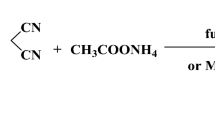Abstract
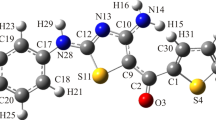
The Gaussian09 program was studied by using the optimized molecular structure with vibrational frequency assignments of 2, 3, 9, and 10 tetrahydroacridin-3-one (abbreviated as THA-3-one). The VEDA software is used to compute the potential energy distribution of the vibrational modes. The optimized geometrical parameters were in agreement with related derivatives. Time-dependent density functional theory estimates oscillator strength and energy, which are almost in line with experimental discoveries. In order to accomplish the gauge-including atomic orbital 1H and 13CNMR chemical shift predictions, the B3LYP functional with 6–311 + + G (d, p) basis sets was utilized. The charge movement within the compound is determined using the HOMO and LUMO investigations. The MEP was revealed using the DFT approach, and the infrared intensities have also been published. Milliken net charges are contrasted with the atomic natural charges. In vitro biological THA-3-one was discovered potent bactericidal activity with a maximum of (20.5 ± 1.0 mm) at 2.5 µg/mL against Yersinia enterocolitica (MTCC 840) and showed a maximum scavenging property of 73.9 ± 1.5% at the highest concentrations of 1000 μg/ML and also showed a maximum scavenging property of 73.9 ± 1.5% at the highest concentrations of 1000 μg/mL, respectively (p < 0.05). In silico molecular docking showed that ligand interacts with (PDB Code: 5AA9) protein by using AutoDock software, which exhibits a lower binding energy assessment of − 6.8 (Kcal/mol) and inhibition constant (ki) value of 10.234 μM.












Similar content being viewed by others
Data Availability
Publisher’s Note: Springer Nature remains neutral with regard to jurisdictional claims in published Maps and institutional affiliations.
References
R. Kavinilavan, P. Mekala, M.J. Raja, M. Arthanari Eswaran, G. Thirumalaisamy, J. Pharmacogn. Phytochem. 6, 749 (2017)
A.H. Gilani, A.U. Rahman, J. Ethanopharmacol. 100, 43 (2005)
N.G. Cuellar, J. Altern. Complement. Med. 13, 179 (2007)
G.J. Christian, M. Subramanian, D. Periyasami, K. Manickavasakam, P. Gunasekaran, S. Sivasubramanian, M. Nijavizhi, Int. J. Pharm. Sci. Rev. Res. 6, 1656 (2015)
K. Anbarasu, K.K. Manisenthil, S. Ramachandran, Asian Pac. J. Trop. Med. 4, 819 (2011)
R. Kalaiarasi, R. Jeeva Gladys, S. Elangovan, D.K. Soundararajan, H. Mubarak, A. Kanakarajan, Int. J. Curr. Res. 5, 978 (2013)
T.S. Ram, M. Munikumar, V.N. Raju, P. Devaraj, N.K. Boiroju, R. Hemalatha, P.V.V. Prasad, M. Gundeti, B.S. Sisodia, S. Pawar, G.P. Prasad, J. Ayurveda Integr. Med. 13, 100413 (2022)
A.J. Gandhi, J.D. Rupareliya, V.J. Shukla, S.B. Donga, R. Acharya, J. Ayurveda Integr. Med. 21, 100343 (2020)
P. Kamalarajan, S. Muthuraman, M.R. Ganesh, M.F. Valan, Eur. J. Med. Plants. 30, 1 (2019)
K. Anbarasu, K.K. Manisenthil, S. Ramachandran, Asian Pac J. Trop Med. 4, 819 (2011)
O.I. Aruoma, Food. Chem. Toxicol. 32, 671 (1994)
L. Seasotiya, P. Siwach, A. Malik, S. Bai, P. Bharti, S. Dlal, Int. J. Adv. Pharm. Biol. Chem. 3, 604 (2014)
M. Mudhafar, H. Alsailawi, M. Abdulrasool, R.K.M. Jawad, A. Mays, Int. J. Chem. Res. 1, 7 (2021)
N. Ozsoy, A. Can, R. Yanardag, N. Akev, Food. Chem. 110, 571 (2008)
R.W. Snow, C.A. Guerra, A.M. Noor, H.Y. Myint, S.I. Hay, Nature 434, 214 (2005)
D.A. Fidock, Nature 465, 297 (2010)
K. Buchholz, T.A. Burke, K.C. Williamson, R.C. Wiegand, D.F. Wirth, M. Marti, J. Infect. Dis. 203, 1445 (2011)
T. Eicher, S. Hauptmann, George Thieme Verlag, Stuttgart, New York. 354 (1995)
K.M. Hujer, A.M. Hujer, E.A. Hulten, S. Bajaksouzian, J.M. Adams, C.J. Donskey, D.J. Ecker, C. Massire, M.W. Eshoo, R. Sampath, J.M. Thomson, Antimicrob. Agents Chemother. 12, 4114 (2006)
A. Mahmood, A.H. Allah, A. Asim, H.I. Balakit, A.A. Salman, Y.S. Abdulridha, J. Adhes. Sci. Technol. 1, 23 (2022)
A.A. Abdulridha, M.A.A.H. Allah, S.Q. Makki, Y. Sert, H.E. Salman, A.A. Balakit, J. Mol. Liq. 315, 113690 (2020)
A.A. Balakit, S.Q. Makki, Y. Sert, F. Ucun, M.B. Alshammari, P. Thordarson, G.A. El-Hiti, Supramol Chem 32, 519 (2020)
N. Dege, H. Gökce, O.E. Doğan, G. Alpaslan, T. Ağar, S. Muthu, Y. Sert, Colloids Surf. A Physicochem. Eng. 638, 128311 (2022)
Y.M. Ayuba, B.C. Okora, M.N. Abubakar, E. Hesanmi, Int. J. Biochem. Biophy. MolBiol. 2, 1 (2017)
M.J. Frisch, G.W. Trucks, H.B. Schlegel, G.E. Scuseria, M.A. Robb, J.R. Cheeseman, G. Scalmani, V. Barone, B. Mennucci, G.A. Petersson, H. Nakatsuji, Revision D. 01, Wallingford, CT 201, Gaussian. Inc., 2009.
J. Irshad Ahamed, K. Francy, A.V. Priya, J. Prema Kumari, R.P. Steiny, P. Kamalarajan, B. Venkatadri, J. Mol. Struct. 1252, 132186 (2022)
H.B. Schlegel, J. Comput. Chem. 2, 214 (1982)
J. Irshad Ahamed, K. Narendran, V.R. Ambika, R. Priya, P. Kamalarajan, T. Sundareswaran, B. Gunasekaran, S. Jayalakshmi, J. Mol. Struct. 1266, 133548 (2022)
J. Irshad Ahamed, M. Priya, P. Vinothkumar, K. Sathyamoorthy, P. Murali- Manohar, J. Liu, M.F. Valan, J. Mol. Struct. 1202, 127241 (2020)
J. Irshad Ahamed, M.F. Valan, K. Pandurengan, P. Agastian, B. Venkatadri, M.R. Rameshkumar, K. Narendran, Res. Chem. Intermed. 47, 759 (2021)
J. Irshad Ahamed, G.R. Ramkumaar, P. Kamalarajan, K. Narendran, M.F. Valan, T. Sundareswaran, T.A. Sundaravadivel, B. Venkatadri, S. Bharathi, J. Mol. Struct. 1248, 131418 (2022)
A.D. Becke, J. Chem. Phys. 98, 5648 (1993)
C. Lee, W. Yang, R.G. Parr, Phys. Rev. B 37, 785 (1988)
H. Ullah, A.-H.A. Shah, K. Ayub, S. Bilal, J. Phys. Chem. C. 117, 4069 (2013)
I. Javed, A. Khurshid, M.N. Arshad, Y. Wang, New J. Chem. 38, 752 (2014)
A. Babar, H. Khalid, K. Ayub, S. Saleem, A. Waseem, T. Mahmood, M.A. Munawar, G. Abbas, A.F. Khan, J. Mol. Struct. 1072, 221 (2014)
T.Q. Hung, N.N. Thang, T.T. Dang, K. Ayub, A. Villinger, A. Friedrich, S. Lochbrunner, G.U. Flechsig, P. Langer, Org. Biomol. Chem. 12, 6151 (2014)
M.A. Hashmi, A. Khan, K. Ayub, U. Farooq, Spectro Chim. Acta A Mol. Biomol. Spectrosc. 128, 225 (2014)
H. Ullah, A. Rauf, Z. Ullah, M. Anwar, G. Uddin, K. Ayub, Spectrochim. Acta A Mol. Biomol. Spectrosc. 118, 210 (2014)
M.K. Abdel-Latif, H.R. Abd El-Mageed, H.S. Mohamed, F.M. Mustafa, J. Mol. Struct. 1200, 127056 (2020)
P. Ramesh, M.L. Caroline, S. Muthu, B. Narayana, M. Raja, S. Aayisha, J. Mol. Struct. 1200, 127123 (2020)
R. Ditchfield, J. Chem. Phys. 56, 5688 (1972)
R.J. Ruch, S.J. Cheng, J.E. Klaunig, Carcinogenesis 10, 1003 (1989)
B. Venkatadri, A. Khusro, C. Aarti, M.R. Rameshkumar, P. Agastian, Asian. Pac. J Trop. Biomed. 7, 782 (2017)
G.M. Morris, D.S. Goodsell, R.S. Halliday, R. Huey, W.E. Hart, R.K. Belew, A.J. Olson, J. Comput. Chem. 19, 1639 (1998)
Visualizer DS. Accelrys software inc. Discovery Studio Visualizer, (2005)
L. J. Bellamy, He IR-Spectra of Complex Molecules, vol. 205. John Wiley and Sons, New York (1975)
A. Spire, M. Barthes, H. Kellouai, G. De Nunzio, Nonlinear Phenomena. 137, 392 (2000)
K.D. Doney, D. Zhao, J.F. Stanton, H. Linnartz, Phys. Chem. Chem. Phys. 20, 5501 (2018)
H. Tanak, Y. Köysal, Y. Ünver, M. Yavuz, S. Işık, K. Sancak, Mol. Phys. 108, 127 (2010)
S. Muthu, E.I. Paulraj, Solid State Sci. 14, 476 (2012)
S. Sakthivel, T. Alagesan, S. Muthu, C.S. Abraham, E. Geetha, J. Mol. Struct. 1156, 645 (2018)
P.J. Brimmer, P.R. Griffiths, Appl. Spectrosc. 42, 242 (1988)
P.J. Larkin, M.P. Makowski, N.B. Colthup, Spectrochim. Acta A 55, 1011 (1999)
K. Wolinski, J.F. Hinton, P. Pulay, J. Am. Chem. Soc. 112, 8251 (1990)
A.A. Rostami, A. Godarzian, J. Phys. Soc. Japan. 74, 1609 (2005)
C.M. Rohlfing, L.C. Allen, R. Ditchfield, Chem. Phys. 87, 9 (1984)
Y. Atalay, D. Avcı, A. Başoğlu, Struct. Chem. 19, 239 (2008)
T. Vijayakumar, I.H. Joe, C.R. Nair, V.S. Jayakumar, Chem. Phys. 343, 83 (2008)
A. Rauk, Orbital interaction theory of organic chemistry, 2nd edn. (John Wiley & Sons, New York, 2001)
A. Teimouri, A.N. Chermahini, K. Taban, H.A. Dabbagh, Spectrochem. Acta A Mol. Biomol Spectrosc. 72, 369 (2008)
E. Rezkallah, A. Ibrahim, A. Dahy, A.A. Hakiem, R. Mahfouz, Z Phys Chem. 233, 1503 (2019)
P. Singh, S.S. Islam, H. Ahmad, A. Prabaharan, J. Mol. Struct. 1154, 39 (2018)
P. Singh, A. Prabaharan, H. Ahmad, S.S. Islam, Int. J. Curr. Res 10, 66290 (2018)
S. Celik, S. Akyuz, A.E. Ozel, J. Mol. Struct. 1258, 132693 (2022)
S. Celik, S. Akyuz, A.E. Ozel, Mol. Cryst. Liq. 1258, 132693 (2022)
W. Tian, C. Chen, X. Lei, J. Zhao, J. Liang, CASTp 3.0: computed atlas of surface topography of proteins. Nucleic acids research, 46, W363 (2018)
Acknowledgements
The authors are thankful to Dr. G. JeyaJothi, Taxnomist, Department of plant Biology and Biotechnology, Loyola College, Chennai, for the authentication of the plant species. The authors thank J. Irshad Ahamed and P. Kamalarajan for kindly providing us access to the all spectral studies for performing the experiments. The authors also thank R. Priya & M. F. Valan for kindly providing the molecular docking studies.
Funding
The authors confirm that they have no known financial or interpersonal conflicts that may have looked to have influenced the research presented in this study.
Author information
Authors and Affiliations
Contributions
J. Irshad Ahamed was responsible for conceptualization, investigation, writing the original draft, and writing, reviewing, and editing. P. Kamalarajan took part in investigation and writing. R. Priya and M. F. Valan contributed to molecular docking and DFT computational studies.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
The authors declare that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kamalarajan, P., Irshad Ahamed, J., Priya, R. et al. Spectroscopic, computational DFT, in vitro, and molecular docking investigations of newly isolated 2, 3, 9, and 10-tetrahydroacridin-3-one from the methanolic extract of nilavembu kudineer chooranam. Res Chem Intermed 49, 2669–2690 (2023). https://doi.org/10.1007/s11164-022-04906-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11164-022-04906-3