Abstract
Ferroptosis, an iron-dependent form of regulated cell death, has been associated with many virus infections. However, the role of ferroptosis in dengue virus (DENV) infection remains to be clarified. In our study, a dengue fever microarray dataset (GSE51808) of whole blood samples was downloaded from the Gene Expression Omnibus (GEO), and a list of ferroptosis related genes (FRGs) was extracted from the FerrDb. We identified 37 ferroptosis-related differentially expressed genes (FR-DEGs) in DENV-infected patient blood samples compared to healthy individuals. Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses as well as protein–protein interaction (PPI) network of FR-DEGs revealed that these 37 FR-DEGs were mainly related to the C-type lectin receptor and p53 signaling pathway. Nine out of the 37 FR-DEGs (HSPA5, CAV1, HRAS, PTGS2, JUN, IL6, ATF3, XBP1, and CDKN2A) were hub genes, of which 5 were validated by qRT-PCR in DENV-infected HepG2 cells. Finally, using miRNA-mRNA regulatory network, we identified has-miR-124-3p and has-miR-16-5p as the most critical miRNAs in regulating the expression of these hub genes. In conclusion, our findings demonstrated that 5 FR-DEGs, JUN, IL6, ATF3, XBP1, and CDKN2A, and two miRNAs, has-miR-124-3p and has-miR-16-5p may implicate an essential role of ferroptosis in DENV infection, and further studies are warranted to explore the underlying mechanisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dengue virus (DENV), with a positive-stranded RNA genome, is a globally circulating flavivirus transmitted through mosquito bites [1]. The incidence of DENV infection (dengue fever, DF) is increasing annually, causing a major public health concern in tropical and subtropical regions [2]. Other than DF, DENV infection may develop into a more severe form designated as dengue hemorrhagic fever (DHF).
DENV infection may induce cell death although the mechanisms may vary. It has been reported that damaged mitochondrial fusion and altered mitochondrial morphology as well as impaired antiviral responses were observed in DENV-infected cells, leading to enhanced viral replication and cell death [3]. In addition, DENV proteins have been shown to induce cell apoptosis through synergistic activation of complements and chemokines, resulting in transient vascular leakage [4]. DENV infection was also reported to be associated with mitochondria-mediated apoptosis by regulating ISG12b2 [5]. Accumulating data also suggest that DENV induced autophagy regulates cellular lipid metabolism to benefit its own replication [6]. Ferroptosis, an iron-dependent form of cell death, is characterized by intracellular iron-dependency and accumulation of lipid ROS, indicating ferroptosis is mainly mediated by ROS [7, 8]. As a new type of cell death, ferroptosis has been involved in the pathogenesis of many viruses. For example, ferroptosis has been shown to be associated with the replication of SARS-CoV-2 [9], Hepatitis C Virus (HCV) [10], Epstein-Barr virus (EBV) [11], Hepatitis A virus (HAV) [12], and Influenza A virus (IAV) [13]. However, little is known about the role of ferroptosis in the pathogenesis of DENV infection.
In our study, we first identified the differentially expressed genes (DEGs) from GSE51808 dataset which hosted the gene expression data from whole blood samples of 28 patients infected with DENV and 9 healthy individuals. Next, these DEGs were integrated with FRGs from the FerrDb database to further obtain FR-DEGs. PPI network analysis based on FR-DEGs yielded 9 hub genes, of which 5 hub genes JUN, IL6, ATF3, XBP1, and CDKN2A were upregulated in DENV-infected patients and they were successfully validated by qRT-PCR in DENV-infected HepG2 cells. Finally, we identified has-miR-124-3p and has-miR-16-5p were the most critical miRNAs in regulating the expression of these ferroptosis-related hub genes during DENV infection.
Materials and methods
Data acquisition and processing
We downloaded the GSE51808 dataset from the GEO database (http:// www.ncbi.nlm.nih.gov/geo, made to public on Jun 27, 2014). The dataset is based on the gene expression data in whole blood samples from 28 patients with DENV acute infection (18 DF and 10 DHF) and 9 healthy individuals performed on GPL13158 platform (Affymetrix HT HG-U133 + PM Array). FerrDb database is the first manually curated database that dedicates to ferroptosis regulators and ferroptosis-disease associations [14]. Ultimately, we obtained 259 corresponding ferroptosis-related genes (FRGs) from FerrDb V2 database (http://www.zhounan.org/ferrdb, accessed on 31 May 2022). Ferroptosis regulators include driver, suppressor, marker, or multi-annotated genes, and details and classification of FRGs are shown in Supplementary Table1. In addition, the details and classification of these 28 patients together with 9 healthy controls are shown in Supplementary Table 2.
Screening of FR-DEGs from DENV-infected whole blood samples
All data were quantified by log2 scaling to ensure that they were normally distributed. Before DEG analysis, a principal component analysis (PCA) was performed for dimensionality reduction using the R package amap [15]. DEGs between patients with DENV infection and the healthy individuals were identified using the DESeq2 package in R software with cutoffs at |log2 FC|> 1 and P-value < 0.05 [16]. Annotations appear on volcano maps were based on the ggplot2 packages [17]. 259 FRGs were obtained from the FerrDb database and then they were intersected with DEGs of GSE51808 to identify FR-DEGs by Venn diagram. Finally, expression heatmap of FR-DEGs was used for visualization by ComplexHeatmap R package [17].
Functional enrichment analysis of FR-DEGs
To further investigate the biological function of FR-DEGs, we performed the GO functional annotation and KEGG pathway analysis using the R package ClusterProfiler [18]. The GO analysis has three categories containing biological process (BP), molecular function (MF) and cellular component (CC) categories. P < 0.05 was considered as statistical significance.
PPI network analysis and identification of the hub genes
PPI network of FR-DEGs was constructed using the STRING database (https://string-db.org/) to obtain protein interactions with a combined score ≥ 0.4, and PPI networks subsequently were visualized and analyzed by Cytoscape software v3.6.1 [19]. In addition, we utilized the CytoHubba plugin in Cytoscape to track the top 10 highly connected genes by four algorithms including Maximal Clique Centrality (MCC), Maximum Neighborhood Component (MNC), Degree, and Edge Percolated Component (EPC) [20]. The hub genes were obtained from the four algorithms by using intersection in Venn diagram. In addition, we tracked the expression levels of these hub genes in GSE51808 dataset.
Cell culture and DENV virus infection
Human hepatocellular carcinoma cell line HepG2 was routinely maintained in our laboratory and the cells were cultured in Dubecco modified Eagle medium (DMEM, Hyclone, United States) supplemented with 10% fetal bovine serum (FBS, AN Biotech, Germany), 100 U/mL of penicillin (Hyclone, United States), and 100 μg/mL of streptomycin (Hyclone, United States) at 37 °C in a 5% CO2 humidified incubator. DENV-2 virus (New Guinea C strain) was generously provided by professor Zhongtian Qi (Naval Medical University, China). HepG2 cells were infected with DENV-2 in a multiplicity of infection of 1 (MOI = 1) for 2 h followed by incubation for 48 h.
RNA isolation and qRT-PCR
Total RNA was extracted from cells using Trizol reagent (Invitrogen, United States) and cDNA was synthesized using high-capacity cDNA Reverse Transcription Kit (Toyobo, Japan) based on the manufacturer’s protocol [21]. qRT-PCR reactions were performed using CFX96 Real-Time PCR System (Bio-Rad, United States) with the 2 × PCR Master Mix (Toyobo, Japan). The 2−ΔΔCt method was used to calculate the fold difference of all test genes normalized to GAPDH as an internal reference. The sequences of qRT-PCR primers designed and synthesized by Shenggong Biotech Co. Ltd. (Shanghai, China) are listed in Table 1.
Construction of miRNA-mRNA regulatory network
In the present research, NetworkAnalyst 3.0 platform using the miRTarBase v8.0 database (https://www.networkanalyst.ca/NetworkAnalyst/home.xhtml) was used to build pathways upstream of hub genes to reveal the potential miRNA-mRNA regulatory networks [22]. And Cytoscape software v3.6.1 was used to construct and visualize the regulatory networks [23].
Statistical analysis
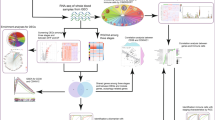
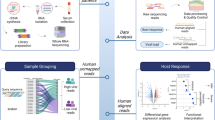
Statistical analysis of qRT-PCR results was performed using GraphPad Prism software. Between-group differences were compared by Student’s t-test. Data were expressed as the mean ± SEM from at least three replicates. * stands for P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001; and ns, not significant. The overall flow chart of this study is shown in Fig. 1.
Results
Identification of FR-DEGs
After batch correction, the median for each sample was essentially located on the horizontal line, indicating a good normalization between samples was obtained (Fig. 2A). Batch variation of the PCA plot shows good clustering between the two groups (Fig. 2B). Volcano plot was drawn to visualize the DEGs (Fig. 2C). These DEGs in GSE51808 were intersected with 259 FRGs screened from FerrDb, and 37 FR-DEGs were identified between the blood samples of DENV-infected patients and healthy individuals (Fig. 2D and Table 2). Heatmap analysis revealed the expression patterns of these 37 FR-DEGs (Fig. 2E). The details of these FR-DEGs including driver, suppressor, and marker genes were shown in Fig. 2F. Interestingly, the same gene can be both driver, suppressor, or marker. TFRC, ATF3 and ALOX5 are both driver and marker genes while SLC40A1 is both suppressor and marker gene. Same analyses were used to identify the FR-DEGs in either DF or DHF patients compared to healthy controls and 34 and 53 FR-DEGs were identified in DF and DHF, respectively (Figure S1 and S2).
The overview process for the identification of FR-DEGs between DENV-infected patients and healthy individuals. A Normalization of GSE51808 dataset. The blue boxes represent samples from healthy individuals (control, n = 9) and the green boxes represent DENV-infected patient samples (DENV, n = 28). Black lines show the median data of each sample. B The visualization of PCA for GSE51808. C Volcano plot of the DEGs, including 884 upregulated and 1289 downregulated genes. Red, upregulation; green, downregulation. Genes without any significant differences are shown in black. |log2FC|> 1 and P < 0.05 were set as cut-off criteria. D Venn diagrams showing the FR-DEGs that intersected between the DEGs in the GSE51808 dataset and FRGs in the FerrDb database. E The heat map of 37 FR-DEGs between healthy individuals and DENV-infected patient samples. Red, up-regulated genes; blue, down-regulated genes. F The 37 FR-DEGs were divided into driver, suppressor, and marker genes related to ferroptosis based on their functions. DENV, Dengue Virus; DEGs, differentially expressed genes; FR-DEGs, ferroptosis-related differentially expressed genes; PCA, principal component analysis
Enrichment analysis of FR-DEGs
To elucidate the potentially enriched pathways of the 37 identified FR-DEGs, GO and KEGG analyses were performed. The top 5 terms in the three GO domains BP, CC, and MF are shown in Table 3. The GO-MF enrichment analysis indicated that these genes were mainly related to transaminase activity, oxidoreductase activity, ubiquitin protein ligase binding, ubiquitin-like protein ligase binding, and antioxidant activity (Fig. 3A). KEGG analysis showed most of these genes were mainly enriched in neurodegeneration-multiple diseases, Alzheimer disease, C-type lectin receptor signaling pathway, apoptosis, and p53 signaling pathway (Fig. 3B). The top 10 terms in KEGG pathway are shown in Table 4.
GO and KEGG enrichment analysis of the 37 FR-DEGs. A The top 5 terms in the three GO domains (BP, CC, MF) are shown in bubble plot. B Top 10 KEGG pathways. GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; BP, Biological Process; CC, Cellular Components; MF, Molecular Functions; KEGG, Kyoto Encyclopedia of Genes and Genomes
Construction of PPI network and identification of the hub genes
The PPI network constructed from 37 FR-DEGs between DENV infected and healthy individuals has a total of 34 nodes and 56 edges (Fig. 4A). Nine common (hub) genes (HSPA5, CAV1, HRAS, PTGS2, JUN, IL6, ATF3, XBP1, and CDKN2A) were identified using any of the 4 different algorithms (MCC, MNC, Degree and EPC) by CytoHubba plugin, and they were displayed in a Venn diagram for easier visualization (Fig. 4B). The genes identified from any of the four algorithms were shown in Table 5. The expression patterns of these 9 hub genes were further validated in GSE51808 dataset. Except for one gene, PTGS2 whose expression was downregulated, all the other 8 hub genes (HSPA5, CAV1, HRAS, JUN, IL6, ATF3, XBP1, and CDKN2A) were upregulated in DENV infected patient samples (Fig. 5). These genes may regulate ferroptosis in DENV infection and further validation in DENV infected cells has been planned in our future study.
PPI network analysis of 37 FR-DEGs by Cytoscape. A PPI network constructed with FR-DEGs, including 34 nodes and 56 edges. Each node represents protein, and each edge shows interactions between these proteins. B Intersection of top 10 hub genesfrom the PPI network analysis ranked by four algorithms (MCC, MNC, Degree, and EPC). PPI, Protein-Potein Interaction; FR-DEGs, ferroptosis-related differentially expressed genes; MCC, Maximal Clique Centrality; MNC, Maximum Neighborhood Component; EPC, Edge Percolated Component
Validation of the expression patterns of the hub genes identified from the FR-DEGs between patients with DENV (n = 28) and healthy individuals (n = 9) in the GES51080 dataset. A HSPA5, B CAV1, C HRAS, D PTGS2, E JUN, F IL6, G ATF3, H XBP1, and I CDKN2A. Each point shows a sample. Statistical analysis was performed with the Wilcoxon rank sum test. *P < 0.05; ** P < 0.01; *** P < 0.001
For 34 FR-DEGs identified in DF patients compared to healthy individuals, eight hub genes (HRAS, NRAS, PTGS2, JUN, ATF3, CDKN2A, HSPA5, and CAV1) were obtained from PPI analysis (Figure S1 and S3). Except for one gene, HRAS whose expression was downregulated, all the other 7 hub genes were upregulated (Figure S3B) in DF patients compared to healthy individuals. Similarly, nine hub genes (ATF3, MAPK1, JUN, DUSP1, HSPA5, PTGS2, HRAS, TLR4, and IL6) were identified by PPI analysis from 53 FR-DEGs in DHF patients (Figures S2 and S3) and six (ATF3, JUN, HSPA5, HRAS, and IL6) were upregulated, while three (MAPK1, DUSP1, PTGS2, and TLR4) were downregulated in DHF patients compared to healthy individuals (Figure S3D). Although 5 common hub genes (HRAS, PTGS2, JUN, ATF3, and HSPA5) were identified and three genes (JUN, ATF3 and HSPA5) were upregulated in both DF and DHF patients, expression patterns of the remaining two common genes, HRAS and PTGS2 are quite different. HRAS gene was downregulated in DF while it was upregulated in DHF, while PTGS2 was upregulated in DF while it was downregulated in DHF (Figure S3B, 3D). These findings may implicate the potential relationship between level of ferroptosis genes expression and severity of dengue infection (DF vs DHF), and further studies are warranted to explore the potential mechanisms.
Validation of the hub genes by qRT-PCR in DENV-infected HepG2 cells
We performed qRT-PCR analyses to validate changes in the expression levels of the hub genes in DENV-infected HepG2 cells. As shown in Fig. 6, the expression levels of JUN, IL6, ATF3, XBP1, and CDKN2A were significantly upregulated in HepG2 cells infected with DENV-2, and these results were consistent with bioinformatics analyses although the expression levels of the remaining 4 genes, HSPA5, CAV1, HRAS, and PTGS2, were not confirmed.
Validation of the expression levels of hub genes in DENV-infected HepG2 cells by qRT-PCR. No significant changes were found in the expression level of (A) HSPA5, (B) CAV1, (C) HRAS, and (D) PTGS2. The expression levels of (E) JUN, (F) IL6, (G) ATF3, (H) XBP1, and (I) CDKN2A were upregulated in DENV-infected cells (n ≥ 3) compared to non-infected controls (n ≥ 3). Data are presented as mean ± SEM; P-values are shown as: ns, not significant; **P < 0.01; ***P < 0.001; ****P < 0.0001. DENV, Dengue Virus; qRT-PCR, quantitative real-time polymerase chain reaction; SEM, standard error of mean
MiRNA-MRNA regulating network analysis
To further investigate the post-transcriptional regulatory network between miRNAs and hub genes, a miRNA-mRNA regulatory network was constructed based on 9 hub genes and this network contained 217 nodes and 251 edges (Fig. 7). We selected top eight miRNAs based on the degree (Table 6). Among these miRNAs, has-miR-124-3p was linked with IL6, PTGS2, XBP1, CDKN2A, and CAV1, and has-miR-16-5p interacted with PTGS2, JUN, CDKN2A, and HSPA5. Thus, we consider has-miR-124-3p and has-miR-16-5p to be the most critical miRNAs that may play a major role in regulating the post-transcriptional expression of the hub genes involved in ferroptosis during DENV infection.
Discussion
DF, a global arthropod-borne viral disease, is caused by DENV infection mainly transmitted through mosquito bites [24]. The pathogenesis of DENV remains partially understood. Many viruses exploit the host responses to benefit their replication. As a new type of cell death, ferroptosis has been involved in the pathogenesis of many viruses, such as SARS-CoV-2, HCV, EBV, HAV, and IAV [9,10,11,12,13]. However, the role of ferroptosis in the pathogenesis of DENV infection has not been explored. In this current study, we employed bioinformatics analysis coupled with the in-vitro cell culture model to elucidate the potential involvement of ferroptosis in DENV infection based on the microarray gene expression dataset from public databases.
GSE51808 dataset, which contains gene expression data from 28 patients (18 DF and 10 DHF) with acute DENV infection and 9 healthy individuals, was downloaded for our analysis. DEGs were identified from GSE51808, and these DEGs were intersected with 259 FRGs screened from FerrDb, and 37 FR-DEGs, including 7 down-regulated genes and 30 up-regulated genes, were obtained (Fig. 2D and Table 2). GO-MF analysis indicated that these genes were largely involved in the antioxidant and oxidoreductase activities, indicating that oxidation is an essential process in ferroptosis which was consistent with previous reports [25, 26]. Results from KEGG analyses indicated that these 37 FR-DEGs were mainly associated with C-type lectin receptor, apoptosis, and p53 signaling pathway.
CLEC5A, a member of C-type lectin receptor family, has been recognized as a critical signaling molecule that mediates the enhanced production of TNF-α in the process of DENV infection [27]. It has been confirmed that the blockade of CLEC5A-mediated signaling could attenuate the production of proinflammatory cytokines by macrophages infected with DENV [28]. Although CLEC5A was not identified as one of the hub genes in our current study, C-type lectin receptor may be involved in the pathogenesis of DENV. DENV infection may induce cell death through different mechanisms. For example, DENV proteins induce apoptosis in infected cells by activating NF-κB signaling [29]. Recent studies have shown that p53 signaling pathway can promote ferroptosis following virus infection [30]. Study from Xie, et al. demonstrated that p53 was a positive regulator of ferroptosis through inhibition of SLC7A11 expression [31]. In addition, p53 signaling was shown to activate type I interferon pathway and promote cell apoptosis to inhibit DENV infection [32]. Considering the fact that our analysis points to the involvement of p53 signaling pathway in the pathogenesis of DENV infection, especially through ferroptosis, it will be interesting to investigate how ferroptosis affect DENV infection in future studies.
To further reveal the mechanism underlying the biological processes, a PPI analysis based on the 37 FR-DEGs was performed. Nine hub genes (HSPA5, CAV1, HRAS, PTGS2, JUN, IL6, ATF3, XBP1, and CDKN2A) were identified by all the 4 different algorithms. Five out of these 9 genes were confirmed by qRT-PCR and they were all up-regulated in DENV-infected HepG2 cells, and these results were consistent with GES51808 dataset. As shown in Fig. 2F, IL-6, ATF3, and XBP1 were classified as marker genes of ferroptosis that may indicate the presence of ferroptosis during DENV infection. JUN was considered as an inhibitor gene of ferroptosis, while ATF3 and CDKN2A were classified as driver genes that may promote ferroptosis [14].
DENV infection induces the increased production of IL6 which has been implicated in the pathogenesis of severe DENV disease [33]. It has been reported that IL6, a pro-inflammatory cytokine, promotes ferroptosis by inducing cellular ROS-dependent lipid peroxidation and disrupting iron homeostasis [34]. Interestingly, IL6 reversed ferroptosis and growth suppression that was induced by xCT knockdown because xCT is a key amino acid antiporter facilitating ferroptosis resistance [35]. Collectively, these results clearly demonstrated that IL-6 plays an important regulatory role in the pathogenesis of DENV infection through modulating ferroptosis. ATF3, another hub genes we identified from this study, is also involved in ferroptosis. It has been reported that as a common stress sensor gene, ATF3 induces ferroptosis by repressing SLC7A11 expression [36]. Elevated ATF3 was also found to induce ferroptosis in gastric carcinoma by inhibiting Nrf2/Keap1/xCT signaling [37]. Another hub gene XBP1, a critical regulator involved in cellular unfolded protein response (UPR) and plasma cell differentiation, has been shown to promote ferroptosis in human T-lymphotropic virus type 1 (HTLV-1) infection [38, 39]. CDKN2A, also named as ARF, stimulated ROS-induced ferroptosis in a p53-independent manner [40]. In our current study we also identified the expression level of CDKN2A was increased in HepG2 cells infected with DENV. All these findings indicated that IL6, ATF3, XBP1, and CDKN2A were closely linked to ferroptosis and virus infection, and these data form the basis for our hypothesis that these genes may be involved in the pathogenesis of DENV infection by regulation of ferroptosis in host cells.
To strengthen our hypothesis that ferroptosis is involved in the pathogenesis and severity of DENV infection, FR-DEGs of DF and DHF patients were further analyzed and compared. Eight hub genes (HRAS, NRAS, PTGS2, JUN, ATF3, CDKN2A, HSPA5, and CAV1) were obtained in DF patients compared to healthy individuals by PPI analysis. HRAS, NRAS, ATF3, and CDKN2A were considered as driver genes that may promote ferroptosis. PTGS2 and ATF3 were considered as marker genes of ferroptosis that may indicate the presence of ferroptosis during DENV infection, and HSPA5, JUN, and CAV1 were considered as inhibitor genes of ferroptosis [14]. Similarly, nine hub genes (ATF3, MAPK1, JUN, DUSP1, HSPA5, PTGS2, HRAS, TLR4, and IL6) were identified in DHF patients compared to healthy individuals by PPI analysis. Although 5 common hub genes (HRAS, PTGS2, JUN, ATF3, and HSPA5) were identified and three genes (JUN, ATF3 and HSPA5) were upregulated in both DF and DHF patients, expression patterns of the remaining two common genes, HRAS and PTGS2 are quite different. HRAS gene was downregulated in DF while it was upregulated in DHF, while PTGS2 was upregulated in DF while it was downregulated in DHF (Figure S3B, 3D). These findings may implicate the potential relationship between level of some ferroptosis-related gene expression, such as HRAS and PTGS2, and severity of dengue infection (DF vs DHF). Taken together, findings not only show a clear relationship between DENV infection and ferroptosis but also implicate the potential involvement of some ferroptosis-related hub gene expression patterns (up or down) in the severity of DENV infection (DF vs DHF).
Many genes are regulated at post-transcriptional level and microRNAs have been shown to play a major role in this regulation process. Therefore, identification of the potential microRNAs that may regulate these ferroptosis-related hub genes may provide novel insights into understanding the gene regulatory network and the potential targets for antiviral drug development. Two microRNAs, has-miR-124-3p and has-miR-16-5p have been identified to target more than one hub genes and they may represent the major non-coding RNAs involved in the regulation of ferroptosis-related hub genes. However, the detailed biological function and regulation role of these microRNAs need further validation in our future study using DENV cell culture model.
Conclusion
Using various bioinformatics analysis tools, we identified and validated 9 hub genes, which were involved in ferroptosis and DENV infection. Our current results indicated the expression patterns and levels of some FRGs may be closely associated with the severity of dengue disease (DF or DHF). In addition, we also found two microRNAs that regulate these hub gene expressions. Although further studies on the biological function of these hub genes and regulatory role of these microRNAs are clearly needed, our data undoubtedly demonstrated an essential role of ferroptosis in the pathogenesis and severity of DENV infection.
Data availability
The datasets supporting the conclusions of this article are available in the Gene Expression Omnibus (GEO) (www.ncbi.nlm.nih.gov/geo) database and the FerrDB (http://www.zhounan.org/ferrdb/current/) database.
References
Kim HO, Na W, Yeom M, Lim JW, Bae EH, Park G et al (2020) Dengue virus-polymersome hybrid nanovesicles for advanced drug screening using real-time single nanoparticle-virus tracking. ACS Appl Mater Interfaces. https://doi.org/10.1021/acsami.9b20492
Eyre NS, Johnson SM, Eltahla AA, Aloi M, Aloia AL, McDevitt CA et al (2017) Genome-wide mutagenesis of dengue virus reveals plasticity of the NS1 protein and enables generation of infectious tagged reporter viruses. J Virol. https://doi.org/10.1128/jvi.01455-17
Yu CY, Liang JJ, Li JK, Lee YL, Chang BL, Su CI et al (2015) Dengue virus impairs mitochondrial fusion by cleaving mitofusins. PLoS Pathogens. https://doi.org/10.1371/journal.ppat.1005350
Avirutnan P, Malasit P, Seliger B, Bhakdi S, Husmann M (1998) Dengue virus infection of human endothelial cells leads to chemokine production, complement activation, and apoptosis. J Immunol 161(11):6338–46
Lu MY, Liao F (2011) Interferon-stimulated gene ISG12b2 is localized to the inner mitochondrial membrane and mediates virus-induced cell death. Cell Death Differ 18(6):925–936. https://doi.org/10.1038/cdd.2010.160
Heaton NS, Randall G (2010) Dengue virus-induced autophagy regulates lipid metabolism. Cell Host Microbe 8(5):422–432. https://doi.org/10.1016/j.chom.2010.10.006
Chen X, Kang R, Kroemer G, Tang D (2021) Ferroptosis in infection, inflammation, and immunity. J Exp Med. https://doi.org/10.1084/jem.20210518
Dixon SJ, Lemberg KM, Lamprecht MR, Skouta R, Zaitsev EM, Gleason CE et al (2012) Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149(5):1060–1072. https://doi.org/10.1016/j.cell.2012.03.042
Han Y, Zhu J, Yang L, Nilsson-Payant BE, Hurtado R, Lacko LA et al (2022) SARS-CoV-2 infection induces ferroptosis of sinoatrial node pacemaker cells. Circ Res 130(7):963–977. https://doi.org/10.1161/circresaha.121.320518
Yamane D, Hayashi Y, Matsumoto M, Nakanishi H, Imagawa H, Kohara M et al (2022) FADS2-dependent fatty acid desaturation dictates cellular sensitivity to ferroptosis and permissiveness for hepatitis C virus replication. Cell Chem Biol 29(5):799-810.e4. https://doi.org/10.1016/j.chembiol.2021.07.022
Burton EM, Voyer J, Gewurz BE (2022) Epstein-Barr virus latency programs dynamically sensitize B cells to ferroptosis. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.2118300119
Komissarov AA, Karaseva MA, Roschina MP, Shubin AV, Lunina NA, Kostrov SV et al (2021) Individual expression of hepatitis a virus 3C protease induces ferroptosis in human cells in vitro. Inter J Mol Sci. https://doi.org/10.3390/ijms22157906
Cheng J, Tao J, Li B, Shi Y, Liu H (2022) Swine influenza virus triggers ferroptosis in A549 cells to enhance virus replication. Virology journal 19(1):104. https://doi.org/10.1186/s12985-022-01825-y
Zhou N, Bao J (2020) FerrDb: a manually curated resource for regulators and markers of ferroptosis and ferroptosis-disease associations. Database. https://doi.org/10.1093/database/baaa021
Nakagawa S, Niimura Y, Gojobori T (2017) Comparative genomic analysis of translation initiation mechanisms for genes lacking the Shine-Dalgarno sequence in prokaryotes. Nucleic Acids Res 45(7):3922–3931. https://doi.org/10.1093/nar/gkx124
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550. https://doi.org/10.1186/s13059-014-0550-8
Honjo K, Won WJ, King RG, Ianov L, Crossman DK, Easlick JL et al (2020) Fc receptor-like 6 (FCRL6) discloses progenitor B cell heterogeneity that correlates with Pre-BCR dependent and independent pathways of natural antibody selection. Front Immunol 11:82. https://doi.org/10.3389/fimmu.2020.00082
Deng M, Chen B, Liu Z, Cai Y, Wan Y, Zhang G et al (2020) YTHDF2 regulates maternal transcriptome degradation and embryo development in goat. Front Cell Develop Biol. https://doi.org/10.3389/fcell.2020.580367
Wang D, Chen J, Ding Y, Kong H, You H, Zhao Y et al (2020) miR-188-5p promotes tumor growth by targeting CD2AP through PI3K/AKT/mTOR signaling in children with acute promyelocytic leukemia. Onco Targets Ther 13:6681–6697. https://doi.org/10.2147/ott.S244813
Nangraj AS, Selvaraj G, Kaliamurthi S, Kaushik AC, Cho WC, Wei DQ (2020) Integrated PPI- and WGCNA-retrieval of hub gene signatures shared between barrett’s esophagus and esophageal adenocarcinoma. Front Pharmacol 11:881. https://doi.org/10.3389/fphar.2020.00881
Wang ZE, Wu D, Zheng LW, Shi Y, Wang C, Chen ZH et al (2018) Effects of glutamine on intestinal mucus barrier after burn injury. Am J Translational Res 10(11):3833–3846
Zhu GD, Cao XJ, Li YP, Li JX, Leng ZJ, Xie LM et al (2021) Identification of differentially expressed genes and signaling pathways in human conjunctiva and reproductive tract infected with Chlamydia trachomatis. Hum Genomics 15(1):22. https://doi.org/10.1186/s40246-021-00313-8
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504. https://doi.org/10.1101/gr.1239303
Meuren LM, Prestes EB, Papa MP, de Carvalho LRP, Mustafá YM, da Costa LS et al (2022) Infection of endothelial cells by dengue virus induces ROS production by different sources affecting virus replication Cellular Activation, Death and Vascular Permeability. Front Immunol. https://doi.org/10.3389/fimmu.2022.810376
Ferrari M, Zevini A, Palermo E, Muscolini M, Alexandridi M, Etna MP et al (2020) Dengue virus targets Nrf2 for NS2B3-mediated degradation leading to enhanced oxidative stress and viral replication. J Virol. https://doi.org/10.1128/jvi.01551-20
Santana-Román ME, Maycotte P, Uribe-Carvajal S, Uribe-Alvarez C, Alvarado-Medina N, Khan M et al (2021) Monitoring mitochondrial function in Aedes albopictus C6/36 Cell line during dengue virus infection. Insects. https://doi.org/10.3390/insects12100934
Cheng YL, Lin YS, Chen CL, Tsai TT, Tsai CC, Wu YW et al (2016) Activation of Nrf2 by the dengue virus causes an increase in CLEC5A, which enhances TNF-α production by mononuclear phagocytes. Sci Rep 6:32000. https://doi.org/10.1038/srep32000
Chen ST, Lin YL, Huang MT, Wu MF, Cheng SC, Lei HY et al (2008) CLEC5A is critical for dengue-virus-induced lethal disease. Nature 453(7195):672–676. https://doi.org/10.1038/nature07013
Lin JC, Lin SC, Chen WY, Yen YT, Lai CW, Tao MH et al (2014) Dengue viral protease interaction with NF-κB inhibitor α/β results in endothelial cell apoptosis and hemorrhage development. J Immunol. https://doi.org/10.4049/jimmunol.1302675
Liu J, Zhang C, Wang J, Hu W, Feng Z (2020) The regulation of ferroptosis by tumor suppressor p53 and its pathway. Int J Mol Sci. https://doi.org/10.3390/ijms21218387
Xie Y, Hou W, Song X, Yu Y, Huang J, Sun X et al (2016) Ferroptosis: process and function. Cell Death Differ 23(3):369–379. https://doi.org/10.1038/cdd.2015.158
Hu YL, Li XS, Xiong S, Ma Q, Liu D, Shi ZQ et al (2017) The inhibiting effect of the transcription factor p53 on dengue virus infection by activating the type I interferon. Oncotarget 8(15):25151–25157. https://doi.org/10.18632/oncotarget.15352
Chaudhary N, Srivastava S, Dave U, Ojha A, Guchhait P, Chandele A et al (2022) High-mobility group box 1 protein promotes dengue virus replication by interacting with untranslated regions of viral genome. Virus Res. https://doi.org/10.1016/j.virusres.2021.198668
Han F, Li S, Yang Y, Bai Z (2021) Interleukin-6 promotes ferroptosis in bronchial epithelial cells by inducing reactive oxygen species-dependent lipid peroxidation and disrupting iron homeostasis. Bioengineered 12(1):5279–5288. https://doi.org/10.1080/21655979.2021.1964158
Li M, Jin S, Zhang Z, Ma H, Yang X (2022) Interleukin-6 facilitates tumor progression by inducing ferroptosis resistance in head and neck squamous cell carcinoma. Cancer Lett 527:28–40. https://doi.org/10.1016/j.canlet.2021.12.011
Wang L, Liu Y, Du T, Yang H, Lei L, Guo M et al (2020) ATF3 promotes erastin-induced ferroptosis by suppressing system Xc(). Cell Death Differ 27(2):662–675. https://doi.org/10.1038/s41418-019-0380-z
Fu D, Wang C, Yu L, Yu R (2021) Induction of ferroptosis by ATF3 elevation alleviates cisplatin resistance in gastric cancer by restraining Nrf2/Keap1/xCT signaling. Cell Mol Biol Lett 26(1):26. https://doi.org/10.1186/s11658-021-00271-y
Dixon SJ, Patel DN, Welsch M, Skouta R, Lee ED, Hayano M et al (2014) Pharmacological inhibition of cystine-glutamate exchange induces endoplasmic reticulum stress and ferroptosis. ELife. https://doi.org/10.7554/eLife.02523
Lew QJ, Chu KL, Lee J, Koh PL, Rajasegaran V, Teo JY et al (2011) PCAF interacts with XBP-1S and mediates XBP-1S-dependent transcription. Nucleic Acids Res 39(2):429–439. https://doi.org/10.1093/nar/gkq785
Chen D, Tavana O, Chu B, Erber L, Chen Y, Baer R et al (2017) NRF2 Is a major target of ARF in p53-independent tumor suppression. Mol Cell 68(1):224–32.e4. https://doi.org/10.1016/j.molcel.2017.09.009
Acknowledgements
Thanks to the Xiantao tool (https://www.xiantao.love/) and all the people in our laboratory for their help.
Funding
This study is supported by Qin Chuangyuan recruited high-level innovation and entrepreneurship talents project of Science and Technology Department of Shaanxi Province (QCYRCXM-2022–56); and Medical Research project of Xi'an Science and Technology Bureau (22YXYJ0120).
Author information
Authors and Affiliations
Contributions
LMC and QYW conceived the study and edited the manuscript. JLL and XPY wrote the manuscript. BL, HX and BLH prepared the figures and tables. XWW and BRH helped to carry out the cell-culture and data analysis. LMC revised the manuscript. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflicts of interest
The authors declared that there was no conflict of interests.
Additional information
Edited by Juergen Richt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, J., Yan, X., Li, B. et al. Identification and validation of ferroptosis-related genes in patients infected with dengue virus: implication in the pathogenesis of DENV. Virus Genes 59, 377–390 (2023). https://doi.org/10.1007/s11262-023-01985-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-023-01985-1