Abstract
Salinity stress is one of the abiotic factors that greatly affect agriculture by limiting plant growth and yield worldwide. A set of 138 barley accessions from different geographical regions was characterized with the aim to identify metabolite, biochemical, and morphological phenotypes under salinity stress. Salt stress resulted in significant increases in the phytochemicals, including the contents of total phenolic (TPC), total flavonoid (TFC), proline (ProC), and total antioxidant capacity (TAC) except soluble protein (SP). Positive relationships between proline content and the secondary metabolites or antioxidants, including total phenolics and flavonoids, were detected among barley accessions, indicating a critical adaptive strategy against free radicals under salt stress. Genome-wide association study (GWAS) revealed 122 significant quantitative trait nucleotides (QTNs) associated with the measured traits which resulted in the identification of 203 potential candidate genes. Interestingly, the QTN G:A was located inside the candidate gene HORVU.MOREX.r3.3HG0291720 at position 501,703,401 on chromosome 3H. This gene encodes Ca-binding protein and contributes to the signalling pathway that in turn triggers the expression of salt-stress responsive genes. The identified QTNs/ candidate genes provide information useful for the genetic improvement of barley genotypes under salt stress.









Similar content being viewed by others
Availability of Data and Materials
All data supporting the findings of this study are available within the paper and within its supplementary materials published online.
References
Alqudah AM, Sallam A, Baenziger PS, Börner AJJoar (2020) GWAS: Fast-forwarding gene identification and characterization in temperate Cereals: lessons from Barley–a review 22:119–135
Alqudah AM, Sharma R, Börner A (2021) Insight into the genetic contribution of maximum yield potential, spikelet development and abortion in barley 3:721–736
Ashraf M, Harris PJC (2004) Potential biochemical indicators of salinity tolerance in plants. Plant Sci 166:3–16
Bates LS, Waldren RP, Teare ID (1973) Rapid determination of free proline for water-stress studies. Plant Soil 39:205–207
Bates D, Mächler M, Bolker B, Walker S (2015) Fitting linear mixed-effects models using lme4. J Stat Softw 67:1–48
Cao JF, Huang JQ, Liu X, Huang CC, Zheng ZS, Zhang XF, Shangguan XX, Wang LJ, Zhang YG, Wendel JF, Grover CE, Chen ZW (2020) Genome-wide characterization of the GRF family and their roles in response to salt stress in Gossypium. BMC Genomics 21:575
Cho K-M, Nguyen HTK, Kim SY, Shin JS, Cho DH, Hong SB, Shin JS, Ok SH (2016) CML10, a variant of calmodulin, modulates ascorbic acid synthesis 209:664–678
DeFalco TA, Marshall CB, Munro K, Kang HG, Moeder W, Ikura M, Snedden WA, Yoshioka K (2016) Multiple calmodulin-binding sites positively and negatively regulate Arabidopsis CYCLIC NUCLEOTIDE-GATED CHANNEL12. Plant Cell 28:1738–1751
El Mahi H, Pérez-Hormaeche J, De Luca A, Villalta I, Espartero J, Gámez-Arjona F, Fernández JL, Bundó M, Mendoza I, Mieulet D, Lalanne E, Lee SY, Yun DJ, Guiderdoni E, Aguilar M, Leidi EO, Pardo JM, Quintero FJ (2019) A critical role of sodium flux via the plasma membrane Na(+)/H(+) exchanger SOS1 in the salt tolerance of rice. Plant Physiol 180:1046–1065
Galon Y, Finkler A, Fromm H (2010) Calcium-Regulated Transcription in Plants Molecular Plant 3:653–669
Hichem H, Mounir D, Naceur EA (2009) Differential responses of two maize (Zea mays L.) varieties to salt stress: changes on polyphenols composition of foliage and oxidative damages. Ind Crops Prod 30:144–151
Hodaei M, Rahimmalek M, Arzani A, Talebi M (2018) The effect of water stress on phytochemical accumulation, bioactive compounds and expression of key genes involved in flavonoid biosynthesis in Chrysanthemum morifolium L. Ind Crops Prod 120:295–304
Isayenkov SV, Maathuis FJM (2019) Plant salinity stress: many unanswered questions remain. Front Plant Sci 10:80
Ishida H, Nishimori Y, Sugisawa M, Makino A, Mae T (1997) The large subunit of ribulose-1,5-bisphosphate carboxylase/oxygenase is fragmented into 37-kDa and 16-kDa polypeptides by active oxygen in the lysates of chloroplasts from primary leaves of wheat. Plant Cell Physiol 38:471–479
Kaewneramit T, Buaboocha T, Sangchai P, Wutipraditkul NJBP (2019) OsCaM1–1 overexpression in the transgenic rice mitigated salt-induced oxidative damage 63:335
Khosravinejad F, Heydari R, Farboodnia T (2009) Effect of salinity on organic solutes contents in barley. Pakistan Journal of Biological Sciences : PJBS 12:158–162
Kovács J (2017) Comparative study of salt stress-induced physiological and molecular responses in tomato (Solanum lycopersicum L.). szte
Kudla J, Batistic O, Hashimoto K (2010) Calcium signals: the lead currency of plant information processing. Plant Cell 22:541–563
Kumar S, Beena AS, Awana M, Singh A (2017) Physiological, biochemical, epigenetic and molecular analyses of wheat (Triticum aestivum) genotypes with contrasting salt tolerance. Front Plant Sci 8:1151
Li Z, Xie Q, Yan J, Chen J, Chen Q (2021) Genome-Wide Identification and Characterization of the Abiotic-Stress-Responsive Grf Gene Family in Diploid Woodland Strawberry (fragaria Vesca) 10:1916
Lipka AE, Tian F, Wang Q, Peiffer J, Li M, Bradbury PJ, Gore MA, Buckler ES, Zhang Z (2012) GAPIT: genome association and prediction integrated tool. Bioinformatics 28:2397–2399
Liu X, Huang M, Fan B, Buckler ES, Zhang Z (2016) Iterative usage of fixed and random effect models for powerful and efficient genome-wide association studies. PLoS Genet 12:e1005767
Liu X-J, Dong Y-H, Liu X, You C-X, Hao Y-J (2019) A C2-domain phospholipid-binding protein MdCAIP1 positively regulates salt and osmotic stress tolerance in apple. Plant Cell. Tissue and Organ Culture (PCTOC) 138:29–39
Mascher M, Wicker T, Jenkins J, Plott C, Lux T, Koh CS, Ens J, Gundlach H, Boston LB, Tulpova Z, Holden S, Hernandez-Pinzon I, Scholz U, Mayer KFX, Spannagl M, Pozniak CJ, Sharpe AG, Simkova H, Moscou MJ, Grimwood J, Schmutz J, Stein N (2021) Long-read sequence assembly: a technical evaluation in barley. Plant Cell 33:1888–1906
Milner SG, Jost M, Taketa S, Mazon ER, Himmelbach A, Oppermann M, Weise S, Knupffer H, Basterrechea M, Konig P, Schuler D, Sharma R, Pasam RK, Rutten T, Guo G, Xu D, Zhang J, Herren G, Muller T, Krattinger SG, Keller B, Jiang Y, Gonzalez MY, Zhao Y, Habekuss A, Farber S, Ordon F, Lange M, Borner A, Graner A, Reif JC, Scholz U, Mascher M, Stein N (2019) Genebank genomics highlights the diversity of a global barley collection. Nat Genet 51:319–326
Munns R (2008) Tester MJARPB. Mechanisms of Salinity Tolerance 59:651–681
Mwando E, Han Y, Angessa TT, Zhou G, Hill CB, Zhang X-Q, Li C (2020) Genome-wide association study of salinity tolerance during germination in barley (Hordeum vulgare L.) 11
Omidbakhshfard MA, Proost S, Fujikura U, Mueller-Roeber B (2015) Growth-regulating factors (grfs): a small transcription factor family with important functions in plant biology. Mol Plant 8:998–1010
Prieto P, Pineda M, Aguilar M (1999) Spectrophotometric quantitation of antioxidant capacity through the formation of a phosphomolybdenum complex: specific application to the determination of vitamin E. Anal Biochem 269:337–341
Raza SH, Athar HR, Ashraf M, Hameed A (2007) Glycinebetaine-induced modulation of antioxidant enzymes activities and ion accumulation in two wheat cultivars differing in salt tolerance. Environ Exp Bot 60:368–376
Rehman S, Abbas G, Shahid M, Saqib M, Umer Farooq AB, Hussain M, Murtaza B, Amjad M, Naeem MA, Farooq A (2019) Effect of salinity on cadmium tolerance, ionic homeostasis and oxidative stress responses in conocarpus exposed to cadmium stress: implications for phytoremediation. Ecotoxicol Environ Saf 171:146–153
Rivandi J, Miyazaki J, Hrmova M, Pallotta M, Tester M, Collins NC (2011) A SOS3 homologue maps to HvNax4, a barley locus controlling an environmentally sensitive Na+ exclusion trait. J Exp Bot 62:1201–1216
Sakuma Y, Maruyama K, Osakabe Y, Qin F, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2006) Functional analysis of an arabidopsis transcription factor, DREB2A, involved in drought-responsive gene expression. Plant Cell 18:1292–1309
Schmidt P, Hartung J, Rath J, Piepho H-P (2019) Estimating Broad-Sense Heritability with Unbalanced Data from Agricultural Cultivar Trials 59:525–536
Shahid MA, Sarkhosh A, Khan N, Balal RM, Ali S, Rossi L, Gómez C, Mattson N, Nasim W, Garcia-Sanchez F (2020) Insights into the Physiological and Biochemical Impacts of Salt Stress on Plant Growth and Development 10:938
Sharma A, Shahzad B, Rehman A, Bhardwaj R, Landi M, Zheng B (2019) Response of phenylpropanoid pathway and the role of polyphenols in plants under abiotic stress. Molecules 24
Shavrukov Y, Gupta NK, Miyazaki J, Baho MN, Chalmers KJ, Tester M, Langridge P, Collins NC (2010) HvNax3–a locus controlling shoot sodium exclusion derived from wild barley (Hordeum vulgare ssp. spontaneum). Funct Integr Genomics 10:277–291
Shen Q, Fu L, Su T, Ye L, Huang L, Kuang L, Wu L, Wu D, Chen ZH, Zhang G (2020) Calmodulin HvCaM1 negatively regulates salt tolerance via modulation of HvHKT1s and HvCAMTA4. Plant Physiol 183:1650–1662
Singh NK, Bracker CA, Hasegawa PM, Handa AK, Buckel S, Hermodson MA, Pfankoch E, Regnier FE, Bressan RA (1987) Characterization of osmotin : a thaumatin-like protein associated with osmotic adaptation in plant cells. Plant Physiol 85:529–536
Song FL, Gan RY, Zhang Y, Xiao Q, Kuang L, Li HB (2010) Total phenolic contents and antioxidant capacities of selected chinese medicinal plants. Int J Mol Sci 11:2362–2372
Sun Y, Zhao J-Y, Li Y-T, Zhang P-G, Wang S-P, Guo J, Chen J, Zhou Y-B, Chen M, Ma Y-Z, Fang Z-W, Xu Z-S (2021) Genome-wide analysis of the C2 domain family in soybean and identification of a putative abiotic stress response gene GmC2–148 12
Tester M, Davenport RJAob (2003) Na+ tolerance and Na+ transport in higher plants 91:503–527
Thabet SG, Alqudah AM (2019) Crops and Drought. eLS, pp 1–8
Thabet SG, Alqudah AM (2023) New genetic insights into improving barley cope with salt stress via regulating mineral accumulation, cellular ion homeostasis, and membrane trafficking. Environ Exp Bot 208:105252. https://doi.org/10.1016/j.envexpbot.2023.105252
Thabet SG, Moursi YS, Karam MA, Graner A, Alqudah AM (2018) Genetic basis of drought tolerance during seed germination in barley. PLoS ONE 13:e0206682
Thabet SG, Moursi YS, Karam MA, Börner A, Alqudah AM (2020) Natural Variation Uncovers Candidate Genes for Barley Spikelet Number and Grain Yield under Drought Stress 11:533
Thabet SG, Alomari DZ, Alqudah AM (2021a) Exploring natural diversity reveals alleles to enhance antioxidant system in barley under salt stress. Plant Physiol Biochem 166:789–798
Thabet SG, Moursi YS, Sallam A, Karam MA, Alqudah AM (2021b) Genetic associations uncover candidate SNP markers and genes associated with salt tolerance during seedling developmental phase in barley. Environ Exp Bot 188:104499
Thabet SG, Alomari DZ, Börner A, Brinch-Pedersen H, Alqudah AM (2022) Elucidating the genetic architecture controlling antioxidant status and ionic balance in barley under salt stress. Plant molecular biology
Tohidi B, Rahimmalek M, Arzani A (2017) Essential oil composition, total phenolic, flavonoid contents, and antioxidant activity of Thymus species collected from different regions of Iran. Food Chem 220:153–161
VanRaden PMJJods (2008) Efficient methods to compute genomic predictions 91:4414–4423
Wei T, Simko V (2017) R package “corrplot”: visualization of a Correlation Matrix (Version 0.84). Vienna
Yokotani N, Ichikawa T, Kondou Y, Maeda S, Iwabuchi M, Mori M, Hirochika H, Matsui M, Oda K (2009) Overexpression of a rice gene encoding a small C2 domain protein OsSMCP1 increases tolerance to abiotic and biotic stresses in transgenic Arabidopsis. Plant Mol Biol 71:391–402
Yoo JH, Park CY, Kim JC, Heo WD, Cheong MS, Park HC, Kim MC, Moon BC, Choi MS, Kang YH, Lee JH, Kim HS, Lee SM, Yoon HW, Lim CO, Yun DJ, Lee SY, Chung WS, Cho MJ (2005) Direct interaction of a divergent CaM isoform and the transcription factor, MYB2, enhances salt tolerance in arabidopsis. J Biol Chem 280:3697–3706
Zhang H, Zeng Y, Seo J, Kim YJ, Kim ST, Kwon SW (2022) Global identification and characterization of C2 domain-containing proteins associated with abiotic stress response in rice (Oryza sativa L.). Int J Mol Sci 23
Zou Y, Lu Y, Wei D (2004) Antioxidant activity of a flavonoid-rich extract of Hypericum perforatum L. in vitro. J Agric Food Chem 52:5032–5039
Acknowledgements
The author (M.D.A) would like to acknowledge Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2023R355), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
Funding
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
SGT and AMA designed the experiment and analyzed the data. SGT, M.D.A, A.A.J, and AMA wrote and edited the manuscript. SGT conceived the idea and participated in the interpretation of the results.
Corresponding authors
Ethics declarations
Ethical Approval
This article does not contain any studies involving animals or human participants performed by any of the authors.
Competing interests
The authors have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Highlights
• This study focuses on characterizing a metabolite-based genome-wide association study (mGWAS) that is involved in salt adaptation in barley.
• The identified QTN/candidate genes provide information useful for the genetic improvement of barley genotypes under salt stress.
• However, the genetic control of metabolomes underlying crop environmental stress adaptation remains elusive.
Supplementary Information
Below is the link to the electronic supplementary material.
11105_2023_1408_MOESM1_ESM.pptx
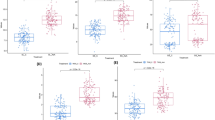
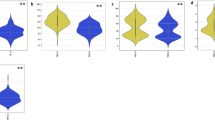
Fig. S1: The number of SNPs within 1 MB window size for barley; Fig. S2: Distribution of accessions for antioxidants under control and salt stress in barley. Proline content (ProC), Soluble protein (SP), Total antioxidant capacity (TAC), Total phenolic content (TPC), and Total flavonoid content (TFC); Fig. S3: Phenotypic distribution of accessions for all agronomic traits under control and salt stress in barley. Spike Length (SL), spikelet per spike (SS), grains per spike (GS), Weight of grains per spike (WGS), and Thousand kernel weight (TKW); Fig. S4: Phenotypic distribution of accessions for all of the studied traits underlie salt-tolerant indices (STI) in barley; Fig. S5: Comparison association analysis for FarmCPU, GLM, MLM models indicates Manhattan plot for; ProC—Proline content, SP—Soluble protein, TAC—Total antioxidant capacity, TFC— Total Flavonoid content, and TPC— Total phenolic content, SL— Spike Length, SS—Spikelets per Spike, GS — Grains per Spike, WGS—Weight Grains per Spike, and TKW—Thousand Kernel Weight in barley. The x-axis shows the chromosomes and the SNP order. The y-axis shows the − Log10 (P-value) for each SNP marker. The x-axis shows the chromosomes and the SNP order. The y-axis shows the − Log10 (P-value) for each SNP marker; Fig. S6: Quantile–quantile scale representing expected versus observed -log10 (p-value) underlying FarmCPU, GLM, MLM models for traits, including ProC—Proline content, SP—Soluble protein, TAC—Total antioxidant capacity, TFC— Total Flavonoid content, and TPC— Total phenolic content, SL— Spike Length, SS—Spikelets per Spike, GS — Grains per Spike, WGS—Weight Grains per Spike, and TKW—Thousand Kernel Weight in barley (PPTX 17788 KB)
11105_2023_1408_MOESM2_ESM.xlsx
Table S1: The detailed information of barley accessions; Table S2: Heritability and analysis of variance for all biochemical traits; Table S3: Heritability and analysis of variance for all morphological traits; Table S4: Marker trait association with the studied traits. Physical position of markers which are passing -log10(4) and R2 10%; Table S5: Marker trait association with the multi traits. Physical position of markers which are passing -log10(4) and R2 10%; Table S6: The list of candidate genes based on the linkage disequilibrium of multi-traits associated marker. (XLSX 54 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Thabet, S.G., Alqahtani, M.D., Jabbour, A.A. et al. Genetic Associations Underpinning the Metabolite-Mediated Salt Stress Tolerance in Barley. Plant Mol Biol Rep (2023). https://doi.org/10.1007/s11105-023-01408-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11105-023-01408-3