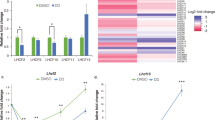
Abstract
Diatoms are a diverse group of photosynthetic unicellular algae with a plastid of red-algal origin. As prolific primary producers in the ocean, diatoms fix as much carbon as all rainforests combined. The molecular mechanisms that contribute to the high photosynthetic productivity and ecological success of diatoms are however not yet fully understood. Using the model diatom Phaeodactylum tricornutum, here we show rhythmic transcript accumulation of plastid psaA, psbA, petB, and atpB genes as driven by a free running circadian clock. Treatment with the electron transport inhibitor 3-(3,4-dichlorophenyl)-1,1-dimethylurea overrides the circadian signal by markedly downregulating transcription of psaA, petB, and atpB genes but not the psbA gene. Changes in light quantity produce little change in plastid gene transcription while the effect of light quality seems modest with only the psaA gene responding in a pattern that is dependent on the redox state of the plastoquinone pool. The significance of these plastid transcriptional responses and the identity of the underlying genetic control systems are discussed with relevance to diatom photosynthetic acclimation.







Similar content being viewed by others
Data availability
The datasets from this study will be assigned a stable DOI and published on the Purdue University Research Repository (PURR): (https://purr.purdue.edu).
References
Allen JF (2015) Why chloroplasts and mitochondria retain their own genomes and genetic systems: colocation for redox regulation of gene expression. Proc Natl Acad Sci USA 112:201500012. https://doi.org/10.1073/pnas.1500012112
Allen AE, LaRoche J, Maheswari U et al (2008) Whole-cell response of the pennate diatom Phaeodactylum tricornutum to iron starvation. Proc Natl Acad Sci USA 105:10438–10443. https://doi.org/10.1073/pnas.0711370105
Annunziata R, Ritter A, Fortunato AE et al (2019) BHLH-PAS protein RITMO1 regulates diel biological rhythms in the marine diatom Phaeodactylum tricornutum. Proc Natl Acad Sci USA 116:13137–13142. https://doi.org/10.1073/pnas.1819660116
Armbrust EV (2009) The life of diatoms in the world’s oceans. Nature 459:185–192
Baginsky S, Tiller K, Pfannschmidt T, Link G (1999) PTK, the chloroplast RNA polymerase-associated protein kinase from mustard (Sinapis alba), mediates redox control of plastid in vitro transcription. Plant Mol Biol 39:1013–1023
Bailleul B, Rogato A, De Martino A et al (2010) An atypical member of the light-harvesting complex stress-related protein family modulates diatom responses to light. Proc Natl Acad Sci USA 107:18214–18219. https://doi.org/10.1073/pnas.1007703107
Bailleul B, Berne N, Murik O et al (2015) Energetic coupling between plastids and mitochondria drives CO2 assimilation in diatoms. Nature 524:366–369. https://doi.org/10.1038/nature14599
Barkan A (2011) Expression of plastid genes: organelle-specific elaborations on a prokaryotic scaffold. Plant Physiol 155:1520–1532. https://doi.org/10.1104/pp.110.171231
Bowler C, Allen AE, Badger JH et al (2008) The Phaeodactylum genome reveals the evolutionary history of diatom genomes. Nature 456:239–244. https://doi.org/10.1038/nature07410
Bruce Cahoon A, Stern DB (2001) Plastid transcription: a menage a trois? Trends Plant Sci 6:45–46. https://doi.org/10.1016/S1360-1385(00)01858-6
Büchel C (2019) How diatoms harvest light. Science 365:447–448
Buck JM, Sherman J, Bártulos CR et al (2019) Lhcx proteins provide photoprotection via thermal dissipation of absorbed light in the diatom Phaeodactylum tricornutum. Nat Commun 10:4167. https://doi.org/10.1038/s41467-019-12043-6
Choquet Y, Vallon O (2000) Synthesis, assembly and degradation of thylakoid membrane proteins. Biochimie 82:615–634
Chow WS, Melis A, Anderson JM (1990) Adjustments of photosystem stoichiometry in chloroplasts improve the quantum efficiency of photosynthesis. Proc Natl Acad Sci USA 87:7502–7506. https://doi.org/10.1073/pnas.87.19.7502
Dagan T, Martin W (2009) Seeing green and red in diatom genomes. Science 324:1651–1652
Deschamps P, Moreira D (2012) Reevaluating the green contribution to diatom genomes. Genome Biol Evol 4:683–688. https://doi.org/10.1093/gbe/evs053
Dodd AN, Salathia N, Hall A et al (2005) Cell biology: plant circadian clocks increase photosynthesis, growth, survival, and competitive advantage. Science 309:630–633. https://doi.org/10.1126/science.1115581
Doran E, Cattolico RA (1997) Photoregulation of chloroplast gene transcription in the chromophytic alga Heterosigma carterae. Plant Physiol 115:773–781
Dorrell RG, Gile G, McCallum G et al (2017) Chimeric origins of ochrophytes and haptophytes revealed through an ancient plastid proteome. Elife. https://doi.org/10.7554/eLife.23717
El Bissati K, Kirilovsky D (2001) Regulation of psbA and psaE expression by light quality in Synechocystis species PCC 6803. A redox control mechanism. Plant Physiol 125:1988–2000. https://doi.org/10.1104/pp.125.4.1988
Escoubas JM, Lomas M, LaRoche J, Falkowski PG (1995) Light intensity regulation of cab gene transcription is signaled by the redox state of the plastoquinone pool. Proc Natl Acad Sci USA 92:10237–10241
Fortunato AE, Jaubert M, Enomoto G et al (2016) Diatom phytochromes reveal the existence of far-red-light-based sensing in the ocean. Plant Cell 28:616–628. https://doi.org/10.1105/tpc.15.00928
Gibbs SP (1981) The chloroplasts of some algal groups may have evolved from endosymbiotic eukarotic algae. Ann NY Acad Sci 361:193–208. https://doi.org/10.1111/j.1749-6632.1981.tb54365.x
Giordano M, Beardall J, Raven JA (2005) CO2 concentrating mechanisms in algae: mechanisms, environmental modulation, and evolution. Annu Rev Plant Biol 56:99–131. https://doi.org/10.1146/annurev.arplant.56.032604.144052
Gross M (2012) The mysteries of the diatoms. Curr Biol 22:R581–R585
Grouneva I, Rokka A, Aro EM (2011) The thylakoid membrane proteome of two marine diatoms outlines both diatom-specific and species-specific features of the photosynthetic machinery. J Proteome Res 10:5338–5353. https://doi.org/10.1021/pr200600f
Harmer SL (2009) The circadian system in higher plants. Annu Rev Plant Biol 60:357–377. https://doi.org/10.1146/annurev.arplant.043008.092054
Herbstová M, Bína D, Koník P et al (2015) Molecular basis of chromatic adaptation in pennate diatom Phaeodactylum tricornutum. Biochim Biophys Acta Bioenerg 1847:534–543. https://doi.org/10.1016/j.bbabio.2015.02.016
Herranen M, Tyystjärvi T, Aro EM (2005) Regulation of photosystem I reaction center genes in Synechocystis sp. strain PCC 6803 during light acclimation. Plant Cell Physiol 46:1484–1493. https://doi.org/10.1093/pcp/pci160
Ibrahim IM, Wu H, Ezhov R et al (2020) An evolutionarily conserved iron-sulfur cluster underlies redox sensory function of the chloroplast sensor kinase. Commun Biol 3:1–11. https://doi.org/10.1038/s42003-019-0728-4
Jovanovic M, Rooney MS, Mertins P et al (2015) Dynamic profiling of the protein life cycle in response to pathogens. Science. https://doi.org/10.1126/science.1259038
Jungandreas A, Costa BS, Jakob T et al (2014) The acclimation of Phaeodactylum tricornutum to blue and red light does not influence the photosynthetic light reaction but strongly disturbs the carbon allocation pattern. PLoS ONE 9:e99727. https://doi.org/10.1371/journal.pone.0099727
Kramer DM, Johnson G, Kiirats O, Edwards GE (2004) New fluorescence parameters for the determination of QA redox state and excitation energy fluxes. Photosynth Res 79:209. https://doi.org/10.1023/B:PRES.0000015391.99477.0d
Lepetit B, Sturm S, Rogato A et al (2013) High light acclimation in the secondary plastids containing diatom Phaeodactylum tricornutum is triggered by the redox state of the plastoquinone pool. Plant Physiol 161:853–865. https://doi.org/10.1104/pp.112.207811
Levitan O, Chen M, Kuang X et al (2019) Structural and functional analyses of photosystem II in the marine diatom Phaeodactylum tricornutum. Proc Natl Acad Sci USA 116:17316–17322. https://doi.org/10.1073/pnas.1906726116
Li G, Woroch AD, Donaher NA et al (2016) A hard day’s night: diatoms continue recycling photosystem II in the dark. Front Mar Sci 3:218. https://doi.org/10.3389/fmars.2016.00218
Mann JE, Myers J (1968) Photosynthetic enhancement in the diatom Phaeodactylum tricornutum. Plant Physiol 43:1991–1995. https://doi.org/10.1104/pp.43.12.1991
Mayfield SP, Yohn CB, Cohen A, Vihai Danon A (1995) Regulation of chloroplast gene expression. Annu Rev Plant Physiol Plant Mol Biol 46:147
McKenzie SD, Ibrahim IM, Aryal UK, Puthiyaveetil S (2020) Stoichiometry of protein complexes in plant photosynthetic membranes. Biochim Biophys Acta Bioenerg 1861:148141. https://doi.org/10.1016/j.bbabio.2019.148141
Miloslavina Y, Grouneva I, Lambrev PH et al (2009) Ultrafast fluorescence study on the location and mechanism of non-photochemical quenching in diatoms. Biochim Biophys Acta Bioenerg 1787:1189–1197. https://doi.org/10.1016/j.bbabio.2009.05.012
Minoda A, Nagasawa K, Hanaoka M et al (2005) Microarray profiling of plastid gene expression in a unicellular red alga, Cyanidioschyzon merolae. Plant Mol Biol 59:375–385. https://doi.org/10.1007/s11103-005-0182-1
Minoda A, Weber APM, Tanaka K, Miyagishima S (2010) Nucleus-independent control of the rubisco operon by the plastid-encoded transcription factor Ycf30 in the red alga Cyanidioschyzon merolae. Plant Physiol 154:1532–1540. https://doi.org/10.1104/pp.110.163188
Moustafa A, Beszteri B, Maier UG et al (2009) Genomic footprints of a cryptic plastid endosymbiosis in diatoms. Science 324:1724–1726. https://doi.org/10.1126/science.1172983
Mullet JE (1993) Dynamic regulation of chloroplast transcription. Plant Physiol 103:309–313
Mulo P, Sakurai I, Aro EM (2012) Strategies for psbA gene expression in cyanobacteria, green algae and higher plants: from transcription to PSII repair. Biochim Biophys Acta Bioenerg 1817:247–257
Murakami A, Fujita Y (1991) Regulation of photosystem stoichiometry in the photosynthetic system of the Cyanophyte Synechocystis PCC 6714 in response to light-intensity. Plant Cell Physiol 32:223
Murakami A, Fujita Y, Nemson JA, Melis A (1997) Chromatic regulation in Chlamydomonas reinhardtii: time course of photosystem stoichiometry adjustment following a shift in growth light quality. Plant Cell Physiol 38:188–193
Nagao R, Kato K, Suzuki T et al (2019) Structural basis for energy harvesting and dissipation in a diatom PSII–FCPII supercomplex. Nat Plants 5:890–901. https://doi.org/10.1038/s41477-019-0477-x
Nagao R, Kato K, Ifuku K et al (2020) Structural basis for assembly and function of a diatom photosystem I-light-harvesting supercomplex. Nat Commun 11:1–12. https://doi.org/10.1038/s41467-020-16324-3
Noordally ZB, Ishii K, Atkins KA et al (2013) Circadian control of chloroplast transcription by a nuclear-encoded timing signal. Science 339:1316–1319. https://doi.org/10.1126/science.1230397
Oka K, Ueno Y, Yokono M et al (2020) Adaptation of light-harvesting and energy-transfer processes of a diatom Phaeodactylum tricornutum to different light qualities. Photosynth Res 1:3. https://doi.org/10.1007/s11120-020-00714-1
Oxborough K, Baker NR (1997) Resolving chlorophyll a fluorescence images of photosynthetic efficiency into photochemical and non-photochemical components—calculation of qP and Fv’/Fm’ without measuring Fo’. Photosynth Res 54:135–142. https://doi.org/10.1023/A:1005936823310
Pfannschmidt T, Nilsson A, Allen JF (1999) Photosynthetic control of chloroplast gene expression. Nature 397:625–628. https://doi.org/10.1038/17624
Prihoda J, Tanaka A, De Paula WBM et al (2012) Chloroplast-mitochondria cross-talk in diatoms. J Exp Bot 63:1543–1557
Puthiyaveetil S, Allen JF (2008) Transients in chloroplast gene transcription. Biochem Biophys Res Commun 368:871–874. https://doi.org/10.1016/j.bbrc.2008.01.167
Puthiyaveetil S, Allen JF (2009) Chloroplast two-component systems: evolution of the link between photosynthesis and gene expression. Proc Biol Sci 276:2133–2145. https://doi.org/10.1098/rspb.2008.1426
Puthiyaveetil S, Kavanagh TA, Cain P et al (2008) The ancestral symbiont sensor kinase CSK links photosynthesis with gene expression in chloroplasts. Proc Natl Acad Sci USA 105:10061–10066. https://doi.org/10.1073/pnas.0803928105
Race HL, Herrmann RG, Martin W (1999) Why have organelles retained genomes? Trends Genet 15:364–370. https://doi.org/10.1016/S0168-9525(99)01766-7
Rochaix JD (2001) Posttranscriptional control of chloroplast gene expression. From RNA to photosynthetic complex. Plant Physiol 125:142–144. https://doi.org/10.1104/pp.125.1.142
Schweer J, Türkeri H, Kolpack A et al (2010) Role and regulation of plastid sigma factors and their functional interactors during chloroplast transcription—recent lessons from Arabidopsis thaliana. Eur J Cell Biol 89:940–946. https://doi.org/10.1016/j.ejcb.2010.06.016
Shimizu M, Kato H, Ogawa T et al (2010) Sigma factor phosphorylation in the photosynthetic control of photosystem stoichiometry. Proc Natl Acad Sci USA 107:10760–10764. https://doi.org/10.1073/pnas.0911692107
Stirbet A, Govindjee (2011) On the relation between the Kautsky effect (chlorophyll a fluorescence induction) and photosystem II: basics and applications of the OJIP fluorescence transient. J Photochem Photobiol B 104:236–257
Strenkert D, Schmollinger S, Gallaher SD et al (2019) Multiomics resolution of molecular events during a day in the life of Chlamydomonas. Proc Natl Acad Sci USA 116:2374–2383. https://doi.org/10.1073/pnas.1815238116
Tullberg A, Alexciev K, Pfannschmidt T, Allen JF (2000) Photosynthetic electron flow regulates transcription of the psaB gene in pea (Pisum sativum L.) chloroplasts through the redox state of the plastoquinone pool. Plant Cell Physiol 41:1045–1054. https://doi.org/10.1093/pcp/pcd031
Valle KC, Nymark M, Aamot I et al (2014) System responses to equal doses of photosynthetically usable radiation of blue, green, and red light in the marine diatom Phaeodactylum tricornutum. PLoS ONE 9:e114211. https://doi.org/10.1371/journal.pone.0114211
Wilhelm C, Büchel C, Fisahn J et al (2006) The regulation of carbon and nutrient assimilation in diatoms is significantly different from green algae. Protist 157:91–124. https://doi.org/10.1016/j.protis.2006.02.003
Wilhelm C, Jungandreas A, Jakob T, Goss R (2014) Light acclimation in diatoms: from phenomenology to mechanisms. Mar Genomics 16:5–15
Acknowledgements
We acknowledge Purdue Biochemistry Department and Center for Plant Biology for funding and Stefan Zauner, Richard Dorrell, and Steven McKenzie for discussions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kayanja, G.E., Ibrahim, I.M. & Puthiyaveetil, S. Regulation of Phaeodactylum plastid gene transcription by redox, light, and circadian signals. Photosynth Res 147, 317–328 (2021). https://doi.org/10.1007/s11120-020-00811-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11120-020-00811-1