Abstract
The rapid development of next-generation sequencing technologies has made it possible to measure the expression profiles of thousands of genes simultaneously. Often, there exist group structures among genes manifesting biological pathways and functional relationships. Analyzing such high-dimensional and structural datasets can be computationally expensive and results in the complicated models that are hard to interpret. To address this, variable selection such as penalized methods are often taken. Here, we focus on the Cox’s proportional hazards model to deal with censoring data. Most of the existing penalized methods for Cox’s model are the group lasso methods that show deficiencies, including the over-shrinkage problem. In addition, the contemporary algorithms either exhibit the loss of efficiency or require the group-wise orthonormality assumption. Hence, efficient algorithms for general design matrices are needed to enable practical applications. In this paper, we investigate and comprehensively evaluate three group penalized methods for Cox’s model: the group lasso and two nonconvex penalization methods—group SCAD and group MCP—that have several advantages over the group lasso. These methods are able to perform group selection in both non-overlapping and overlapping cases. We have developed the fast and stable algorithms and a new package grpCox to fit these models without the initial orthonormalization step. The runtime of grpCox is improved significantly over the existing packages, such as grpsurv (for the non-overlapping case), grpregOverlap (overlapping), and SGL. In addition, grpCox is better than grpsurv and comparable with SGL in terms of variable selection performances. Comprehensive studies on both simulation and real-world cancer datasets demonstrate the statistical properties of our grpCox implementations with the group lasso, SCAD, and MCP regularization terms.








Similar content being viewed by others
References
Ahmed, M., Rahman, N.: Atm and breast cancer susceptibility. Oncogene 25(43), 5906–11 (2006)
Alsina-Sanchis, E., Figueras, A., Lahiguera Vidal, A., Casanovas, O., Graupera, M., Villanueva, A., Viñals, F.: The tgf pathway stimulates ovarian cancer cell proliferation by increasing igf1r levels. Int. J. Cancer 139(8), 1894–903 (2016)
Alsina-Sanchis, E., Figueras, A., Gil-Martín, M., Pardo, B., Piulats, J.M., Martí, L., Ponce, J., Matias-Guiu, X., Vidal, A., Villanueva, A., Viñals, F.: Tgf controls ovarian cancer cell proliferation. Int. J. Mol. Sci. 18(8) (2017)
Andersen, P.K., Gill, R.D.: Cox’s regression model for counting processes: a large sample study. Ann. Stat. 10(4), 1100–1120 (1982)
Assefnia, S., Dakshanamurthy, S., Guidry-Auvil, J.M., Hampel, C., Anastasiadis, P.Z., Kallakury, B., Uren, A., Foley, D.W., Brown, M.L., Shapiro, L., Brenner, M., Haigh, D., Byers, S.: Cadherin-11 in poor prognosis malignancies and rheumatoid arthritis: common target, common therapies. Oncotarget 5(6), 1458–74 (2014)
Belhechmi, S., De Bin, R., Rotolo, F., Michiels, S.: Accounting for grouped predictor variables or pathways in high-dimensional penalized Cox regression models. BMC Bioinf. 21(277) (2020)
Benjamini, Y., Hochberg, Y.: Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995)
Benjamini, Y., Yekutieli, D.: The control of the false discovery rate in multiple testing under dependency. Ann. Stat.B 29, 1165–1188 (2001)
Bertucci, F., Nasser, V., Granjeaud, S., Eisinger, F., Adelaïde, J., Tagett, R., Loriod, B., Giaconia, A., Benziane, A., Devilard, E., Jacquemier, J., Viens, P., Nguyen, C., Birnbaum, D., Houlgatte, R.: Gene expression profiles of poor-prognosis primary breast cancer correlate with survival. Hum. Mol. Genet. 11(8), 863–72 (2002)
Blighe, K., Lasky-Su, J.: Regparallel: Standard regression functions in r enabled for parallel processing over large data-frames (2021)
Breheny, P., Huang, J.: Group descent algorithms for nonconvex penalized linear and logistic regression models with grouped predictors. Stat. Comput. 25, 173–187 (2015)
Brisson, B.K., Mauldin, E.A., Lei, W., Vogel, L.K., Power, A.M., Lo, A., Dopkin, D., Khanna, C., Wells, R.G., Pure, E.: Estimation of mean sojourn time in breast cancer screening using a Markov chain model of entry to and exit from preclinical detectable phase. Am. J. Pathol. 185(5), 1471–86 (2015)
Cox, D.R.: Regression models and life-tables. J. R. Stat. Soc. B 34(1), 187–220 (1972)
Dang, X.: grpCox: Penalized Cox model for high-dimensional data with grouped predictors. (2020) https://CRAN.R-project.org/package=grpCox, R package version 1.0-1
Etemadmoghadam, D., deFazio, A., Beroukhim, R., Mermel, C.: Integrated genome-wide dna copy number and expression analysis identifies distinct mechanisms of primary chemoresistance in ovarian carcinomas. Clin. Cancer Res. 15(4), 1417–27 (2009)
Fan, J., Li, R.: Variable selection for cox’s proportional hazards model and frailty model. Ann. Stat. 6, 74–99 (2002)
Fan, J., Li, R.: Variable selection via nonconcave penalized likelihood and its oracle properties. J. Am. Stat. Assoc. 96(456), 1348–1360 (2001)
Gatcliffe, T.A., Monk, B.J., Planutis, K., Holcombe, R.F.: Wnt signaling in ovarian tumorigenesis. Int. J. Gynecol. Cancer 18(5), 954–62 (2008)
Gee, M.E., Faraahi, Z., McCormick, A., Edmondson, R.: Dna damage repair in ovarian cancer: unlocking the heterogeneity. J. Ovarian Res. 11(50),(2018)
Goldgar, D.E., Healey, S., Dowty, J.G., Da-Silva, L., Chen, X., Spurdle, A.B., Terry, M.B., Daly, M.J., Buys, S.M., Southey, M.C., Andrulis, I., John, E.M., Khanna, K.K., Hopper, J.L., Oefner, P.J., Lakhani, S., Chenevix-Trench, G.: Rare variants in the atm gene and risk of breast cancer. Breast Cancer Res. 13(4) (2011)
Gui, J., Li, H.: Penalized cox regression analysis in the high-dimensional and low-sample size settings, with applications to microarray gene expression data. Biofinformatics 21(13), 3001–3008 (2005)
Hänzelmann, S., Castelo, R., Guinney, J.: GSVA: gene set variation analysis for microarray and RNA-Seq data. BMC Bioinf. 14(7) (2013)
Hochberg, Y.: A sharper bonferroni procedure for multiple tests of significance. Biometrika 75, 800–80 (1988)
Holm, S.: A simple sequentially rejective multiple test procedure. Scand. J. Stat. 6, 65–70 (1979)
Hommel, G.: A stagewise rejective multiple test procedure based on a modified bonferroni test. Biometrika 75, 383–386 (1988)
Huang, J., Breheny, P., Ma, S.: A selective review of group selection in high-dimensional models. Stat. Sci. 27(4), 481–499 (2012)
Hunter, D., Lange, K.: A tutorial on mm algorithms. Am. Stat. 58(1), 30–37 (2004)
Jacob, L., Obozinski, G., Vert, J.: Group lasso with overlap and graph lasso. In International Conference on Machine Learning, Montreal, Canada, Proceedings of the 26th annual international conference on machine learning, pp. 433–440, (2009)
Jenatton, R., Mairal, G., Obozinski, G., Bach, F.: Proximal methods for hierarchical sparse coding. J. Mach. Learn. Res. 12, 2297–2334 (2011)
Jones, S., Zhang, X., Parsons, D.W., Lin, J.C., Leary, R.J., Angenendt, P., Mankoo, P., Carter, H., Kamiyama, H., Jimeno, A., Hong, S.M., Fu, B., Lin, M.T., Calhoun, E.S., Kamiyama, M., Walter, K., Nikolskaya, T., Nikolsky, Y., Hartigan, J., Smith, D.R., Hidalgo, M., Leach, S.D., Klein, A.P., Jaffee, E.M., Goggins, M., Maitra, A., IacobuzioDonahue, C., Eshleman, J.R., Kern, S.E., Hruban, R.H., Karchin, R., Papadopoulos, N., Parmigiani, G., Vogelstein, B., Velculescu, V.E., Kinzler, K.W.: Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science 321, 1801–1806 (2008)
Kim, Y., Kim, J., Kim, Y.: Blockwise sparse regression. Stat. Sin. 16, 375–390 (2006)
Lange, K., Hunter, D., Yang, I.: Optimization transfer using surrogate objective functions (with discussion). J. Comput. Graph. Stat. 9(1), 1–20 (2000)
Li, Y., Chao, F., Huang, B.: Hoxc8 promotes breast tumorigenesis by transcriptionally facilitating cadherin-11 expression. Oncotarget 5(9), 2596–607 (2014)
Lin, Z., Zhu, G., Tang, D., Bu, J., Zou, J.: High expression of col6a1 correlates with poor prognosis in patients with breast cancer. Int. J. Clin. Exp. Med. 11(11), 12157–12164 (2018)
Loss, L.A., Sadanandam, A., Durinck, S., Nautiyal, S., Flaucher, D., Carlton, V.E., Moorhead, M., Lu, Y., Gray, J.W., Faham, M., Spellman, P., Parvin, B.: Prediction of epigenetically regulated genes in breast cancer cell lines. BMC Bioinf. 11(305) (2010)
Ma, S., Song, X., Huang, J.: Supervised group lasso with applications to microarray data analysis. BMC Bioinf. 8, 60–76 (2007)
Mairal, J., Yu, B.: Complexity analysis of the lasso regularization path (2012)
McCormick, A., Donoghue, P., Dixon, M., O’Sullivan, R., O’Donnell, R., Murray, J., Kaufmann, A., Curtin, N., Edmondson, R.: Ovarian cancers harbour defects in non-homologous end joining resulting in resistance to rucaparib. Clin. Cancer Res. 23(8), 2050–2060 (2017)
Meir, L., Van de Geer, S., Buhlmann, P.: The group lasso for logistic regression. J. R. Stat. Soc. Ser. B (Methodol.) 70(1), 53–71 (2008)
Miller, L.D., Smeds, J., George, J., Vega, V.B., Vergara, L., Ploner, A., Pawitan, Y., Hall, P., Klaar, S., Liu, E.T., Bergh, J.: An expression signature for p53 status in human breast cancer predicts mutation status, transcriptional effects, and patient survival. Proc. Natl. Acad. Sci. U.S.A. 102(38), 13550–13555 (2005)
Molecular signatures database v7.4. (2021) https://www.gsea-msigdb.org/gsea/msigdb/collections.jsp
Obozinski, G., Jacob, L., Vert, J.: Group lasso with overlaps: the latent group lasso approach. arXiv (2011)
Otsuka, A., de Paolis, A., Tocchini-Valentini, G.P.: Ribonuclease “xlai,” an activity from xenopus laevis oocytes that excises intervening sequences from yeast transfer ribonucleic acid precursors. Mol. Cell. Biol. 1(3), 269–280 (1981)
Park, M.Y., Hastie, T.: Penalized logistic regression for detecting gene interactions. Tech report, Stanford University, United States, Tech. Rep (2006)
Puig, A., Wiesel, A., Fleury, G., Hero, A.: Multidimensional shrinkage-thresholding operator and group lasso penalties. IEEE Signal Process. Lett. 18, 363–366 (2011)
Sarrio, D., Rodriguez-Pinilla, S.M., Hardisson, D., Cano, A., Moreno-Bueno, G., Palacios, J.: Epithelial-mesenchymal transition in breast cancer relates to the basal-like phenotype. Cancer Res. 68(4), 989–997 (2008)
Sengupta, P.K., Smith, E.M., Kim, K., Murnane, M.J., Smith, B.D.: Dna hypermethylation near the transcription start site of collagen alpha2(i) gene occurs in both cancer cell lines and primary colorectal cancers. Can. Res. 63, 1789–1797 (2003)
Simon, N.: Regularization paths for coxś proportional hazards model via coordinate descent. J. Stat. Softw. 39(5), 53–66 (2012)
Simon, N., Tibshiran, R.: Standardization and the group lasso penalty. Stat. Sin. 22, 983–1001 (2011)
Simon, N., Friedman, J., Hastie, T., Tibshirani, R.: A sparse-group lasso. J. Comput. Graph. Stat. 22(2), 231–245 (2013)
Subramanian, A., Tamayo, P., Mootha, V.K., Mukherjee, S., Ebert, B.L., Gillette, M.A., Paulovich, A., Pomeroy, S.L., Golub, T.R., Lander, E.S., Mesirov, J.P.: Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. U.S.A. 102(43), 15545–50 (2005)
Szkandera, J., Kiesslich, T., Haybaeck, J., Gerger, A., Pichler, M.: Hedgehog signaling pathway in ovarian cancer. Int. J. Mol. Sci. 14(1), 1179–1196 (2013)
Ternes, N., Rotolo, F., Michiels, S.: Empirical extensions of the lasso penalty to reduce the false discovery rate in high-dimensional Cox regression models. Stat. Med. 35(15), 2561–73 (2016)
Therneau, T.M.: A package for survival analysis in R. https://CRAN.R-project.org/package=survival, R package version 3.2-11 (2021)
Tibshirani, R.: The lasso method for variable selection in the cox model. Stat. Med. 16(4), 385–395 (1996)
Van de Vijer, M.J., He, Y.D., van’t Veer, L.J., Dai, H., Hart, A.A., Voskuil, D., Schreiber, G.J., Peterse, J.L., CW, R., Marton, M.J., Parrish, M., Atsma, D., Witteveen, A., Glas, A., Delahaye, L., van der Velde, T.W., Bartelink, H., Rodenhuis, S., Rutgers, E.T., Friend, S.H., Bernards, R.: A gene-expression signature as a predictor of survival in breast cancer. New Engl. J. Med. 347(25), 1999–2009 (2002)
Verweij, P.J., Houwelingen, H.C.: Cross-validation in survival analysis. Stat. Med. 12(24), 385–395 (1993)
Wang, L., Chen, G., Li, H.: Group scad regression analysis for microarray time course gene expression data. Bioinformatics 23(12), 1486–1494 (2007)
Wang, L., Li, H., Huang, J.: Variable selection in nonparametric varying-coefficient models for analysis of repeated measurements. J. Am. Stat. Assoc. 103(484), 1556–1569 (2008)
Wu, T., Wang, S.: Doubly regularized cox regression for high-dimensional survival data with group structures. Stat. Interface 6, 175–186 (2013)
Xiong, G., Deng, L., Zhu, J., Xu, R.: Prolyl-4-hydroxylase subunit 2 promotes breast cancer progression and metastasis by regulating collagen deposition. BMC Cancer 14(1) (2014)
Yang, Y., Zou, H.: A fast unified algorithm for solving group-lasso penalize learning problems. Stat. Comput. 25, 1129–1141 (2015)
Yuan, M., Lin, Y.: Model selection and estimation in regression with grouped variables. J. R. Stat. Soc. Ser. B (Methodol.) 68(1), 49–67 (2006)
Zeng, Y., Breheny, P.: Overlapping group logistic regression with applications to genetic pathway selection. Cancer Inf. 15, 179–187 (2016)
Zhang, H., Lu, W.: Adaptive lasso for cox’s proportional hazards model. Biometrika 94(3), 691–703 (2007)
Zhang, C.: Nearly unbiased variable selection under minimax concave penalty. Ann. Stat. 38(2), 894–942 (2010)
Zhao, P., Rocha, G., Yu, B.: The composite absolute penalties family for grouped and hierarchical variable selection. Ann. Stat. 37(6A), 3468–3497 (2009)
Zou, H.: A note on path-based variable selection in the penalized proportional hazards model. Biometrika 95(1), 241–247 (2008)
Acknowledgements
We are grateful to three anonymous reviewers for their helpful comments and constructive feedback, which help significantly improve the preliminary version of this paper. We thank Texas A&M High Performance Research Computing for providing computational resources to perform experiments in this work. This work was supported in part by the National Science Foundation (NSF)–Division of Communication & Computing Foundations (CCF) awards #1553281, #1718513, #1715027, NSF–Division of Information & Intelligent Systems (IIS) award #1812641, and the JDRF award #2-SRA-2018-513-S-B.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Appendices
Appendix 1
We have studied the statistical properties of the estimators: consistency and convergence rate as follows.
The partial likelihood
where the penalty term \(P_{\lambda , \gamma }(\beta )\) can be denoted as \(P_{\lambda _n}(\beta )\) since \(\gamma \) for group SCAD and group MCP are fixed. Here, \(\ell _n(\beta ), \lambda _n\) denote the partial likelihood and tuning parameter changing with the sample size n, respectively.
Let the true parameter be \(\beta _0 = \big ( \beta _{01}^T, \beta _{02}^T \big )^T\) where \(\beta _{01}\) consists of all nonzero groups and \(\beta _{02}\) consists of all remaining zero groups. The objective function is
Correspondingly, the minimizer of \({\mathcal {Q}}_n(\beta , \lambda _n)\) is \(\beta _n = \big ( \beta _{n1}^T, \beta _{n2}^T \big )^T\) where \({\beta _n} = \underset{\beta }{\text {argmin }}{\mathcal {Q}}_n(\beta , \lambda _n)\).
Define \(a_n = \text {max} \{ P^{'}_{\lambda _n}(\Vert \beta _{j0}\Vert ): \Vert \beta _{j0}\Vert \ne 0 \}\) and \(b_n = \text {max} \{ P^{''}_{\lambda _n}(\Vert \beta _{j0}\Vert ): \Vert \beta _{j0}\Vert \ne 0 \}\).
Theorem 1
(Consistency and convergence rate) If \( P_{\lambda _n}(\Vert \beta \Vert )\) simultaneously satisfies two conditions: \(a_n = O_p(n^{-1/2})\) and \(b_n \rightarrow 0\), then \(\beta _n\) is a root-n consistent estimator for \(\beta _0\) with rate \(n^{-1/2}\), i.e. \(\Vert \beta _n - \beta _0\Vert = O_p(n^{-1/2})\).
Proof
According to Theorem 3.2 in Andersen and Gill (1982) two results hold
where \(\Sigma \) is the positive definite Fisher information matrix.
Consider a constant ball, \(B(C) = \{ \beta _0 + \alpha _n\mathbf{u }: \Vert \mathbf{u }\Vert \le C \}\) and its boundary \(\partial B(C)\) where \(C>0\) and \(\alpha _n = n^{-1/2} + a_n\). Therefore, \(O_p(\alpha _n) = O_p(a_n) = O_p(n^{-1/2})\). To prove \(\Vert \beta _n - \beta _0\Vert = O_p(n^{-1/2})\), it is sufficient to prove that for any \(\epsilon >0\), there exists a large constant C such that
This implies that with probability at least \(1-\epsilon \) (or goes to 1), \(Q_n(\beta , \lambda _n)\) has a local minimum in the ball B(C) for a given \(\lambda _n\).
Denote \(D_n(\mathbf{u }) = Q_n(\beta , \lambda _n) - Q(\beta _0, \lambda _n)\), we have
Consider that
Consider \(D_2\) using Taylor expansion, we have
Because \(b_n \rightarrow 0\), \(D_2 \rightarrow O_p(C\alpha _n^2)\). By choosing a sufficiently large C, \(D_1\) dominates \(D_2\). Thus, inequality (15) holds.
Appendix 2
We present the simulation studies of the second cross-validation approach described in Section 2.7 to select the tuning parameters \(\lambda \) and evaluate its variable selection performance.
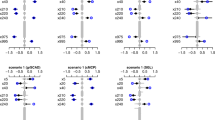
In Fig. 8, each dot represents the logarithm of the \(\lambda \) values along the solution path, and the error bars provide the confidence intervals for the cross-validation log-partial-likelihood. The left vertical bar indicates the maximum cross-validation partial-log-likelihood using the first method Verweij and Houwelingen (1993) while the right one shows the maximum cross-validation log-partial-likelihood using the second method Ternes et al. (2016).
We continue considering \(N=100\) observations and \(P=400\) covariates with 40 groups, each with 10 elements. There are two non-zero groups. The coefficient magnitude \(|\beta | = 0.5\), the values of the population correlation \(\rho \) are 0, 0.2 and 0.5, the censoring rates are 0% and 20%. The results are summarized in Tables 11, 12, and 13 . It can be seen that using the second cross-validation method always results in smaller models than using the first cross-validation method. For group lasso, it produces better variable selection results with much smaller FPR values. For group SCAD and MCP, it often gives better results, but sometimes suppresses too much, e.g., in group MCP case with 20% censoring, \(\rho =0.5\). Therefore, the second cross-validation method should be used with caution.
Appendix 3
We present additional settings based on the reviewer’s suggestions: settings with a large number of overlapping covariates and the number of zero groups being more than the number of non-zero groups. More specifically, we have performed an additional experiment using the simulated data with \(N=100\), \(P=55\), in which there are 10 groups of size 10 and 50% covariates overlap between two successive groups. The “correct” underlying group structure is given by
We set the population correlation \(\rho =0.5\) with 30% censoring rate. The corresponding coefficients are
Then we consider four setups with the misspecified group structures for inference. In the first setup, the number of groups are incorrect because the overlapping groups are collapsed as follows:
In the second setup, the misspecified group structure deviates from the ground truth more significantly will all the overlapping covariates put into one group:
Similar as the first setup, the third and fourth setups are defined as follows:
and
The results shown in Table 14 confirm our expectation: the setup with the collapsed groups including several non-zero (active) groups produces worse results than the cases with the collapsed groups with none or only one non-zero group. More clearly, the first setup in the table including two collapsed groups (group3 and group5), where each of them consists of two non-zero groups, has the worst variable selection performance. Both the second and third misspecification setups including only one group (group5) that is collapsed from two non-zero groups have almost the same performance, better than the first misspecification setup. The fourth mispecification setup with no misspecified group collapsed from two non-zero groups has the best performance. We hypothesize that the probability of variables being incorrectly selected increases due to the ignorance of the overlapping property of active elements in the collapsed groups and the larger group sizes of these collapsed groups. In other words, FPR increases and then corresponding RMSE increases.
Rights and permissions
About this article
Cite this article
Dang, X., Huang, S. & Qian, X. Penalized Cox’s proportional hazards model for high-dimensional survival data with grouped predictors. Stat Comput 31, 77 (2021). https://doi.org/10.1007/s11222-021-10052-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11222-021-10052-4