Abstract
Canine parvovirus (CPV-2) modified-live virus vaccine strain can replicate in lymphoid tissues and intestinal mucosa after administration, being shed through canine faeces. Detection of vaccine strains has been reported in the bloodstream and faeces, potentially interfering with molecular diagnostic tests. The persistence of these strains in canine tissues has not yet been described. With this aim, canine tissues were tested during a molecular survey to screen for the presence of canine enteric viruses. Tissue samples from 165 dead dogs were tested by a conventional PCR assay. Positive samples and five commercial vaccines were subjected to sequence analysis. Vaccinal strains were detected and virus load was measured by using a set of real-time PCR assays using minor-groove binder (MGB) probes. Seventy-five dogs (45.4%) tested positive for CPV-2. Strains from 70 dogs were characterised as field variants. The presence of CPV sequences of vaccine origin was observed in the spleen, intestine, and mesenteric lymph nodes of five young dogs. Vaccinal strains were detected from 12 to 24 days after the last vaccine administration. Viral loads comprised between 6.3 × 102 and 9.95 × 104 DNA copies/10 µl of template. This study confirms that CPV vaccinal strains can be detected in canine tissues after vaccination, so post-mortem diagnosis of CPV infection needs further molecular analyses to assess the viral type (vaccine or field strains). The present study updates the current information on the persistence of CPV vaccine strains in canine tissues and their possible interference with molecular assays.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Canine parvovirus type 2 (CPV-2) is the causative agent of canine parvovirosis, an acute and often fatal viral disease characterised by severe gastroenteritis and lymphopenia, mainly affecting young dogs. CPV is a small, non-enveloped single‐stranded DNA virus, included in the species Carnivore protoparvovirus 1, a member of the Protoparvovirus genus (Family Parvoviridae, Subfamily Parvovirinae) (Cotmore et al. 2019). The original CPV-2 type was first identified as a canine pathogen in the late 1970s and has since been replaced by three antigenic and genetic variants, namely CPV-2a, CPV-2b, and CPV-2c (Parrish et al. 1991; Buonavoglia et al. 2001). While the three CPV-2 variants are still distributed worldwide, the original CPV-2 type is no longer circulating in the field and is present only in some vaccines (Decaro and Buonavoglia 2017).
To prevent canine parvovirus (CPV) infection in dogs, CPV vaccines are highly recommended to stimulate a protective antibody and cell-mediated immune response and, thus, considered as core vaccines (WSAVA 2016; AAHA). Currently, modified-live virus (MLV) vaccine formulations, including the original CPV-2 type or the CPV-2b variant, are licensed and available in the market. After primary vaccination protocol with MLV vaccines, revaccination is required at intervals to warrant the long duration of immunity (Day et al. 2016; Ford et al. 2017).
After administration of MLV vaccines, CPV replicates in lymphoid tissues and intestinal mucosa, with associated viraemia and fecal shedding for a short (12–19 mean days) period (Carmichael et al. 1981; Decaro et al. 2014). Detection of vaccine strains has been reported in the bloodstream and faeces, potentially interfering with diagnostic tests (Decaro et al. 2017; Freisl et al. 2017). The persistence and distribution of CPV vaccinal strains in canine tissues and, thus, the potential occurrence of misleading laboratory diagnoses in tissue samples have not yet been described. During a molecular survey of canine enteric viruses for diagnostic purposes, the presence of CPV vaccine strains in tissue samples was evidenced and herein described.
Materials and methods
CPV vaccines and clinical samples
A total number of six vaccine formulations licensed in Italy, including four CPV type 2 (154, CPV780916, NL-35-D, CAG2) and two CPV type 2b (CPV39, CPV-2b Bio 12/B) vaccine strains, were tested (more details in Supplementary file 1).
Between 2019 and 2021, a total of 165 dead dogs were submitted to Istituto Zooprofilattico Sperimentale della Sicilia “A. Mirri” (Palermo, Italy) for diagnostic purposes and subjected to necropsy. Tissue samples (brain, lungs, heart, kidneys, liver, spleen, intestine, mesenteric lymph nodes) were collected and stored at -80 °C until use.
Nucleic acid extraction and PCR
Lyophilised vaccines were resuspended in 1ml of PBS and total DNA was extracted from 200 µl of the suspensions or ready-to-use liquid formulations using the DNeasy Blood & Tissue Kit (Qiagen S.p.A.), according to the manufacturer’s instructions.
All organ homogenates were obtained as previously described (Purpari et al. 2018). DNA and RNA were extracted using the DNeasy Blood & Tissue Kit and the QIAmp Viral RNA Mini Kit (Qiagen S.p.A.), respectively, according to the manufacturer’s instructions.
A diagnostic PCR assay using the primers pair VP2-850-Forward/VP2-1550-Reverse targeting the CPV VP2 gene (Touihri et al. 2009) was performed to evaluate the presence of CPV DNA, as previously described (Mira et al. 2018). Extracted DNA and RNA were also tested for the presence of other canine viral pathogens such as canine adenoviruses (CAdV-1 and CAdV-2) (Hu et al. 2001), canine coronavirus (CCoV) (Pratelli et al. 2001), canine distemper virus (CDV) (Barrett et al. 1993), canine rotavirus (CRoV) (Freeman et al. 2008), and norovirus (NoV) (Bodnar et al. 2017), as described in Schirò et al. (2022).
Sequence analyses
Sequence analysis of the vaccinal CPV genome sequence encompassing the VP2 coding region was carried out using primers pairs 2161 F/4823R designed by Pérez et al. (2014), as previously described (Mira et al. 2019).
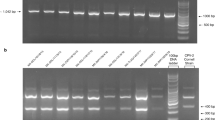
After electrophoresis on agarose gel supplemented with ethidium bromide, positive amplicons obtained from CPV vaccine strains and from CPV-2 screening were purified with llustra™ GFX™ PCR DNA and Gel Band Purification Kit (GE Healthcare Life Sciences, UK). Purified PCR products were submitted to BMR Genomics srl (Padova, Italy) for direct Sanger sequencing with the same primers for CPV-2 screening or, for CPV vaccines analysis, with a set of primers as previously described (Mira et al. 2018). Sequences were assembled and analysed using BioEdit ver 7.2.5 software (Hall 1999). Sequences were analysed with nBLAST (Zhan et al. 2000) to search related sequences in public domain databases (Supplementary file 2). Viral typing was based on the analysis of VP2 amino-acid (aa) residues discriminating the viral type (CPV-2) and the CPV-2 variants (Martella et al. 2005). Sequence data were submitted to the DDBJ/EMBL/GenBank databases under accession numbers reported in Table 1.
To elucidate the genetic relationships of CPV vaccinal strains with other strains, a phylogenetic tree based on partial VP2 gene sequences was constructed with the MEGA X software (Kumar et al. 2018), using the maximum-likelihood method according to the Tamura 3‐parameter (T92) model with discrete Gamma distribution (five rate categories) (bootstrap analyses with 1,000 replicates). The model selection was performed using the best‐fit model of nucleotide substitution test with MEGA X software.
MGB probe assays
CPV vaccinal strains from tissue samples were confirmed and virus load was measured by using a set of real-time PCR assays with minor-groove binder (MGB) probes (Decaro et al. 2006a, b, c). Reactions were carried out following the protocols and the oligonucleotides reported in Decaro et al. (2007a).
Virus isolation
To evaluate the potential presence of vaccinal infectious virus, virus isolation in cell cultures was performed for all vaccinal strains from tissue samples, as previously described (Ogbu et al. 2021). Briefly, samples were inoculated on subconfluent A-72 cell monolayers and left in contact for 30 min at 37 °C and 5% CO2. Culture medium, Eagle’s Minimum Essential Medium (EMEM) (Sigma–Aldrich®, Milan, Italy) supplemented with an antibiotic and antimycotic solution (100 U/mL penicillin G sodium salt, 0.1 mg/mL streptomycin sulfate, 0.25 µg/mL amphotericin B; EuroClone®, Milan, Italy), 1% sodium pyruvate (EuroClone®, Milan, Italy) and 10% fetal bovine serum (FBS; EuroClone®, Milan, Italy), was then added and the incubation was performed at 37 °C and 5% CO2 for 6–7 days. Inoculated cells were monitored daily and, according to standard laboratory procedures, four additional blind passages were carried out before considering virus isolation as unsuccessful. Viral growth was evaluated by the detection of cytopathic effect (CPE). Moreover, inoculated cell monolayers were subjected to three cycles of freeze-thawing, centrifuged at 1,500 x g for 15 min at 4 °C, and the collected supernatants were tested for CPV-2 DNA by PCR assays (Touihri et al. 2009) to confirm the presence of infectious virus.
Results
Vaccines strains analysis
All vaccine samples tested positive for canine parvovirus. The complete VP2 gene sequence (1,755 nucleotides) was obtained for each strain. Based on the analysis of VP2 aa residues discriminating the CPV variants, 4 vaccine strains were characterized as CPV-2 (87 M, 101I, 297 S, 300 A, 305D, 426 N) and 2 strains as CPV-2b (87 L, 101T, 297 S/A, 300G/A, 305Y, 426D). Sequence analysis revealed amino acid changes specific for each vaccinal strain, reported in Table 1.
Detection and characterization of CPV
Among the samples collected from 165 dead dogs, samples from 75 dogs (45.4%) tested positive for CPV-2. Partial VP2 sequences (700 nucleotides) were obtained for each positive dog. Based on the analysis of VP2 amino acid residues discriminating the CPV variants, strains were characterized as CPV-2 (n = 5), CPV-2a (n = 15), CPV-2b (n = 21), and CPV-2c (n = 34).
Based on the sequence analysis, strains from 70 dogs were characterized as field variants, whereas those from five young dogs (6.7%), aging from 2 to 12 months, as vaccinal strains. According to the available anamnestic information, vaccinal strains were detected from 12 to 24 days after the last vaccine administration. CPV DNA of vaccine origin was obtained from spleen, intestine, and mesenteric lymph nodes samples. Other samples of the same dogs tested negative for CPV and all samples from the same dogs tested negative for the other viruses. For these five dogs, death was attributed to causes different from CPV-2 infection. More details on these vaccinal strains and their origin are provided in Table 2. Samples of the other dogs tested also positive, with lower frequencies, to other viruses (more details in Supplementary file 3).
Sequence analyses
CPV sequences from five dogs (2019PA15831, 2020PA59036, 2020PA72035, 2021PA30130, and 2021PA59813) showed high nucleotide identity (99.85–100%) with CPV-2 vaccine strain sequences: three dogs (2019PA15831, 2020PA59036, and 2021PA30130) with vaccine strain 154; one dog (2021PA59813) with strain NL-35-D; one dog (2020PA72035) with strains CAG2 or CPV780916. These sequences revealed specific amino acid residues common to those of vaccinal strains (Table 1).
Vaccine strains showed high nucleotide identity (100%) with other CPV strains (2020PA72035 with accession number KY242639; 2021PA59813 with MK357724 and MK357733; 2019PA15831 with MN104204 and FJ222824; 2020PA57036 and 2021PA30130 with MH491851 and MN810900) obtained from rectal/faecal swabs or faeces collected in Australia in 2015 (Meggiolaro et al. 2017), Vietnam in 2017 (Hoang et al. 2019), Italy in 2005, 2008, and 2014 (Decaro et al. 2005; Franzo et al. 2019; Tucciarone et al. 2018), and in China in 2018/2019 (Jiang et al. 2021).
Phylogenetic analysis inferred from partial VP2 gene sequences indicated that analysed strains from tissue samples were more related to the original CPV-2 type, clustering along with vaccine strains and related CPV strains from dogs in Vietnam, Italy, China, and Australia in a separate clade from the CPV-2 variants (Fig. 1).
Maximum-likelihood tree based on 78 partial VP2 gene sequences of canine parvovirus type 2 strains (T92 + G; bootstrap 1,000 replicates). Black dots markings indicate CPV strains of vaccine origin and black square markings indicate CPV vaccine strains. Each sequence is indicated with virus variant, country and year of collection, and accession number. 1VP2-297Ala amino acid, 2VP2-297Ser amino acid
MGB probe assays
The presence of CPV vaccine strains in tested samples was confirmed by MGB probe assays. Viral loads comprised between 6.3 × 102 and 9.95 × 104 DNA copies/10 µl of a template. More details on viral loads are provided in Table 2 and depicted in Fig. 2. CPV field strains were not detected in samples positive for CPV vaccines.
Virus isolation
The five vaccinal strains from tissue samples were inoculated into A-72 cell monolayers. After multiple passages, no CPE was observed and all samples tested negative by PCR assay. Thus, the isolation was considered as unsuccessful.
Discussion and conclusions
A core vaccine is thet one recommended in order to provide life-long protection against infectious diseases of global significance for dog, such as those determined by CPV-2, CDV, and CAV types 1 and 2 infections, and that the worldwide dog population must receive (Day et al. 2016). For these reasons and to prevent threatening diseases, vaccines for dogs including these viral agents are widely used in the current veterinary practice.
Similarly to natural infection (Meunier et al. 1985), CPV-2 MLV vaccines induce viraemia, viral replication in lymphoid tissues and viral faecal shedding (Decaro et al. 2017; Freisl et al. 2017). Vaccinal CPV-2 viraemia and faecal shedding were observed for a short period of time after vaccine administration, up to 24 and 20–28 days, respectively (Decaro et al. 2014; Freisl et al. 2017; Segev et al. 2022). Viral DNA load in blood and faeces of vaccinated dogs, detected with molecular methods, has been observed to be considerably lower and DNA present for a shorter period compared to samples of dogs naturally infected with CPV-2 field strains (Decaro et al. 2014, 2017). Therefore, vaccination against CPV could potentially interfere with diagnostic tests, especially when recently administered in dogs showing clinical signs of acute gastroenteritis (Decaro et al. 2007a; Segev et al. 2022).
In this molecular survey, CPV-2 remains the most frequent canine enteric virus detected in tissue samples, as previously observed in other studies investigating its role as causative agent of infection or death in dogs (Duijvestijn et al. 2016; Dowgier et al. 2017). Despite this potential occurrence, currently, there are no molecular studies that evaluated the presence of CPV vaccine virus in canine tissues. In this study, we report evidence of CPV vaccine strains in tissue samples analysed during routine molecular analyses supporting diagnostic necropsies. The genome of CPV MLV vaccine strains was detected only in the intestine, spleen and mesenteric lymph nodes of five young dead dogs. The vaccinal origin of the CPV DNA was confirmed by the sequence analysis, viral typing, and the comparison with VP2 gene sequences of vaccinal strains. All CPV vaccine strains were characterized as original CPV-2 type (VP2 87 M, 101I, 297 S, 300 A, 305D, 426 N) and four strains were identified as related to 154 or NL-35-D strains; information of vaccine administration for dog id. 2020PA72035 was not available and, thus, the vaccine of origin could not been ascertained.
By using a set of real-time PCR assays, the vaccinal origin of CPV strains was confirmed and CPV-2 viral loads reached different titers in spleen, intestine, and mesenteric lymph nodes; the highest CPV DNA titer (9.95 × 104 DNA copies/10 µl of a template) was detected in the spleen of dog id. 2021PA30130, whereas the lowest CPV DNA titer (6.3 × 102 DNA copies/10 µl of template) was detected in the intestine of dog id. 2020PA72035. Viral loads in organs, similarly to those detected in blood and faeces of vaccinated dogs (Decaro et al. 2014), were lower than those reported in faeces and tissues of dogs naturally infected with field strains (Decaro et al. 2005, 2007b). The presence of CPV DNA was observed up to 24 days after last vaccination but, since this was not an experimental assay and the date of last vaccination for some dogs was missing, it was not possible to define the post-vaccination loads trend in organs. Nonetheless, compared to available data on CPV post-vaccinal faecal shedding in immunized pups (Decaro et al. 2014), DNA of CPV strains of vaccine origin in tissue samples was detected for a slightly longer period with higher viral loads (up to 24 days, 9.95 × 104 DNA copies/10 µl of a template), compared to faeces.
To the best of our knowledge, these results represent the first description of the occurrence of CPV vaccinal strains in tissue samples of recently vaccinated dogs. Indeed, CPV post-vaccinal faecal shedding was experimentally described (Decaro et al. 2014; Freisl et al. 2017) and, occasionally, detected (Meggiolaro et al. 2017; Hoang et al. 2019; Decaro et al. 2005; Tucciarone et al. 2018; Franzo et al. 2019; Jiang et al. 2021) in rectal/faecal swabs or faeces. Similarly, post-vaccine feline panleukopenia virus faecal shedding was experimentally described in cats (Bergmann et al. 2019; Jacobson et al. 2022) or detected in diagnostic samples (Decaro et al. 2008; Van Brussel et al. 2019; Smith et al. 2021), suggesting the need for attention when interpreting FPV PCR positive results in recently vaccinated cats.
This data updates the current information on the persistence of CPV vaccine strains in canine tissues and, their persistence, may interfere with laboratory diagnostic testing (Decaro et al. 2017; Freisl et al. 2017; Segev et al. 2022). Therefore, this study underlines the need for an assay (quantitative real-time PCR assays, MGB probe assays and/or molecular analyses) to differentiate vaccine virus from field strains for an accurate diagnosis of CPV when the virus is detected from canine tissues. Albeit quantitative real-time PCR could be helpful to differentiate naturally infected from vaccinated dogs (Veir et al. 2009), MGB probes or other molecular assays and sequence analysis are more reliable to identify and characterise vaccine and field strains. The use of MGB assays to characterize samples that have been already subjected to sequencing could seem needless since sequence analysis should provide a full characterization of the viral type. However, the advantage of the MGB assays is that they are able to detect the presence of more than one CPV strains in the same sample, even in the presence of low viral titers. In contrast, in the presence of mixed populations sequence analysis may take into account only the predominant strains, thus misdiagnosing eventual coinfections with low-titer strains.
Following the diagnostic approach herein described, the sequence analysis of the CPV VP2 gene fragment made it possible to obtain useful information to elucidate the significance of the PCR positive results. The high degree of nucleotide identity with sequences of viruses currently used in vaccines and the evidence of specific amino acid residues in the same genomic fragment allowed to assess the origin of the CPV strains. Moreover, the determination of the nucleotide sequence of vaccine strains allowed to obtain further genomic information, particularly for the vaccinal CPV strains for which a specific molecular assay is not yet available. Some potential diagnostic limitations (i.e., the load of detection of vaccinal strains with the classic PCR assay, the occurrence of double peaks in the chromatogram during sequencing when co-infections occurrs, the availability or use of newly produced or different vaccines) could be considered following this diagnostic approach.
In conclusions, the present study describes the evidence of vaccine-derived CPV-2 DNA in a small percentage of tissue sample tested for diagnostic purposes, updating the current knowledge on the possible interference of residual DNA of vaccinal origin with molecular assays, which could lead to laboratory misdiagnoses, especially in those cases involving dogs with unknown anamnesis and vaccination status.
Data Availability
Sequence data generated during and analysed during the current study are available in the DDBJ/EMBL/GenBank databases under accession numbers ON479057-ON479067.
References
AAHA (2022) Vaccination recommendations for general practice https://www.aaha.org/aaha-guidelines/vaccination-canine-configuration/vaccination-recommendations-for-general-practice/ Accessed 12
Barrett T, Visser IKG, Mamaev L et al (1993) Dolphin and Porpoise Morbilliviruses Are Genetically Distinct from Phocine Distemper Virus. Virology 193:1010–1012. https://doi.org/10.1006/viro.1993.1217
Bergmann M, Schwertler S, Speck S et al (2019) Faecal shedding of parvovirus deoxyribonucleic acid following modified live feline panleucopenia virus vaccination in healthy cats. Vet Rec 185:83–83. https://doi.org/10.1136/vr.104661
Bodnar L, Lorusso E, Di Martino B et al (2017) Identification of a novel canine norovirus. Infect Gen Evol 52:75–81. https://doi.org/10.1016/j.meegid.2017.04.020
Buonavoglia C, Martella V, Pratelli A et al (2001) Evidence for evolution of canine parvovirus type 2 in Italy. J Gen Virol 82:3021–3025. https://doi.org/10.1099/0022-1317-82-12-3021
Carmichael LE, Joubert JC, Pollock RV (1981) A modified live canine parvovirus strain with novel plaque characteristics. I. Viral attenuation and dog response. Cornell Vet 71:408–427
Cotmore SF, Agbandje-McKenna M, Canuti M et al (2019) ICTV Virus Taxonomy Profile: Parvoviridae. J Gen Virol 100:367–368. https://doi.org/10.1099/jgv.0.001212
Day MJ, Horzinek MC, Schultz RD, Squires RA (2016) WSAVA Guidelines for the vaccination of dogs and cats: WSAVA Vaccination Guidelines. J Small Anim Pract 57:E1–E45. https://doi.org/10.1111/jsap.2_12431
Decaro N, Desario C, Campolo M et al (2005) Clinical and Virological Findings in Pups Naturally Infected by Canine Parvovirus Type 2 Glu-426 Mutant. J Vet Diagn Invest 17:133–138. https://doi.org/10.1177/104063870501700206
Decaro N, Elia G, Martella V et al (2006a) Characterisation of the canine parvovirus type 2 variants using minor groove binder probe technology. J Virol Methods 133:92–99. https://doi.org/10.1016/j.jviromet.2005.10.026
Decaro N, Elia G, Desario C et al (2006b) A minor groove binder probe real-time PCR assay for discrimination between type 2-based vaccines and field strains of canine parvovirus. J Virol Methods 136:65–70. https://doi.org/10.1016/j.jviromet.2006.03.030
Decaro N, Martella V, Elia G et al (2006c) Diagnostic tools based on minor groove binder probe technology for rapid identification of vaccinal and field strains of canine parvovirus type 2b. J Virol Methods 138:10–16. https://doi.org/10.1016/j.jviromet.2006.07.011
Decaro N, Desario C, Elia G et al (2007a) Occurrence of severe gastroenteritis in pups after canine parvovirus vaccine administration: A clinical and laboratory diagnostic dilemma. Vaccine 25:1161–1166. https://doi.org/10.1016/j.vaccine.2006.10.020
Decaro N, Martella V, Elia G et al (2007b) Tissue distribution of the antigenic variants of canine parvovirus type 2 in dogs. Vet Microbiol 121:39–44. https://doi.org/10.1016/j.vetmic.2006.11.005
Decaro N, Desario C, Miccolupo A et al (2008) Genetic analysis of feline panleukopenia viruses from cats with gastroenteritis. J Gen Virol 89:2290–2298. https://doi.org/10.1099/vir.0.2008/001503-0
Decaro N, Crescenzo G, Desario C et al (2014) Long-term viremia and fecal shedding in pups after modified-live canine parvovirus vaccination. Vaccine 32:3850–3853. https://doi.org/10.1016/j.vaccine.2014.04.050
Decaro N, Buonavoglia C (2017) Canine parvovirus post-vaccination shedding: Interference with diagnostic assays and correlation with host immune status. Vet J 221:23–24. https://doi.org/10.1016/j.tvjl.2017.01.020
Dowgier G, Lorusso E, Decaro N et al (2017) A Molecular Survey for Selected Viral Enteropathogens Revealed a Limited Role of Canine Circovirus in the Development of Canine Acute Gastroenteritis. Vet Microbiol 204:54–58. https://doi.org/10.1016/j.vetmic.2017.04.007
Duijvestijn M, Mughini-Gras L, Schuurman N et al (2016) Enteropathogen Infections in Canine Puppies: (Co-)Occurrence, Clinical Relevance and Risk Factors. Vet Microbiol 195:115–122. https://doi.org/10.1016/j.vetmic.2016.09.006
Ford RB, Larson LJ, McClure KD et al (2017) 2017 AAHA Canine Vaccination Guidelines. J Am Anim Hosp Assoc 53:243–251. https://doi.org/10.5326/JAAHA-MS-6741
Franzo G, Tucciarone CM, Casagrande S et al (2019) Canine parvovirus (CPV) phylogeny is associated with disease severity. Sci Rep 9:11266. https://doi.org/10.1038/s41598-019-47773-6
Freeman MM, Kerin T, Hull J et al (2008) Enhancement of detection and quantification of rotavirus in stool using a modified real-time RT-PCR assay. J Med Virol 80:1489–1496. https://doi.org/10.1002/jmv.21228
Freisl M, Speck S, Truyen U et al (2017) Faecal shedding of canine parvovirus after modified-live vaccination in healthy adult dogs. The Vet J 219:15–21. https://doi.org/10.1016/j.tvjl.2016.11.011
Hall T(1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acid Symposium Series, 41:95–98
Hoang M, Lin W-H, Le VP et al (2019) Molecular epidemiology of canine parvovirus type 2 in Vietnam from November 2016 to February 2018. Virol J 16:52. https://doi.org/10.1186/s12985-019-1159-z
Hu RL, Huang G, Qiu W et al (2001) Detection and Differentiation of CAV-1 and CAV-2 by Polymerase Chain Reaction. Vet Res Commun 25:77–84. https://doi.org/10.1023/A:1006417203856
Jacobson LS, Janke KJ, Ha K et al (2022) Feline panleukopenia virus DNA shedding following modified live virus vaccination in a shelter setting. Vet J 279:105783. https://doi.org/10.1016/j.tvjl.2021.105783
Jiang H, Yu Y, Yang R et al (2021) Detection and molecular epidemiology of canine parvovirus type 2 (CPV-2) circulating in Jilin Province, Northeast China. Comp Immunol Microbiol Dis 74:101602. https://doi.org/10.1016/j.cimid.2020.101602
Kumar S, Stecher G, Li M et al (2018) MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Martella V, Decaro N, Elia G, Buonavoglia C (2005) Surveillance Activity for Canine Parvovirus in Italy. J Vet Med Series B 52:312–315. https://doi.org/10.1111/j.1439-0450.2005.00875.x
Meggiolaro MN, Ly A, Rysnik-Steck B et al (2017) MT-PCR panel detection of canine parvovirus (CPV-2): Vaccine and wild-type CPV-2 can be difficult to differentiate in canine diagnostic fecal samples. Mol Cell Probes 33:20–23. https://doi.org/10.1016/j.mcp.2017.02.007
Meunier PC, Cooper BJ, Appel MJG, Slauson DO (1985) Pathogenesis of Canine Parvovirus Enteritis: The Importance of Viremia. Vet Pathol 22:60–71. https://doi.org/10.1177/030098588502200110
Mira F, Purpari G, Lorusso E et al (2018) Introduction of Asian canine parvovirus in Europe through dog importation. Transbound Emerg Dis 65:16–21. https://doi.org/10.1111/tbed.12747
Mira F, Canuti M, Purpari G et al (2019) Molecular Characterization and Evolutionary Analyses of Carnivore Protoparvovirus 1 NS1 Gene. Viruses 11(4):308. https://doi.org/10.3390/v11040308
Ogbu KI, Chukwudi IC, Mira F et al (2021) Current status and risk factors of canine parvovirus type 2 in North Central Nigeria. Comp Immunol Microbiol Dis 74:101578. https://doi.org/10.1016/j.cimid.2020.101578
Parrish CR, Aquadro CF, Strassheim ML et al (1991) Rapid antigenic-type replacement and DNA sequence evolution of canine parvovirus. J Virol 65:6544–6552. https://doi.org/10.1128/jvi.65.12.6544-6552.1991
Pérez R, Calleros L, Marandino A et al (2014) Phylogenetic and Genome-Wide Deep-Sequencing Analyses of Canine Parvovirus Reveal Co-Infection with Field Variants and Emergence of a Recent Recombinant Strain. PLoS ONE 9:e111779. https://doi.org/10.1371/journal.pone.0111779
Pratelli A, Martella V, Elia G et al (2001) Variation of the sequence in the gene encoding for transmembrane protein M of canine coronavirus (CCV). Mol Cell Probes 15:229–233. https://doi.org/10.1006/mcpr.2001.0364
Purpari G, Mira F, Di Bella S et al(2018) Investigation on Canine parvovirus circulation in dogs from Sicily (Italy) by biomolecular assay. Acta Vet. 68:80–94. https://doi.org/10.2478/acve-2018-0007
Schirò G, Gambino D, Mira F et al (2022) Antimicrobial Resistance (AMR) of Bacteria Isolated from Dogs with Canine Parvovirus (CPV) Infection: The Need for a Rational Use of Antibiotics in Companion Animal Health. Antibiotics 11:142. https://doi.org/10.3390/antibiotics11020142
Segev G, Yaaran T, Maurice S, Baneth G (2022) Effect of sampling site on the diagnosis of canine parvovirus infection in dogs using polymerase chain reaction. J Vet Intern Med 36:591–598. https://doi.org/10.1111/jvim.16373
Smith SL, Afonso MM, Roberts L et al (2021) A virtual biobank for companion animals: A parvovirus pilot study. Vet Rec 189(6):e556. https://doi.org/10.1002/vetr.556
Touihri L, Bouzid I, Daoud R et al (2009) Molecular characterization of canine parvovirus-2 variants circulating in Tunisia. Virus Genes 38:249–258. https://doi.org/10.1007/s11262-008-0314-1
Tucciarone CM, Franzo G, Mazzetto E et al (2018) Molecular insight into Italian canine parvovirus heterogeneity and comparison with the worldwide scenario. Infect Genet Evol 66:171–179. https://doi.org/10.1016/j.meegid.2018.09.021
Van Brussel K, Carrai M, Lin C et al (2019) Distinct Lineages of Feline Parvovirus Associated with Epizootic Outbreaks in Australia, New Zealand and the United Arab Emirates. Viruses 11:1155. https://doi.org/10.3390/v11121155
Veir JK, Duffy AL, Dow SW, Lappin MR (2009) Comparison of quantitative PCR and conventional endpoint PCR for amplification of parvovirus DNA in blood from naturally infected and recently vaccinated dogs. J Vet Intern Med 23:673–786
WSAVA (2016) Vaccination Guidelines. https://wsava.org/global-guidelines/vaccination-guidelines/ Accessed 12 April 2022
Zhang Z, Schwartz S, Wagner L, Miller W (2000) A Greedy Algorithm for Aligning DNA Sequences. J Comput Biol 7:203–214. https://doi.org/10.1089/10665270050081478
Funding
This work was funded by the Ministero della Salute (Italy), Ricerca Corrente IZS SI 03/18 RC “Studio del potenziale zoonosico e caratterizzazione genomica dei virus enterici del cane”.
Author information
Authors and Affiliations
Contributions
GS and FM conceptualized the study. GS, ND, CD, GC, SD, VC, GP, GV, VR, DV, FG, and CC carried out the analyses. GS, FM, ND, CD, VC, GP, and DV analyzed data. GS, FM, ND, VC, GP, DV, AG wrote the manuscript; FM, ND, and AG revised the manuscript for important intellectual content. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Ethics approval
Not applicable.
Declaration of conflict of interest
none.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Schirò, G., Mira, F., Decaro, N. et al. Persistence of DNA from canine parvovirus modified-live virus in canine tissues. Vet Res Commun 47, 567–574 (2023). https://doi.org/10.1007/s11259-022-10008-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11259-022-10008-7