. https://doi.org/10.1126/science.1179689
Wang YP, Lei QY (2018) Regulation of cell metabolism by lysine acetylation. Chin Bull Life Sci 30:447–454. https://doi.org/10.13376/j.cbls/2018054
Dickman KG, Hempson SJ, Anderson J, Lippe S, Zhao L, Burakoff R, Shaw RD (2000) Rotavirus alters paracellular permeability and energy metabolism in Caco-2 cells. Am J Physiol Gastrointest Liver Physiol 279:G757–G766. https://doi.org/10.1152/ajpgi.2000.279.4.G757
Gaunt ER, Cheung W, Richards JE, Lever A, Desselberger U (2013) Inhibition of rotavirus replication by downregulation of fatty acid synthesis. J Gen Virol 94:1310–1317. https://doi.org/10.1099/vir.0.050146-0
Seidler NW (2013) Compartmentation of GAPDH. Adv Exp Med Biol 985:61–101. https://doi.org/10.1007/978-94-007-4716-6_3
Bond ST, Howlett KF, Kowalski GM, Mason S, Connor T, Cooper A, Streltsov V, Bruce CR, Walder KR, McGee SL (2017) Lysine post-translational modification of glyceraldehyde 3 phosphate dehydrogenase regulates hepatic and systemic metabolism. FASEB J 31:2592–2602. https://doi.org/10.1096/fj.201601215R
Krasnov GS, Dmitriev AA, Snezhkina AV, Kudryavtseva AV (2013) Deregulation of glycolysis in cancer: glyceraldehyde-3-phosphate dehydrogenase as a therapeutic target. Expert Opin Ther Targets 17:681–693. https://doi.org/10.1517/14728222.2013.775253
Li T, Liu M, Feng X, Wang Z, Das I, Xu Y, Zhou X, Sun Y, Guan KL, Xiong Y, Lei QY (2014) Glyceraldehyde-3-phosphate dehydrogenase is activated by lysine 254 acetylation in response to glucose signal. J Biol Chem 289:3775–3785. https://doi.org/10.1074/jbc.M113.531640
Balmer ML, Ma EH, Bantug GR, Grahlert J, Pfister S, Glatter T, Jauch A, Dimeloe S, Slack E, Dehio P, Krzyzaniak MA, King CG, Burgener AV, Fischer M, Develioglu L, Belle R, Recher M, Bonilla WV, Macpherson AJ, Hapfelmeier S, Jones RG, Hess C (2016) Memory CD8(+) T cells require increased concentrations of acetate induced by stress for optimal function. Immunity 44:1312–1324. https://doi.org/10.1016/j.immuni.2016.03.016
Xie D, Zhu J, Liu Q, Li J, Song M, Wang K, Zhou Q, Jia Y, Li T (2019) Dysregulation of HDAC9 represses trophoblast cell migration and invasion through TIMP3 activation in preeclampsia. Am J Hypertens 32:515–523. https://doi.org/10.1093/ajh/hpz006
Salian-Mehta S, Xu M, McKinsey TA, Tobet S, Wierman ME (2015) Novel interaction of class IIb histone deacetylase 6 (HDAC6) with class IIa HDAC9 controls gonadotropin releasing hormone (GnRH) Neuronal cell survival and movement. J Biol Chem 290:14045–14056. https://doi.org/10.1074/jbc.M115.640482
Choi HJ, Kwon S, Kim DW (2016) A post-translational modification cascade employing HDAC9-PIASy-RNF4 axis regulates chondrocyte hypertrophy by modulating Nkx3.2 protein stability. Cell Signal 28:1336–1348. https://doi.org/10.1016/j.cellsig.2016.06.006
Chatterjee TK, Idelman G, Blanco V, Blomkalns AL, Piegore MJ, Weintraub DS, Kumar S, Rajsheker S, Manka D, Rudich SM, Tang Y, Hui DY, Bassel-Duby R, Olson EN, Lingrel JB, Ho SM, Weintraub NL (2011) Histone deacetylase 9 is a negative regulator of adipogenic differentiation. J Biol Chem 286:27836–27847. https://doi.org/10.1074/jbc.M111.262964
Chen J, Zhang Z, Wang N, Guo M, Chi X, Pan Y, Jiang J, Niu J, Ksimu S, Li JZ, Chen X, Wang Q (2017) Role of HDAC9-FoxO1 axis in the transcriptional program associated with hepatic gluconeogenesis. Sci Rep 7:6102. https://doi.org/10.1038/s41598-017-06328-3
Li X, Zhang Q, Ding Y, Liu Y, Zhao D, Zhao K, Shen Q, Liu X, Zhu X, Li N, Cheng Z, Fan G, Wang Q, Cao X (2016) Methyltransferase Dnmt3a upregulates HDAC9 to deacetylate the kinase TBK1 for activation of antiviral innate immunity. Nat Immunol 17:806–815. https://doi.org/10.1038/ni.3464
Singh S, Singh PK, Suhail H, Arumugaswami V, Pellett PE, Giri S, Kumar A (2020) AMP-activated protein kinase restricts Zika virus replication in endothelial cells by potentiating innate antiviral responses and inhibiting glycolysis. J Immunol 204:1810–1824. https://doi.org/10.4049/jimmunol.1901310
Findlay JS, Ulaeto D (2015) Semliki Forest virus and Sindbis virus, but not vaccinia virus, require glycolysis for optimal replication. J Gen Virol 96:2693–2696. https://doi.org/10.1099/jgv.0.000226
Chi PI, Huang WR, Chiu HC, Li JY, Nielsen BL, Liu HJ (2018) Avian reovirus sigmaA-modulated suppression of lactate dehydrogenase and upregulation of glutaminolysis and the mTOC1/eIF4E/HIF-1alpha pathway to enhance glycolysis and the TCA cycle for virus replication. Cell Microbiol 20:e12946. https://doi.org/10.1111/cmi.12946
van den Brink J, Canelas AB, van Gulik WM, Pronk JT, Heijnen JJ, de Winde JH, Daran-Lapujade P (2008) Dynamics of glycolytic regulation during adaptation of Saccharomyces cerevisiae to fermentative metabolism. Appl Environ Microbiol 74:5710–5723. https://doi.org/10.1128/AEM.01121-08
Sokolov SS, Balakireva AV, Markova OV, Severin FF (2015) Negative feedback of glycolysis and oxidative phosphorylation: mechanisms of and reasons for it. Biochemistry (Mosc) 80:559–564. https://doi.org/10.1134/S0006297915050065
Zhang JY, Zhang F, Hong CQ et al (2015) Critical protein GAPDH and its regulatory mechanisms in cancer cells. Cancer Biol Med 12(1):10–22
Acknowledgements
This study was funded by National Natural Science Foundation of China (No. 81973548 and No. 81473401), Key Project of Social Science and Technology Development of Dongguan (NO. 20185071521658), Zhanjiang Science and Technology Planning Project (2021B01139), Guangdong Innovation Team Project for Regular Institutions of Higher Learning (2022KCXTD011) and Discipline construction project of Guangdong Medical University (No. 4SG22009G).
Author information
Authors and Affiliations
Contributions
Conceptualization, L. S. and P. Z.; methodology, L. S.; software, Y. Z.; validation, Y. Q.; formal analysis, S. Y.; investigation, H. Y.; resources, Z.Y.; data curation, Y. Y.; writing—original draft, P. Z. and Y. Y.; writing—review and editing, W. Z. ; visualization, Y. Z; supervision, R. Y.; project administration, W. Z.; All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
No conflict of interest exits in the submission of this manuscript, and manuscript is approved by all authors for publication. I would like to declare on behalf of my co-authors that the work described was original research that has not been published previously, and not under consideration for publication elsewhere, in whole or in part.
Additional information
Edited by Juergen Richt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Song, L., Zhong, P., Yu, R. et al. Effect of HDAC9-induced deacetylation of glycolysis-related GAPDH lysine 219 on rotavirus replication in rotavirus-infected Caco-2 cells. Virus Genes (2024). https://doi.org/10.1007/s11262-024-02104-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11262-024-02104-4