Abstract
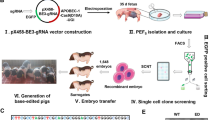
Insulin-like growth factor 2 (IGF2) plays an important role in the development of the foetus and in post-natal growth and development. A SNP within intron 3 of porcine IGF2 disrupts a binding site for the repressor, zinc finger BED-type containing 6 (ZBED6), leading to up-regulation of IGF2 in skeletal muscle and major effects on muscle growth, heart size, and fat deposition. This favourable mutation is common in Western commercial pig populations, but is not present in most indigenous Chinese pig breeds. Here, we described the efficient disruption of the ZBED6 binding site motif in intron 3 of IGF2 by CRISPR/Cas9 in porcine embryonic fibroblasts (PEFs) from the indigenous Chinese pig breed, Liang Guang Small Spotted pig. Disruption of the binding motif led to a drastic up-regulation of IGF2 expression in PEFs and enhanced myogenic potential and cell proliferation of PEFs. IGF2-edited pigs were then generated using somatic cell nuclear transfer. Enhanced muscle development was evident in one pig with biallelic deletion of the ZBED6 binding site motif, implying that the release of ZBED6 repression has a major effect on porcine muscle development. Our study confirmed the important effect of a mutation in the ZBED6 binding site motif on IGF2 expression and myogenesis, thus providing the basis for breeding a new line of Liang Guang Small Spotted pigs with improved lean meat percentage, a trait of great commercial value to pig producers.




Similar content being viewed by others
References
Adli M (2018) The CRISPR tool kit for genome editing and beyond. Nat Commun 9:1911. https://doi.org/10.1038/s41467-018-04252-2
Braunschweig MH, Van Laere AS, Buys N, Andersson L, Andersson G (2004) IGF2 antisense transcript expression in porcine postnatal muscle is affected by a quantitative trait nucleotide in intron 3. Genomics 84:1021–1029. https://doi.org/10.1016/j.ygeno.2004.09.006
Estelle J, Mercade A, Noguera JL, Perez-Enciso MOC, Sanchez A, Folch JM (2005) Effect of the porcine IGF2-intron3-G3072A substitution in an outbred Large White population and in an Iberian × Landrace cross. J Anim Sci 83:2723–2728
Florini JR, Ewton DZ, Coolican SA (1996) Growth hormone and the insulin-like growth factor system in myogenesis. Endocr Rev 17:481–517. https://doi.org/10.1210/edrv-17-5-481
He Z et al (2016) Comparison of surrogate reporter systems for enrichment of cells with mutations induced by genome editors. J Biotechnol 221:49–54. https://doi.org/10.1016/j.jbiotec.2016.01.009
Hsu HH, Zdanowicz MM, Agarwal VR, Speiser PW (1997) Expression of myogenic regulatory factors in normal and dystrophic mice: effects of IGF-1 treatment. Biochem Mol Med 60:142–148
Jeon JT et al (1999) A paternally expressed QTL affecting skeletal and cardiac muscle mass in pigs maps to the IGF2 locus. Nat Genet 21:157–158. https://doi.org/10.1038/5938
Ji Q et al (2013) Improvement of porcine cloning efficiency by trichostain A through early-stage induction of embryo apoptosis. Theriogenology 79:815–823. https://doi.org/10.1016/j.theriogenology.2012.12.010
Ji Q et al (2014) Exogenous expression of OCT4 facilitates oocyte-mediated reprogramming in cloned porcine embryos. Mol Reprod Dev 81:820–832. https://doi.org/10.1002/mrd.22351
Jungerius BJ, Van Laere A-S, Te Pas MFW, Van Oost BA, Andersson L, Groenen MAM (2004) The IGF2-intron3-G3072A substitution explains a major imprinted QTL effect on backfat thickness in a Meishan × European white pig intercross. Genet Res 84:95–101. https://doi.org/10.1017/s0016672304007098
Kim H, Um E, Cho SR, Jung C, Kim JS (2011) Surrogate reporters for enrichment of cells with nuclease-induced mutations. Nat Methods 8:941–943. https://doi.org/10.1038/nmeth.1733
Markljung E et al (2009) ZBED6, a novel transcription factor derived from a domesticated DNA transposon regulates IGF2 expression and muscle growth. PLoS Biol 7:e1000256. https://doi.org/10.1371/journal.pbio.1000256
Nezer C et al (1999) An imprinted QTL with major effect on muscle mass and fat deposition maps to the IGF2 locus in pigs. Nat Genet 21:155–156. https://doi.org/10.1038/5935
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F (2013) Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8:2281–2308. https://doi.org/10.1038/nprot.2013.143
Tan W, Proudfoot C, Lillico SG, Whitelaw CBA (2016) Gene targeting, genome editing: from Dolly to editors. Transgenic Res 25:273–287. https://doi.org/10.1007/s11248-016-9932-x
Van Laere AS et al (2003) A regulatory mutation in IGF2 causes a major QTL effect on muscle growth in the pig. Nature 425:832–836. https://doi.org/10.1038/nature02064
Xiang G et al (2018) Editing porcine IGF2 regulatory element improved meat production in Chinese Bama pigs. Cell Mol Life Sci 75:4619–4628. https://doi.org/10.1007/s00018-018-2917-6
Yang GC, Ren J, Guo YM, Ding NS, Chen CY, Huang LS (2006) Genetic evidence for the origin of an IGF2 quantitative trait nucleotide in Chinese pigs. Anim Genet 37:179–180. https://doi.org/10.1111/j.1365-2052.2006.01416.x
Younis S et al (2018) The ZBED6-IGF2 axis has a major effect on growth of skeletal muscle and internal organs in placental mammals. Proc Natl Acad Sci USA 115:E2048–E2057. https://doi.org/10.1073/pnas.1719278115
Acknowledgements
This work was jointly supported by National Transgenic Major Program (2016ZX08006003-006), the Natural Science Foundation of Guangdong Province (2016A030313310 and 2014030312011) and the Program for Guangdong YangFan Introducting Innovative and Enterpreneurial Teams (2014YT02H042).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, X., Liu, H., Wang, M. et al. Disruption of the ZBED6 binding site in intron 3 of IGF2 by CRISPR/Cas9 leads to enhanced muscle development in Liang Guang Small Spotted pigs. Transgenic Res 28, 141–150 (2019). https://doi.org/10.1007/s11248-018-0107-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11248-018-0107-9