Abstract
Receptor-like cytoplasmic kinases (RLCKs) form a large subfamily of proteins in plants. RLCKs are known to regulate plant immunity to bacterial and fungal pathogens. In this study, we analyzed the genome-wide complement of maize RLCK genes and conducted detailed studies on one maize RLCK. The maize genome encodes 192 RLCKs that largely mirror the RLCK family in other plants. Previous studies implicated Arabidopsis BOTRYTIS INDUCED KINASE1 (BIK1) and TOMATO PROTEIN KINASE 1b (TPK1b) in plant resistance to the bacterial pathogen Pseudomonas syringae and the fungal pathogen Botrytis cinerea. A novel maize RLCK, Zea Mays BIK1-LIKE KINASE 1 (ZmBLK1), was identified based on sequence similarity to the tomato and Arabidopsis RLCKs. We demonstrated that ZmBLK1 displays protein kinase activity in vitro and the protein localizes to the plasma membrane. Importantly, expression of ZmBLK1 partially rescued the growth and disease phenotypes of the Arabidopsis bik1 mutant plants. The expression of ZmBLK1 was induced in maize at 12 h after inoculation with Clavibacter michiganensis subsp. nebraskensis (CMN), the bacterial pathogen causing Goss’s wilt. Interestingly, overexpression of ZmBLK1 in transgenic maize increased resistance to CMN but did not impact resistance to Aspergillus ear rot caused by the fungal pathogen Aspergillus flavus and the associated aflatoxin contamination. These findings support our hypothesis that ZmBLK1 contributes to plant resistance to bacterial pathogens likely by modulating events early after pathogen infection, implying that the protein may interact with other membrane proteins early in the immune response pathway.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Receptor-like kinases (RLKs) belong to a large superfamily of plant proteins, with 610 members in Arabidopsis and 1100 members in rice (Shiu et al. 2004). In many plant species, RLKs participate in various signal transduction pathways underlying development, defense against pathogens, and self-incompatibility (Walker and Zhang 1990; Stein et al. 1991; Torii 2000; Becraft 2002). Generally, RLK proteins contain a ligand-binding extracellular domain, a transmembrane region, and a C-terminal kinase domain (Shiu and Bleecker 2001a). Within the RLK superfamily, plants evolved a distinct subfamily, receptor-like cytoplasmic kinases (RLCKs), which lack the extracellular and transmembrane domains that are characteristic of the RLK family of proteins (Shiu et al. 2004). RLCKs have a serine/threonine protein kinase domain, including 11 highly conserved kinase subdomains (I to XI), a short N-terminal region, and a short C-terminal region (Hanks and Hunter 1995). Arabidopsis contains 147 RLCKs, and rice has 379 members (Vij et al. 2008). RLCKs are predicted to be localized to the plasma membrane through a post-translational N-terminal myristoylation motif. The myristic acid, which is attached to the N-terminal Gly residue, facilitates the anchoring of the proteins to membranes (Maurer-Stroh et al. 2002; Podell and Gribskov 2004). This plasma membrane localization enhances interaction between RLCKs and other membrane proteins. Frequently, RLCKs associate with RLKs to function in signaling of extracellular cues perceived by RLKs (Lin et al. 2013; Yamaguchi et al. 2013b).
RLCKs are involved in plant immune responses in several plant species, including Arabidopsis, tomato, and rice (Veronese 2006; AbuQamar et al. 2008; Vij et al. 2008; Ao et al. 2014). A well-established model in Arabidopsis describes the interaction between the RLCK, BIK1, and two leucine-rich repeat-receptor kinases, FLS2 and BAK1 (Chinchilla et al. 2007; Lu et al. 2010). The interaction between BIK1, FLS2, and BAK1 happens during pathogen- or microbe-associated molecular pattern (PAMPs/MAMPs)-triggered immunity (PTI) (Jones and Dangl 2006). FLS2 is the receptor for the bacterial flagellin protein, a common bacterial PAMP. Upon perception of flagellin, FLS2 and BAK1 then form a protein complex in which the activated BAK1 phosphorylates BIK1, which in turn phosphorylates the FLS2/BAK1 complex. The phosphorylated FLS2/BAK1 complex further phosphorylates BIK1. These transphosphorylation events are suggested to be required for the flagellin-induced immune signaling (Chinchilla et al. 2007; Lu et al. 2010). BIK1 also directly interacts with the pattern-recognition receptors PEPR1, EF-Tu receptor EFR, LysM receptor kinase CERK1, and with the NADPH oxidase RbohD which is required for production of reactive oxygen species (ROS) (Zhang et al. 2010; Liu et al. 2013; Li et al. 2014). These PRRs recognize bacterial and fungal elicitors of various natures (de Wit 2007; Zipfel 2008). In addition, BIK1 was found to increase Arabidopsis susceptibility to aphids through suppression of the aphid resistance and senescence-promoting Arabidopsis PAD4 (Lei et al. 2014). These studies provide evidence for a central role of BIK1 in plant innate immune responses.
In other plant species, RLCKs have been reported to have similar regulatory functions. Tomato Protein Kinase 1b (TPK1b) encodes an RLCK that is localized to the plasma membrane and required for tomato resistance against the fungal pathogen B. cinerea and the chewing insect, tobacco hornworm (AbuQamar et al. 2008). In rice, OsRLCK185 encodes an RLCK that is directly phosphorylated by a lysine motif-containing PAMP-receptor OsCERK1. Suppression of OsRLCK185 expression resulted in reduced MAP kinase activation and reduced expression of chitin-induced genes PBZ1 and PAL1 (Yamaguchi et al. 2013a; Wang et al. 2017). In addition, the OsRLCK185/ OsCERK1 interaction is suppressed by Xoo1488, an effector produced by Xanthomonas oryzae, the causal pathogen of bacterial blight disease in rice, suggesting an essential role for the OsRLCK185/OsCERK1 complex during PTI responses. Similarly, OsRLCK176/OsCERK1 interaction is required for chitin-induced ROS production (Ao et al. 2014). These previous studies indicate the significant function of RLCKs in plant immune responses. Furthermore, based on protein structure and phylogenetic analyses of RLCKs in Arabidopsis and rice, most immunity-related RLCKs belong to the RLCK VII subfamily (Shiu et al. 2004; Rao et al. 2018). Therefore, one can hypothesize that RLCK-VII members, which are highly conserved in terms of their protein structures, are also conserved in functions across dicot and monocot species.
The maize RLCK subfamily has not been classified and few studies have focused on individual maize RLCKs and their functions in disease resistance. The objectives of this study were to identify putative RLCKs in maize and to study the function of ZmBLK1 (Zm00001d034662) that was identified by a screen for maize orthologs of Arabidopsis BIK1 and tomato TPK1b. We describe features of ZmBLK1 and provide evidence for its role in disease resistance through gain of function lines that overexpress ZmBLK1.
Materials and Methods
Identification of Maize RLCKs and ZmBLK1
Maize RLCKs were identified by comparing maize kinases with Arabidopsis RLCK subfamily members in a phylogenetic analysis. All maize kinases were selected in maize B73 RefGen_v4 databases (Jiao et al. 2017) with the hmmersearch tool (https://www.ebi.ac.uk/Tools/hmmer/search/hmmsearch). The hidden Markov model (HMM) profile of eukaryotic protein kinases (PF00069) was used for the search. The 610 reported Arabidopsis RLKs were retrieved from TAIR database (Shiu and Bleecker 2001b; Berardini et al. 2015). All the maize kinases and Arabidopsis RLKs were aligned using ClustalW (Larkin et al. 2007). The kinase domains of these sequences were used to generate a phylogenetic tree by the neighbor-joining method with 500 bootstrap replicates (Saitou and Nei 1987). Maize sequences in the same cluster as known Arabidopsis RLCKs were classified into maize RLCK subfamilies. The maize BIK1-like RLCKs were identified by BLASTp analysis in the B73 RefGen_v4 translation database with the amino acid sequences of BIK1 (NP_181496.1) and TPK1b (NP_181496.1). Annotation information was retrieved from Phytozome v12.1 (https://phytozome.jgi.doe.gov/pz/portal.html). Sequence alignments were obtained with Clustal Omega (https://www.ebi.ac.uk/Tools/msa/clustalo/) and visualized with the Mega 7 tool (Sievers et al. 2011; Kumar et al. 2016). Maize gene Zm00001d034662 was assigned with the name ZmBLK1.
Nucleic Acid Purification, cDNA Synthesis, and qRT-PCR Protocols
DNA was extracted from maize leaves by the standard CTAB method (Minsavage et al. 1994). Total RNA was extracted with Trizol (Invitrogen) following the manufacturer’s protocols and then purified with an RNeasy Mini Kit (Qiagen). Total RNA was extracted from kernel samples by a phenol-chloroform method. Kernels were ground with mortar and pestle in liquid nitrogen and approximately 3 g of fine powder was mixed with 10 mL of Tris-saturated phenol (pH 4.3) and 10 mL of 1 M Tris-HCl (pH 8.0). The extracts were centrifuged at 10,000×g for 10 min at 4 °C and the supernatant was then extracted with an equal volume of chloroform/phenol (1:1), followed by an extraction with an equal volume of chloroform/isoamyl alcohol (24:1). RNA was precipitated overnight at − 20 °C and pelleted by centrifugation. The pellet was dissolved in DEPC-treated water and precipitated again with ethanol. After centrifugation, the RNA pellet was washed with 70% ethanol and dissolved in DEPC-treated water. Total RNA was then purified with an RNeasy Mini Kit (Qiagen)
cDNA was synthesized with SuperScript™ III reverse transcriptase (Invitrogen) according to standard protocols (Reese et al. 2011). For qRT-PCR, a reaction mixture containing 7.5 μl of SYBR Green Supermix (Bio-rad), 1 μl of each primer (10 μM), 2 μl of cDNA template, and 3.5 μl of nuclease-free water. The reaction cycling consisted of 3 min at 95 °C, 40 cycles of 5 s at 95 °C, and 30 s at 57 °C. The ∆∆Ct method was used to calculate relative gene expression as described (Flaherty et al. 2003) with the α-tubulin gene as the internal normalizer. Briefly, to compare the expression of ZmBLK1 in non-transgenic and transgenic plants, average Ct values of each sample were calculated from three replicates for both ZmBLK1 and α-tubulin gene (TUBA4; GRMZM2G152466). The ∆Ct values were calculated by subtracting the average Ct values of TUBA4 from those of ZmBLK1. The ∆∆Ct value of each comparison was calculated by subtracting the ∆Ct value of the non-transgenic line (control group) from ∆Ct values of each transgenic line. Similarly, the expression of ZmBLK1 in different maize tissues and different kernel developmental stages were compared using husk sample and R2 stage kernel sample, respectively, as the control group in ∆∆Ct calculations. ZmBLK1-specific primers were qZmBLK1F2/qZmBLK1R2 and the α-tubulin gene-specific primers were AlphaTUBF/AlphaTUBR (Table 1).
Construction of ZmBLK1 Vectors
The coding region of full-length ZmBLK1 was amplified by PCR with cDNA generated from RNA purified from B73 maize leaves. PCR primers were ZmBLK1F and ZmBLK1R (Table 1). The reaction product was cloned into the AscI and XmaI sites of the binary vector pTF101HA. The resulting construct (pTFZBLK1) contained a T-DNA cassette consisting of ZmBLK1 coding sequence with a triple HA tag (5′-TACCCATACGATGTTCCTGACTATGCGGGCTATCCCTATGACG TCCCGGCCTATGCAGGATCCTATCCATATGACGTTCCAGATTACGCTGCT-3′) driven by the cauliflower mosaic virus 35S promoter. The cassette also contained the bar gene as glufosinate-ammonium resistance for callus selection (Schröder et al. 1994; Rajasekaran et al. 2017). A green fluorescent protein (GFP)-tagged ZmBLK1 vector (pTFZBLK1G) was also constructed. The GFP was amplified from pCAMBIA99 (Mang et al. 2009) by PCR with primers GFPF and GFPR (Table 1). The PCR product was cloned into pTFZBLK1 and into pTF101HA replacing the HA tag in both vectors. The resulting vectors were named pTFZBLK1G and pTFGFP, respectively. All vectors were transformed into Agrobacterium tumefaciens strain GV3101 (Sheludko et al. 2006).
Transformation of Maize with ZmBLK1
Binary vector pTFZBLK1 was used to transform maize Hi-II plants by the Plant Transformation Facility at Iowa State University (Ames, IA, USA). Transformed callus tissue was sent to the University of Arkansas (Fayetteville, AR, USA), and after plantlet formation, sent to Purdue University (West Lafayette, IN, USA). The transgenic plantlets were grown in ground-beds (16 h of light daily) at the Purdue Lilly Greenhouse Facility. Transgenic plants were identified by their resistance to glufosinate-ammonium herbicide. At the fourth-leaf stage, a solution (500 mg/L) of the herbicide was applied to leaves as described by Rajasekaran et al. (2017). Expression of the ZmBLK1 was determined by Western analysis. Total proteins were isolated from herbicide-resistant plants with extraction buffer (50 mM HEPES, 50 mM EGTA, 25 mM NaF, 1 mM Na3VO4, 50 mM β-glycerophosphate, 100 mM NaCl, 1 mM PMSF, 2 mM DTT, 20% glycerol, 1× proteinase inhibitor cocktail (Sigma)) and separated by 12% SDS-PAGE. Blots were probed with an anti-HA antibody as described (Wood et al. 2006).
Out of the 4 transgenic events, two regenerated ZMBLK1 lines (ZMBLK1-1 and ZMBLK1-3) were identified. The T0 transgenic plants were crossed to B73, and the BC1 transgenic plants were backcrossed to B73. BC2 transgenic plants were used for the Goss’s wilt assay. The non-transgenic control plants (WT1-1) were generated from non-transgenic Hi-II plants by the same transformation and crossing procedures.
Location of the insertion into the maize genome was determined by Wideseq. Genomic DNA was purified from ZMBLK1-1 and ZMBLK1-3 leaf tissues by the CTAB method as described (Minsavage et al. 1994). DNA (2 μg) was digested with fragmentase (NEB), run on 2% agarose electrophoresis gels, and the 500- to 1500-bp fragments were gel-purified. The recovered fragments were ligated with a 5′ terminal adapter SeqAdapterL and a 3′ terminal adapter SeqAdapterR (Table 1). Subsequently, two rounds of PCR were conducted with the ligation products to enrich the T-DNA and genome junction region. The first PCR primer set was AP1-out/pTF-out and the second PCR primer set was AP1-nest/pTF-nest (Table 1). After the second PCR step, the products were separated by electrophoresis on 2% agarose gels, and the 500- to 1500-bp products were purified and used for Wide-seq high throughput sequencing at the Purdue Genomics Core Facility. Sequence reads that covered the T-DNA and maize genome junctions were manually identified by searching the pTFZBLK1 left border sequence against all the sequencing readings with Notepad++ software. The selected reads contained the left border sequences of pTFZBLK1 and a flanking maize genome sequence. The maize sequences were used to search the maize genome by the BLASTn analysis of the maize genome database (MaizeGDB). For the transgenic lines ZMBLK1-1 and ZMBLK1-3, 24 out of 82,146 reads and 47 out of 62,830 reads, respectively, were identified containing the T-DNA and genome junction sequence. The analysis indicated that the T-DNA cassette was inserted in the same position at chromosome 4 (36,635,505) in both transgenic lines.
Analysis of ZmBLK1 Kinase Activity
Agrobacterium tumefaciens strain GV3101 carrying pTFZBLK1 was grown at 25 °C in LB medium supplemented with 25 mg/mL rifampicin and 100 mg/mL spectinomycin. After 2 days, the cultures were centrifuged and the cells washed twice with the infiltration medium (10 mM MES, 10 mM MgCL2, and 100 μM acetosyringone). Bacterial suspensions were then maintained at room temperature for 3 h. Leaf infiltrations were applied to the abaxial surface of fully expanded Nicotiana benthamiana leaves with a 1-mL disposable syringe. Two days after incubation under constant illumination at 25 °C, infiltrated leaves were ground in liquid nitrogen and suspended in cold immunoprecipitation buffer (50 mM HEPES, 50 mM EGTA, 25 mM NaF, 1 mM Na3VO4, 50 mM β-glycerophosphate, 100 mM NaCl, 1 mM PMSF, 2 mM DTT, 20% glycerol, 1× proteinase inhibitor cocktail (Sigma-Aldrich)). Monoclonal anti-HA-agarose (Sigma) was used to immunoprecipitate the ZmBLK1-HA protein. The protein was washed three times with the immunoprecipitation buffer and twice with kinase reaction buffer (50 mM Tris, pH 7.5, 20 mM MnCl2, 2 mM EGTA, and 2 mM DTT). The protein was then tested for kinase activity in a 50-μL reaction containing 25 μg of myelin basic protein (MBP) substrate, 200 mM of ATP, and 1 μCi [γ-32P] ATP. After 30 min, the reaction mixture was boiled for 5 min and the proteins were separated by 8% SDS–PAGE. 32P-labeled products were visualized by autoradiography.
Subcellular Localization of ZmBLK1
In vivo localization of ZmBLK1 was examined in both intact tissues and protoplasts. A. tumefaciens carrying pTFPK1G was infiltrated into 3-week-old Nicotiana benthamiana plants on the 4th or 5th leaves as described (Goodin et al. 2002). Microscopic observation was carried out 2 days after infiltration. Prior (30 min) to the observation, N. benthamiana leaves that received Agrobacterium infiltration were infiltrated with 50 µM 4′,6-diamidino-2′-phenylindole, dihydrochloride (DAPI), and 1 µg/mL FM 4-64.
Protoplasts were isolated from infiltrated N. benthamiana leaf tissues as described with slight modifications (Ibrahim et al. 2012). Briefly, whole infiltrated leaves were cut into approximately 1-cm2 pieces and digested with gentle shaking in an enzyme mixture containing 2% cellulysin (Calbiochem), 0.1% pectolyase Y-23 (Seishin Pharmaceutical), and 400 mM mannitol for 90 min at 30 °C. Protoplasts were collected by centrifugation at 50×g for 2 min and then resuspended in mannitol/CaCl2 solution (400 mM mannitol and 70 mM CaCl2). Protoplast suspensions were layered onto 20% sucrose solution and centrifuged at 50×g for 10 min. Protoplasts were collected and washed twice with mannitol/CaCl2 solution, pelleted, and resuspended in 4 mL mannitol/MgCl2 solution (400 mM mannitol, 15 mM MgCl2, and 5 mM MES, pH 5.7).
Water-mounts prepared from the lower epidermis of infiltrated leaves were examined with a Nikon A1R confocal laser scanning microscope system, and protoplasts were examined with a Zeiss LSM 880 upright confocal system. For both systems, blue fluorescence of DAPI-stained nuclei was detected in the 455 nm channel. Green fluorescence of the ZmBLK1::GFP fusion protein was detected in the 488 nm channel. Red fluorescence of FM4-64-stained plasma membrane was detected in the 560 nm channel.
Complementation of Arabidopsis bik1 Plants
Agrobacterium tumefaciens strain GV3101 carrying pTFZPK1 was incubated at 28 °C for 2 days in 100 mL of LB medium with 25 mg/mL rifampicin and 100 mg/mL spectinomycin. Cultures were then diluted by 1:100 in the same medium and incubated for another 2 days. Floral-dip transformation was used to generate Arabidopsis plants expressing ZmBLK1 (Clough and Bent 1998). Briefly, A. tumefaciens cells were collected by centrifuging for 15 min at 7000×g and then resuspended in 5% sucrose solution with 0.01% Silwet L-77. Arabidopsis flower buds on the bik1 mutants were submerged into the inoculum for 30 s with gentle shaking. After transformation, plants were kept in darkness for 24 h and then grown in a 12-h photoperiod until siliques turn brown and dry. Transgenic plants are selected by spraying glufosinate-ammonium herbicide (250 mg/L) and Western blot revealing the protein expression. Leaf and stem morphology of transgenic plants was photographed with the wild-type and bik1 plants for comparison.
Alternaria brassicicola Disease Assay
Alternaria brassicicola strain MUCL20297 was grown on potato dextrose agar medium. A spore suspension with 5 × 105 spores/mL was prepared in distilled water. Detached Arabidopsis leaves were obtained from 6-week-old plants. A single 5-mL spore suspension was spotted onto the surface of each detached leaf. Inoculated leaves were placed on water-soaked filter papers and sealed in large Petri dishes to maintain high humidity. Disease lesions were measured 4 days and 7 days post-inoculation.
Analysis of Goss’s Wilt
C. michiganensis subsp. nebraskensis (CMN) was cultured on NBY (Nutrient Broth Yeast extract) agar medium at room temperature. After 4 days of growth, bacteria were harvested from the plates with sterile water, and the concentration was adjusted to OD640 = 0.3 (108 CFU/mL). Transgenic maize ZMBLK1-1 and ZMBLK1-3 and the non-transgenic line (WT1-1) were grown in 4-inch pots in the greenhouse. At the V2 stage, leaf tissue was analyzed by Western blots to verify expression of ZmBLK1. Subsequently, the third leaf of ten V4-stage plants was cut at the tip and inoculated with CMN by submerging the cut end into the inoculum for 5 s. Lesion length on each inoculated leaf was measured every 24 h. After 8 days, bacteria within the inoculated leaves were measured by a modified 6 × 6 drop plate method (Chen et al. 2003; Mbofung et al. 2015). From each leaf, a 15-cm segment, measured from the inoculation site, was collected. The leaves were flattened and scanned. The area of each leaf was analyzed with ImageJ software. The tissue was surface sterilized in 10% bleach solution for 30 s, and immediately washed three times with sterile distilled water. The tissue was placed in a sterile tube containing 15 mL of phosphate-buffered saline (PBS), placed in a sonication bath for 7 min, and then vortexed for 15 s. The resulting extract was serially diluted, and 10-µL drops of each dilution were transferred to two replicate plates of NBY agar media. The plates were incubated in the dark at room temperature, and the bacterial colonies were counted after 4 days. The number of CFU/mm2 leaf tissue was calculated.
Inoculation of Ears and Kernels with Aspergillus flavus
Ears of field-grown transgenic lines were harvested at the R3 stage. For the ear assay, three whole ears of each transgenic line were inoculated with an A. flavus conidial suspension (107 conidia/mL) by pin bars as described (King and Scott 1982). The pin bars were surface sterilized with bleach and then washed three times with sterile water before they were dipped into the A. flavus conidial suspension. The dipped pin bar was used to penetrate the kernels on the ears. The kernel screening assay was as described by Cary et al. (2011). Briefly, 30 random individual kernels from each transgenic line were removed from each transgenic line and placed in foil cups (10 kernels per cup). Each kernel was wounded with an 18-gauge needle and inoculated with 10 µL of an A. flavus conidial suspension (105 conidia/mL) at the wounding site. Inoculated ears or kernels were incubated in covered plastic boxes. Aspergillus infection and colonization on ears and kernels were photographed each day after inoculation. Five days after inoculation, inoculated kernels were removed from the ears or foil cups and kept in − 80 °C for further analysis.
Aflatoxin Analysis
Kernel samples of ear or kernel assay were ground in a coffee grinder (Hamilton Beach, Southern Pines, NC). From each sample, 0.5 g of the ground kernel was extracted by shaking overnight in 2 mL of chloroform/methanol (1:1) solution. Extracts were centrifuged and filtered through Whatman 1001055 filter papers (GE Healthcare Life Sciences, Boston, MA). Aflatoxins were analyzed by thin layer chromatography (TLC) as described by Narendrakumar and Dhandapani (2011). Briefly, 10 µL of the extracted samples was spotted on the silica gel 60 F254 plates (Merck KGaA, Darmstadt, Germany). TLC plates were developed in chloroform/acetone/water (88:12:1), photographed under UV, and analyzed with ImageJ software (https://imagej.nih.gov/ij/index.html). Quantification was obtained with a standard curve from a serial of aflatoxin standards (10 µg, 20 µg, 50 µg, and 100 µg) spotted on each TLC plate.
Results
Identification of Maize RLCKs and ZmBLK1
Of the 1512 protein kinases identified in maize, 192 clustered with the Arabidopsis RLCKs (Fig. S1). Ten of the Arabidopsis RLCK subfamilies (I, II, IV, V, VI, VII, VIII, IX, X, and XI) were identified in maize. There were no maize RLCKs that clustered with RLCK subfamily III of Arabidopsis. Also, three Arabidopsis RLCK IX members (AT3G21450, AT5G65500, and AT3G26700) failed to cluster with the other RLCK IX members in our phylogenetic analysis.
ZmBLK1 was identified as a putative maize ortholog of BIK1 and TPK1b. Amino acid sequences of BIK1 (At2g39660) and TPK1b (Solyc06g005500) were used to search the maize genome database, and five proteins shared the highest sequence identity and closest phylogenetic relationship with both BIK1 and TPK1b (Table 2; Fig. 1a). These five proteins were designated as ZmBLK1 (Zm00001d034662), ZmBLK2 (Zm00001d012958), ZmBLK3 (Zm00001d011066), ZmBLK4 (Zm00001d011779), and ZmBLK5 (Zm00001d028613). Although no previous reports about the function of these putative RLCKs were published, the Phytozome v12.1 database annotates ZmBLK1, ZmBLK2, and ZmBLK5 as cytoplasmic protein tyrosine kinases and annotates ZmBLK3 and ZmBLK4 as chloroplastic-related protein kinase APK1A. In the phylogenetic tree, these five proteins were all in the same clade and branched out from BIK1. ZmBLK1 and ZmBLK2 are two proteins which branched from TPK1b. ZmBLK1 and ZmBLK2 were therefore selected as candidates for functional analyses. An RNA-seq analysis of R3-stage maize kernels, 6 days post-inoculation with Aspergillus flavus, indicated that expression of ZmBLK1 was higher in the inoculated kernels (RPKM 33.82) and the non-inoculated control (RPKM 27.68) compared to the ZmBLK2 (RPKM 4.05 and 6.69, respectively). Thus, we hypothesized that ZmBLK1 is regulated after fungal infection similar to Arabidopsis BIK1 and tomato TPK1, suggesting a functional conservation between these RLCKs.
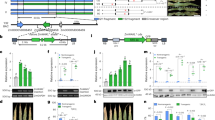
ZmBLK1 is a maize BIK1-like receptor-like protein kinase. a Rooted unweighted pair group method with arithmetic mean (UPGMA) tree of 20 top BIK1 related RLCKs in the maize B73 genome, TPK1b and BIK1. Sequences were aligned with Clustal Omega and the tree was developed with Mega 7.0. Numbers at the branches indicate the percentage of bootstrap values with 1000 replicates. Zea mays PTI1 protein was used as an outgroup. b Amino acid sequence alignment of ZmBLK1, BIK1, and TPK1b. Bars indicate the 11 kinase subdomains (I-XI). Red frame indicates the conserved protein kinase activation regions. c Kinase activity of ZmBLK1. Protein was extracted from N. benthamiana leaves transiently expressing ZmBLK1, and ZmBLK1 protein was immuno-purified. A Western blot of ZmBLK1 probed with anti-HA antibody. Autoradiographs of SDS-PAGE gels, showing B autophosphorylation of ZmBLK1 and C phosphorylation of MBP substrate. Control lanes (WT) contain protein extracts from non-treated N. benthamiana leaf tissues
ZmBLK1 Is a Functional RLCK
ZmBLK1 is a single copy gene on chromosome 1, encoding a 419 amino acid protein with an estimated molecular weight of 46.02 kDa. In the phylogenetic analysis, ZmBLK1 clustered with the Arabidopsis RLCK VII subfamily. The amino acid sequence of ZmBLK1 shows all features of an RLCK. ZmBLK1 has a protein kinase domain (residues 76 to 361) containing all 11 conserved protein kinase subdomains I to XI (Fig. 1b) (Hanks and Hunter 1995; Angermayr and Bandlow 2002). The subdomain II contains a protein kinase ATP-binding signature (residues 82 to 114). Subdomain VI contains a serine/threonine protein kinases active-site signature. In subdomain VII and VIII, ZmBLK1 has a conserved sequence between the DFG and APE motif as in BIK1 and TPK1b (Fig. 1b) (AbuQamar et al. 2008; Laluk et al. 2011). This motif has been reported as the activation segment of a protein kinase (Taylor and Radzio-Andzelm 1994; Johnson et al. 1996). To determine kinase activity of ZmBLK1, an in vitro assay was performed with transiently expressed ZmBLK1 in N. benthamiana leaf tissues. The immunopurified ZmBLK1 displayed self-phosphorylation and phosphorylation of the myelin basic protein (MBP), an artificial kinase substrate (Fig. 1c).
Expression of ZmBLK1 in various maize tissues was determined by qRT-PCR. Expression in leaves was over 7-fold higher than in silks, husks, and roots (Fig. 2a). ZmBLK1 expression was detectable in all stages of kernel development (Fig. 2b). The expression increased from the R2 stage (blister) to a maximum at the R5 stage (dent).
Relative expression of ZmBLK1 in different maize tissues. RNA was isolated from a leaves, silks, husks, and roots at the silk (R1) stage of development and b kernels at the blister (R2) milk (R3) dough (R4) dent (R5) and maturity (R6) stages of development. Expression was measured by qPCR, normalized to α-tubulin, and calculated relative to expression in a husk tissue or b blister kernel tissue. The relative expression of each gene was calculated as 2ΔΔCt. Bars indicate standard deviations of three technical replicates. The analysis was repeated on at least two biological replicates with similar results
ZmBLK1 Is Localized to the Plasma Membrane
Although no trans-membrane motif was identified in ZmBLK1, it does have an N-myristoylation motif at the N terminus. N-myristoylated proteins are often targeted to membranes for binding (Resh 1999; Maurer-Stroh et al. 2002; De Vries et al. 2006). To assess the subcellular localization of ZmBLK1, a ZmBLK1::GFP fusion protein was constructed and transiently expressed in epidermal cells of N. benthamiana leaves. The abaxial epidermal tissues were examined by confocal laser scanning microscopy. The cell shape is presented under transmission light (Fig. 3a). DAPI is a blue fluorescence marker that stains nuclei (Fig. 3b) and FM4-64 is a red fluorescence marker for plasma membranes (Fig. 3d). The green fluorescence signal from ZmBLK1::GFP was observed along the plasma membrane (Fig. 3c) and the FM4-64 signal overlapped with ZmBLK1::GFP but not with the DAPI-stained nuclei (Fig. 3e). Localization was also investigated in protoplasts isolated from infiltrated leaves of N. benthamiana. The shape of the protoplasts is presented in Fig. 3f and the green fluorescence of the ZmBLK1::GFP fusion protein (Fig. 3h) overlapped with the red fluorescence of FM4-64 (Fig. 3g, i), which demonstrated that ZmBLK1 is localized to the plasma membrane.
Subcellular localization of ZmBLK1. Epidermal cells (a–e) and protoplasts (f–i) of N. benthamiana leaves are shown. The epidermal cells and protoplasts were also imaged under transmission light channel (a, f). The green fluorescence indicated the transiently expressed ZmBLK1-GFP fusion protein (c, h). The nuclei were stained with DAPI (b). The plasma membrane was stained with FM4-64 (d, g). The GFP fluorescence merged with FM4-64 signal indicated the plasma membrane localization of ZmBLK1 (e, i)
ZmBLK1 Rescues the Disease Response and Growth Phenotypes of Arabidopsis bik1 Mutant
The Arabidopsis bik1 mutant plants lose resistance to the fungal pathogen Alternaria brassicicola and display altered growth traits, including early flowering and weak stem strength leading to lodging of plants (Veronese 2006). In this study, we examined whether ZmBLK1 can functionally substitute for BIK1 through the transgenic expression of a 35S:ZmBLK1 construct in Arabidopsis bik1 plants. The resulting Arabidopsis bik1;35S:ZmBLK1 lines displayed reduced susceptibility to A. brassicicola (Fig. 4b, c). After inoculation of detached leaves, the size of disease lesions on wild-type and bik1:ZmBLK1 leaves was limited primarily to the site of inoculation. Compared to wild type and ZmBLK1:bik1, the lesions on bik1 leaves were significantly larger and chlorosis was evident on the margin of the lesions. This recovery of resistance to A. brassicicola indicated that the BIK1 and ZmBLK1 share similar functions during plant immune responses.
Expression of ZmBLK1 rescues the increased susceptibility of Arabidopsis bik1 mutant to A. brassicicola. a PCR and Western blot showing the expression of ZmBLK1 in wild type, bik1 and bik1;ZmBLK1 transgenic plants. b, c bik1;ZmBLK1 transgenic plants showed wild-type level of resistance to A. brassicicola at 7 days post-inoculation. Experiments were repeated twice with similar results. In c, capitalized letters indicate significant groups and sample size n = 24. WT: wild-type Arabidopsis line Col-0
In addition, in bik1;35S:ZmBLK1 lines, the leaf margins were less serrated and the surface of leaves was less wrinkled compared to bik1 plants, but they were not complimented fully to the wild-type phenotype (Fig. 5a). Stem strength of bik1;ZmBLK1 plants was similar to bik1 plants, resulting in a similar lodging phenotype (Fig. 5b). The siliques produced by bik1;ZmBLK1 plants were similar to those of wild-type plants, while the siliques of bik1 plants were smaller (Fig. 5c). Under the same 16:8 photoperiod condition, bik1 plants flowered 10 to 12 days earlier than wild-type plants, whereas ZmBLK1 plants flowered 0 to 2 days earlier than wild-type plants. These results indicate that ZmBLK1 can partially complement the developmental functions of BIK1 in Arabidopsis.
ZmBLK1 partially rescues some but not all of the altered growth traits in Arabidopsis bik1 mutant plants. a Leaves of bik1;ZmBLK1 plants showing less serrated margins and less wrinkled surfaces than bik1 plants. b Stems on bik1;ZmBLK1-expressing plants showing lodging due to weaker stem strength similar to bik1 plants. c Inflorescences of bik1;ZmBLK1 plants showing siliques with similar size to the wild-type plants. WT: wild-type Arabidopsis line Col-0
Overexpression of ZmBLK1 Increases Resistance to Goss’s Wilt
Expression of ZmBLK1 was studied through qRT-PCR, which indicated that ZmBLK1 expression was up to 50 times higher in the non-transgenic plants before inoculation (Fig. 6a). When the leaves on Hi-II×B73 hybrid maize were inoculated with the Goss’s wilt pathogen C. michiganensis subsp. nebraskensis (CMN), a disease lesion formed at the inoculation site and spread to adjacent areas. Two days post-inoculation with CMN, symptoms (necrosis and water soaking) were visible at the inoculation site on the two transgenics ZMBLK1-1 and ZMBLK1-3 and the non-transformed plants. Over the next 7 days, disease lesion development on the ZMBLK1-1 and ZMBLK1-3 plants was significantly slower (P < 0.05) than the non-transgenic plants (Fig. 6b). The rate of lesion spread on the non-transgenic line was 23.96 mm/day (± 1.18) and 9.48 mm/day (± 1.37) and 10.99 mm/day (± 1.26) on ZMBLK1-1 and ZMBLK1-3, respectively (Fig. 6b). At 8 days post inoculation (DPI), the inoculated leaves from the transgenic plants developed distinctively smaller disease lesions compared to those from the non-transgenic plants (Fig. 6c). Although the number of bacteria in the inoculated leaves varied greatly, significantly fewer bacteria were observed in the transgenic lines (ZMBLK1-1, P = 0.0026; ZMBLK1-3, P < 0.001) than the non-transgenic line. Mean bacterial counts from the infected leaves of ZMBLK1-1 and ZMBLK1-3 were 3.5 × 104 CFU/mm2 of leaf (± 3.4 × 105) and 3.8 × 104 CFU/mm2 of leaf (± 3.8 × 104), respectively, which were lower than from the non-transgenic plants 4.9 × 106 CFU/mm2 of leaf (± 3.9 × 105). These results indicate that overexpression of ZmBLK1 provides resistance to Goss’s wilt in maize plants by restricting disease symptoms and bacterial growth.
Transgenic maize expressing ZmBLK1 show reduced disease lesion size after inoculation with Clavibacter michiganensis subsp. nebraskensis (CMN). a ZmBLK1 expression is higher in the ZMBLK1-1 and ZMBLK1-3 plants than the non-transgenic (WT1-1) plants. b Disease development of Goss’s wilt lesion on non-transgenic (WT-1), ZMBLK1-1 and ZMBLK1-3 plants. Disease lesion length was measured from the inoculation point. Data are the mean from 10 plants and the bars represent the standard error. In a, expression was measured by qRT-PCR. Data were normalized to α-tubulin, and expression was calculated relative to expression in the non-transgenic plant (WT1-1). The relative expression of ZmBLK1 was calculated as 2ΔΔCt. Bars indicate standard deviations of three technical replicates. c Disease lesions on non-transgenic (WT1-1), ZMBLK1-1, and ZMBLK1-3 plants at 8 DPI. ZMBLK1-1 and ZMBLK1-3 showed restricted disease lesion phenotype compared to the non-transgenic control
Overexpression of ZmPK1 Does Not Affect Resistance to Aspergillus Ear Rot or Aflatoxin Contamination
When maize kernels were inoculated with A. flavus conidia, disease symptoms were observed at the inoculation site after 2 days. Over a 5-day incubation period, no differences in disease were observed between the transgenic lines overexpressing ZmBLK1 and the kernels of non-transgenic plants (Fig. S2). Also, no difference in aflatoxin accumulation between transgenic and non-transgenic kernels was found after 5 days post-inoculation (Table S1).
Discussion
We compared 1512 protein kinases in maize B73 with the 610 reported Arabidopsis RLKs and identified a large number (195) of putative maize RLCKs. These maize RLCKs also clustered with ten of the 11 Arabidopsis subfamilies. Subfamily III cluster, which is similar to the rice RLCKs, was not identified in maize (Shiu et al. 2004). The Arabidopsis RLCK subfamily III contains ZRK3 (AT3G57720), which is required for recognition of the Pseudomonas syringae type III effector HopF2a (Seto et al. 2017) and RKS1 (AT3G57710), which is associated with broad-spectrum resistance to Xanthomonas campestris (Huard-Chauveau et al. 2013).
In Arabidopsis, 46 RLCKs, including BIK1, have been classified into the RLCK-VII subfamily (Shiu et al. 2004; Lin et al. 2013). As in Arabidopsis, RLCK VII is also the largest RLCK subfamily in maize. In our phylogenetic analysis, which included BIK1 and TPK1b, none of the 20 top maize RLCK candidates tightly clustered with BIK1, while ZmBLK1 and ZmBLK2 were most closely associated with TPK1b. ZmBLK1 and ZmBLK2 are also the most closely related to the rice RLCK, OsRLCK118 (83% identity with ZmBLK1 and 82% with ZmBLK2), which is necessary for Xanthomonas leaf blight resistance (Zhou et al. 2016).
In rice, RLCK VII subfamily proteins have overlapping functions (Shiu et al. 2004; Zhou et al. 2016). Both ZmBLK1 and ZmBLK2 are predicted to belong in the RLCK-VII family, and we speculate that they have roles in regulating disease resistance in maize. ZmBLK1 and ZmBLK2, which are on chromosome 1 and 5, respectively, share 90.74% identity in amino acid sequence. Thus, ZmBLK1 and ZmBLK2 may share some functional redundancy.
In our study, ZmBLK1 exhibited kinase activities in vitro. Most protein kinases in the eukaryotic kingdoms have 11 conserved kinase catalytic subdomains (Hanks et al. 1988; Hanks and Hunter 1995). The kinase activation region (from DGP through APE) in subdomain VII and VIII is required as a structural component for the catalytic ability (Johnson et al. 1996). In ZmBLK1, two threonine residues Thr-246 and Thr-251, which are conserved among ZmBLK1, BIK1, and TPK1b, are in the kinase activation region. These two threonine residues were identified to have essential roles for kinase activity in BIK1 and TPK1b. Substitutions of any of these two residues resulted in loss of kinase activity and resistance to B. cinerea (AbuQamar et al. 2008; Laluk et al. 2011). These results suggest that Thr-246 and Thr-251 are phosphorylatable residues in ZmBLK1.
We confirmed that ZmBLK1 is localized to the plasma membrane and the N-terminal myristate residue likely facilitates localization as described by Resh (1999). Like BIK1, the localization of ZmBLK1 suggests that the enzyme may directly interact with other RLKs or RLCKs. Song et al. (2015) performed bioinformatics analysis of the entire maize genome and classified a series of maize immune-related LRR-containing kinases, including a maize homolog of FLS2, which in Arabidopsis recognizes flagellin and activates signaling cascades by binding to BIK1 (Gómez-Gómez and Boller 2000; Zhang et al. 2010). Any signaling cascade linked to ZmBLK1 remains unknown.
BIK1 has been implicated in plant development and immune response functions. The bik1 mutant has wrinkled leaf surfaces with serrated leaf margins (Veronese 2006). During the reproductive stages, stems of bik1 mutant are weak and small siliques are produced. It is unknown whether ZmBLK1 is involved in plant development. We found that overexpression of ZmBLK1 in maize did not reveal altered plant morphology or development. A maize line (UFMu-00071), containing a Mu insertion in the 5′-UTR of ZmBLK1, has been identified (McCarty and Meeley 2009). However, this line was not available for our study. When ZmBLK1 was expressed in Arabidopsis bik1 mutant plants, we found that several growth phenotypes of bik1 were rescued. The wrinkled leaf surfaces and serrated leaf margins were partially recovered in the ZmBLK1-expressing plants. The lodging phenotype was not recovered in the ZmBLK1-expressing plants, but the flowering time and siliques morphology were restored to wild-type level. In addition, bik1 plants lost resistance to both B. cinerea and A. brassicicola (Veronese 2006). We found that the expression of ZmBLK1 rescued the A. brassicicola susceptibility of bik1 plants to the wild-type level. These results indicate that ZmBLK1 may play some roles in regulating plant development and disease resistance in maize plant. Phylogenetically, five ZmBLKs (ZmBLK1 to ZmBLK5) are closely related to BIK1. These five kinases may have overlapping functions but also divisions in their functions.
We found that the expression of ZmBLK1 increased in whole kernel tissues during maize seed development. The results are consistent with expression profile data available in the MaizeGDB database (Sekhon et al. 2011). In a microarray analysis carried out by Liu et al. (2008), genes encoding starch metabolic enzymes and storage proteins were most commonly upregulated during seed development, but expression patterns of protein kinases were variable. For example, some cyclin-dependent kinases involved in cell division were highly expressed after pollination but downregulated after the blister stage (Liu et al. 2008). Some putative protein kinases involved in abscisic acid signaling were upregulated during seed development (Liu et al. 2008). There have been no previous studies that examine the role of RLCKs in seed development. However, ZmBLK1 may be involved in plant hormone pathways that regulate seed development, and its expression level may change along with changes in hormone levels in seeds.
Genetic data reveal that BIK1 is a regulator of resistance to bacterial and fungal pathogens (Veronese 2006; Laluk et al. 2011). Knockout of BIK1 resulted in decreased resistance to B. cinerea and type III-secretion mutants of P. syringae pv tomato. (Lu et al. 2010). In tomato, TPK1b RNAi plants showed no impact on resistance to P. syringae (AbuQamar et al. 2008). Veronese (2006) showed that BIK1 expression in Arabidopsis leaves increases about 6.5-fold after inoculation with B. cinerea, and expression remains high for at least 72 h post-inoculation. AbuQamar et al. (2008) reported that TPK1b is induced within 12 h in tomato leaves inoculated with B. cinerea or P. syringae. In maize, we observed an increase in ZmBLK1 expression 12 h after inoculation with CMN, and the expression decreased to the basal level by 72 h. Our finding that the ZmBLK1-overexpressing lines provided resistance to Goss’s wilt suggests that ZmBLK1 is involved in the bacterial resistance pathways. Although BIK1 and TPK1b were required for resistance to the fungal pathogen B. cinerea, respectively, in Arabidopsis and tomato plants (Veronese 2006; AbuQamar et al. 2008), our ZmBLK1-overexpressing maize lines did not display any inhibition of Aspergillus ear rot and aflatoxin accumulation in kernels. This result indicates different roles of ZmBLK1 during Goss’s wilt and Aspergillus ear rot disease development. Clearly, kernels contain highly specialized tissues that have different developmental and transcriptional patterns compared to leaves and other vegetative tissues (Lopes and Larkins 2007). Therefore, the immune responses to A. flavus in kernels may differ from the responses to CMN in leaves.
Conclusions
This report classifies the receptor-like cytoplasmic kinase protein family in maize, including 195 members that were further classified into ten subfamilies. One maize RLCK, ZmBLK1, is the putative maize ortholog of tomato TPK1b. The protein is localized to the plasma membrane and exhibits kinase activity in vitro. Expression of ZmBLK1 complemented the early flowering and small silique phenotypes in bik1 Arabidopsis to wild-type levels and partially restored leaf morphology. Overexpression of ZmBLK1 in maize increased resistance to Goss’s wilt but had no impact on Aspergillus ear rot and aflatoxin accumulation. We propose that ZmBLK1 may regulate the signal transduction in maize immune responses. Further studies are needed to determine how ZmBLK1 and potential interacting receptor-like cytoplasmic kinases function in the signaling pathways.
Availability of Data and Material
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
AbuQamar S, Chai M-F, Luo H, Song F, Mengiste T (2008) Tomato protein kinase 1b mediates signaling of plant responses to necrotrophic fungi and insect herbivory. Plant Cell Online 20:1964–1983
Angermayr M, Bandlow W (2002) RIO1, an extraordinary novel protein kinase. FEBS Lett 524:31–36
Ao Y, Li Z, Feng D, Xiong F, Liu J, Li JF, Wang M et al (2014) OsCERK1 and OsRLCK176 play important roles in peptidoglycan and chitin signaling in rice innate immunity. Plant J 80:1072–1084
Becraft PW (2002) Receptor kinase signaling in plant development. Annu Rev Cell Dev Biol 18:163–192
Berardini TZ, Reiser L, Li D, Mezheritsky Y, Muller R, Strait E, Huala E (2015) The Arabidopsis information resource: making and mining the “gold standard” annotated reference plant genome. Genesis 53:474–485
Cary JW, Brown RL, Luo M, Bhatnagar D, Chen ZY, Rajasekaran K (2011) Developing resistance to aflatoxin in maize and cottonseed. Toxins (Basel) 3:678–696
Chen CY, Nace GW, Irwin PL (2003) A 6x6 drop plate method for simultaneous colony counting and MPN enumeration of Campylobacter jejuni, Listeria monocytogenes, and Escherichia coli. J Microbiol Methods 55:475–479
Chinchilla D, Zipfel C, Robatzek S, Kemmerling B, Nürnberger T, Jones JDG, Felix G et al (2007) A flagellin-induced complex of the receptor FLS2 and BAK1 initiates plant defence. Nature 448:497–500
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
De Vries JS, Andriotis VME, Wu AJ, Rathjen JP (2006) Tomato Pto encodes a functional N-myristoylation motif that is required for signal transduction in Nicotiana benthamiana. Plant J 45:31–45
de Wit Pierre JGM (2007) How plants recognize pathogens and defend themselves. Cell Mol Life Sci 64:2726–2732
Flaherty JE, Pirttilä AM, Bluhm BH, Woloshuk CP (2003) PAC1, a pH-regulatory gene from Fusarium verticillioides. Appl Environ Microbiol 69:5222–5227
Gómez-Gómez L, Boller T (2000) FLS2: An LRR receptor-like kinase involved in the perception of the bacterial elicitor flagellin in Arabidopsis. Mol Cell 5:1003–1011
Goodin MM, Dietzgen RG, Schichnes D, Ruzin S, Jackson AO (2002) pGD vectors: versatile tools for the expression of green and red fluorescent protein fusions in agroinfiltrated plant leaves. Plant J 31:375–383
Hanks SK, Hunter T (1995) The eukaryotic protein kinase superfamily: kinase (catalytic) domain structure and classification. FASEB J 9:576–96
Hanks SK, Quinn AM, Hunter T (1988) The kinase family: conserved protein phylogeny features and deduced domains of the catalytic. Science 241:42–52
Huard-Chauveau C, Perchepied L, Debieu M, Rivas S, Kroj T, Kars I, Bergelson J et al (2013) An atypical kinase under balancing selection confers broad-spectrum disease resistance in Arabidopsis. PLoS Genet 9:e1003766
Ibrahim A, Hutchens HM, Howard Berg R, Sue Loesch-Fries L (2012) Alfalfa mosaic virus replicase proteins, P1 and P2, localize to the tonoplast in the presence of virus RNA. Virology 433:449–461
Jiao Y, Peluso P, Shi J, Liang T, Stitzer MC, Wang B, Campbell MS et al (2017) Improved maize reference genome with single-molecule technologies. Nature 546:524–527
Johnson LN, Noble MEM, Owen DJ (1996) Active and inactive protein kinases: structural basis for regulation. Cell 85:149–158
Jones JDG, Dangl JL (2006) The plant immune system. Nature 444:323–329
King SB, Scott GE (1982) Field inoculation techniques to evaluate maize for reaction to kernel infection by Aspergillus flavus. Phytopathology 72:782
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Laluk K, Luo H, Chai M, Dhawan R, Lai Z, Mengiste T (2011) Biochemical and genetic requirements for function of the immune response regulator BOTRYTIS-INDUCED KINASE1 in plant growth, ethylene signaling, and PAMP-triggered immunity in Arabidopsis. Plant Cell 23:2831–2849
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F et al (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Lei J, Finlayson AS, Salzman RA, Shan L, Zhu-Salzman K (2014) BOTRYTIS-INDUCED KINASE1 modulates Arabidopsis resistance to Green Peach Aphids via PHYTOALEXIN DEFICIENT4. Plant Physiol 165:1657–1670
Li L, Li M, Yu L, Zhou Z, Liang X, Liu Z, Cai G et al (2014) The FLS2-associated kinase BIK1 directly phosphorylates the NADPH oxidase RbohD to control plant immunity. Cell Host Microbe 15:329–338
Lin W, Ma X, Shan L, He P (2013) Big roles of small kinases: the complex functions of receptor-like cytoplasmic kinases in plant immunity and development. J Integr Plant Biol 55:1188–1197
Liu X, Fu J, Gu D, Liu W, Liu T, Peng Y, Wang J, Wang G (2008) Genome-wide analysis of gene expression profiles during the kernel development of maize (Zea mays L.). Genomics 91:378–387
Liu Z, Wu Y, Yang F, Zhang Y, Chen S, Xie Q, Tian X, Zhou J-M (2013) BIK1 interacts with PEPRs to mediate ethylene-induced immunity. Proc Natl Acad Sci 110:6205–6210
Lopes MA, Larkins BA (2007) Endosperm origin, development, and function. Plant Cell 5:1383
Lu D, Wu S, Gao X, Zhang Y, Shan L, He P (2010) A receptor-like cytoplasmic kinase, BIK1, associates with a flagellin receptor complex to initiate plant innate immunity. Proc Natl Acad Sci 107:496–501
Mang HG, Laluk KA, Parsons EP, Kosma DK, Cooper BR, Park HC, AbuQamar S et al (2009) The Arabidopsis RESURRECTION1 gene regulates a novel antagonistic interaction in plant defense to biotrophs and necrotrophs. Plant Physiol 151:290–305
Maurer-Stroh S, Eisenhaber B, Eisenhaber F (2002) N-terminal N-myristoylation of proteins: refinement of the sequence motif and its taxon-specific differences. J Mol Biol 317:523–540
Mbofung G, Sernett J, Horner HT, Robertson AE (2015) Comparison of susceptible and resistant maize hybrids to colonization by Clavibacter michiganensis subsp. nebraskensis. Plant Dis 100:711–717
McCarty D, Meeley R (2009) Transposon resources for forward and reverse genetics in maize. In: Ed. Bennetzen, J.L. and Hake, S. (Eds.) Handbook of Maize. Berlin, Germany: Springer pp. 561-584
Minsavage GV, Thompson CM, Hopkins DL, Leite R, Stall RE (1994) Development of a polymerase chain reaction protocol for detection of Xylella fastidiosa in plant tissue. Phytopathology 84:456–461
Narendrakumar G, Dhandapani R (2011) Characterization of aflatoxin B1 from Aspergillus species and biocompatibility studies. J Pharm Res 4:621–623
Podell S, Gribskov M (2004) Predicting N-terminal myristoylation sites in plant proteins. BMC Genomics 5:1–15
Rajasekaran K, Majumdar R, Sickler C, Wei Q, Cary J, Bhatnagar D (2017) Fidelity of a simple Liberty leaf-painting assay to validate transgenic maize plants expressing the selectable marker gene, bar. J Crop Improv 31:628–636
Rao S, Zhou Z, Miao P, Bi G, Hu M, Wu Y, Feng F et al (2018) Roles of receptor-like cytoplasmic kinase VII members in pattern-triggered immune signaling. Plant Physiol 177:1679–1690
Reese BN, Payne GA, Nielsen DM, Woloshuk CP (2011) Gene expression profile and response to maize kernels by Aspergillus flavus. Phytopathology 101:797–804
Resh MD (1999) Fatty acylation of proteins: new insights into membrane targeting of myristylated and palmitoylated proteins. Biochim. Biophys. Acta Mol Cell Res 1451:1–16
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Schröder M, Dixelius C, Råhlén L, Glimelius K (1994) Transformation of Brassica napus by using the aadA gene as selectable marker and inheritance studies of the marker genes. Physiol Plant 92:37–46
Sekhon RS, Lin H, Childs KL, Hansey CN, Robin Buell C, De Leon N, Kaeppler SM (2011) Genome-wide atlas of transcription during maize development. Plant J 66:553–563
Seto D, Koulena N, Lo T, Menna A, Guttman DS, Desveaux D (2017) Expanded type III effector recognition by the ZAR1 NLR protein using ZED1-related kinases. Nat Plants 3:25–28
Sheludko YV, Sindarovska YR, Gerasymenko IM, Bannikova MA, Kuchuk NV (2006) Comparison of several Nicotiana species as hosts for high-scale Agrobacterium-mediated transient expression. Biotechnol Bioeng 96:608–14
Shiu S-H, Bleecker AB (2001a) Plant receptor-like kinase gene family: diversity, function, and signaling. Sci STKE 113:Re22-Re22
Shiu S-H, Bleecker AB (2001b) Receptor-like kinases from Arabidopsis form a monophyletic gene family related to animal receptor kinases. Proc. Natl Acad Sci 98:10763–10768
Shiu S-H, Karlowski W, Pan R (2004) Comparative analysis of the receptor-like kinase family in Arabidopsis and rice. Plant Cell 16:1220–1234
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R et al (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7
Song W, Wang B, Li X, Wei J, Chen L, Zhang D, Zhang W et al (2015) Identification of immune related LRR-containing genes in maize (Zea mays L.) by genome-wide sequence analysis. Int J Genomics 2015:1–11
Stein JC, Howlett B, Boyes DC, Nasrallah ME, Nasrallah JB (1991) Molecular cloning of a putative receptor protein kinase gene encoded at the self-incompatibility locus of Brassica oleracea. Proc Natl Acad Sci USA 88:8816–8820
Taylor SS, Radzio-Andzelm E (1994) Three protein kinase structures define a common motif. Structure 2:345–355
Torii KU (2000) Receptor kinase activation and signal transduction in plants: an emerging picture. Curr Opin Plant Biol 3:361–367
Veronese P (2006) The membrane-anchored BOTRYTIS-INDUCED KINASE1 plays distinct roles in Arabidopsis resistance to necrotrophic and biotrophic pathogens. Plant Cell Online 18:257–273
Vij S, Giri J, Dansana PK, Kapoor S, Tyagi AK (2008) The receptor-like cytoplasmic kinase (OsRLCK) gene family in rice: organization, phylogenetic relationship, and expression during development and stress. Mol Plant 1:732–750
Walker JC, Zhang R (1990) Relationship of a putative receptor protein kinase from maize to the S-locus glycoproteins of Brassica. Nature 345:743–746
Wang C, Wang G, Zhang C, Zhu P, Dai H, Yu N, He Z et al (2017) OsCERK1-Mediated chitin perception and immune signaling requires receptor-like cytoplasmic kinase 185 to activate an MAPK cascade in rice. Mol Plant 10:619–633
Wood CC, Robertson M, Tanner G, Peacock WJ, Dennis ES, Helliwell CA (2006) The Arabidopsis thaliana vernalization response requires a polycomb-like protein complex that also includes VERNALIZATION INSENSITIVE 3. Proc Natl Acad Sci 103:14631–14636
Yamaguchi K, Yamada K, Ishikawa K, Yoshimura S, Hayashi N, Uchihashi K, Ishihama N et al (2013a) A receptor-like cytoplasmic kinase targeted by a plant pathogen effector is directly phosphorylated by the chitin receptor and mediates rice immunity. Cell Host Microbe 13:347–357
Yamaguchi K, Yamada K, Kawasaki T (2013b) Receptor-like cytoplasmic kinases are pivotal components in pattern recognition receptor-mediated signaling in plant immunity. Plant Signal Behav 8:e25662
Zipfel C (2008) Pattern-recognition receptors in plant innate immunity. Curr Opin Immunol 20:10–16
Zhang J, Li W, Xiang T, Liu Z, Laluk K, Ding X, Zou Y et al (2010) Receptor-like cytoplasmic kinases integrate signaling from multiple plant immune receptors and are targeted by a Pseudomonas syringae effector. Cell Host Microbe 7:290–301
Zhou X, Wang J, Peng C, Zhu X, Yin J, Li W, He M et al (2016) Four receptor-like cytoplasmic kinases regulate development and immunity in rice. Plant Cell Environ 39:1381–1392
Acknowledgements
We gratefully thank Braham Deep Singh Dhillon, Sujoung Shim, Siming Xu, Xinyan Zhang, and Longfei Wang for their technical assistance.
Funding
This study was funded by the Agriculture and Food Research Initiative Competitive Grants Program of the US Department of Agriculture National Institute of Food and Agriculture (No. 2013-68004-20359) and by the National Corn Growers Association through the Aflatoxin Mitigation Center of Excellence (AMCOE, No. 207459, 00068383 and 14055103).
Author information
Authors and Affiliations
Contributions
Charles P. Woloshuk, Tesfaye Mengiste, and Burt H. Bluhm conceived the study. Weiran Li developed and performed the experiments and analyzed the data. Chao-Jan Liao contributed significantly to the experimental design, experimental techniques, and data analysis. Weiran Li and Charles P. Woloshuk drafted the manuscript and all authors provided edits and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Key Message
• We identified 192 putative receptor-like cytoplasmic kinases (RLCK) in maize which clustered into 10 subfamilies. ZmBLK1 was identified as an ortholog of Arabidopsis BIK1 and tomato TPK1b. ZmBLK1 encodes a functional protein kinase and overexpression of ZmBLK1 in maize increased resistance to Goss’s wilt
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, W., Liao, CJ., Bluhm, B.H. et al. A Maize (Zea mays L.) BIK1-Like Receptor-Like Cytoplasmic Kinase Contributes to Disease Resistance. Plant Mol Biol Rep 40, 28–42 (2022). https://doi.org/10.1007/s11105-021-01299-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-021-01299-2