Abstract
Background and aims
Recent research has recognized the presence of metal-resistant bacteria in plants and their role in phytoremediation intensification. However, information on the antibiotic resistance profile of those bacteria remains scarce. This study, describes the first isolation of endophytic bacteria from green parts of Armeria maritima growing on mine-tailing soil in southern Poland, and presents the resistance profile of these microorganisms.
Methods
Bacteria were isolated from internal tissues of Armeria maritima and characterized. Minimal Inhibitory Concentration (MIC) of metals was determined by the plate dilution method using (CH3COO)2Pb and ZnSO4 supplemented medium; antibiotic susceptibility was determined by disk diffusion method according to EUCAST version 11.0; the whole genome sequencing was performed using the MiSeq platform (Illumina). The physicochemical properties of soil were evaluated according to European Standards.
Results
Toxic metal-resistant bacteria were isolated from the green parts of Armeria maritima. The endophytes were identified as Pseudomonas spp. The annotated bacterial genomes carried genes encoding numerous metal ion transporters, metal reducing enzymes and efflux pump components. The bacteria were resistant to streptomycin, fosfomycin and ß-lactams. Moreover, genome analysis revealed the presence of MacAB-TolC efflux pump genes conferring resistance to macrolides, the multidrug efflux pumps AcrAB-TolC and MexAB-OprM.
Conclusion
Armeria maritima is inhabited by endophytic bacteria identified as Pseudomonas species that are resistant to metals and to antibiotics. Under the One Health concept the contamination of soil and plants with ARB and ARGs should be monitored and limited and a regulatory framework for safety use of bacterial bioinoculants should be established.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Careless management of agricultural and industrial activities can result in serious contamination of soils by metals (He et al. 2015; Sharma et al. 2007; Walker et al. 2003). If unchecked, this can pose significant risk to public health. Therefore, new biosafety and effective technologies intended to reduce such contamination are needed. A commonly used technology to remove metals from soil is phytoextraction: a phytoremediation method based on the application of hyperaccumulating plants that can decrease metal level in contaminated areas (Kumar et al. 1995). Hyperaccumulators are capable of sequestering extremely high levels of metals in their tissues. Although phytoextraction is an eco-friendly, low-cost method, it tends to have low efficiency because of slow growth of the plants and the low mobility and bioavailability of the metals in soil (Khan et al. 2000; Liu et al. 2020). Hence, recent years have seen growing interest in developing new phytoextraction efficiency approaches.
Recently, one promising technology for enhancing phytoextraction based on the use of plant-growth promoting bacteria (PGPB) to increase plant biomass production and tolerance to metals has been approved (Ahemad 2015; Kong and Glick 2017; Silambarasan et al. 2020). These PGPB include rhizosphere microorganisms inhabiting plant roots (PGPR) and endophytes inhabiting internal plant tissues without causing them any harm (PGPE). PGPB protect plants and promote their growth mainly by producing antibiotics, and phytohormones, and by inducing the Induced Systemic Resistance (ISR) system of the plant; they also support the dissolution of mineral nutrients, such as phosphorus or potassium, and support iron chelation (Olanrewaju et al. 2017; Gamalero and Glick 2011). PGPB can also stimulate metal uptake and bacteria resistance by various mechanisms, such as metal sorption (Kloepper et al. 1980), enzymatic reduction (Glick 2012), oxidation or extracellular precipitation via active efflux pumping (Alves et al. 2022; Bargaz et al. 2018; Kong and Glick 2017). The most commonly used PGPB species are Azospirillum, Azotobacter, Bacillus, Burkholderia, Pseudomonas or Rhizobium (Alves et al. 2022).
The use of PGPB to enhance phytoremediation may well be a common biotechnology in the near future. Various strains of PGPB have been tested. Wu et al. (2018) confirmed that the endophytic strain Buttiauxella sp. SaSR13 significantly enhanced cadmium accumulation in Sedum alfredii. Inoculation with this bacterium resulted in root elongation and, stimulated the secretion of organic acids and increased Cd uptake by S. alfredii compared to controls during a seven-day pot experiment.
Endophyte assisted phytoremediation has also been studied in Sedum alfredii by Zhang et al. (2013). The findings indicate that the tested Burkholderia sp., Sphingomonas sp., and Variovorax sp. strains significantly promoted Zn and Cd-extraction and had plant growth promoting properties. The experiment was conducted in pots for 60 days (Zhang et al. 2013). Similarly, Wang et al. (2023) revealed that inoculation of Miscanthus floridulus with an endophytic strain Bacillus cereus BL4 significantly strengthen Cd phytoremediation.
Nowadays, PGPB are commonly used in agriculture as bioinoculants. However, it is important to note that such PGPB may enhance the spread of antibiotic resistance genes (ARGs) in soil and plants because they themselves very often harbour ARGs (Chen et al. 2019; Zhang et al. 2020; Mahdi et al. 2022). Furthermore, ARGs can be located on mobile genetic elements (MGE) and they can be easily transferred among indigenous soil bacteria by horizontal gene transfer (HGT) (Arber 2014; Forsberg et al. 2012). This can represent a potential threat to public health because agricultural soil and agricultural plants act as huge reservoir and propagation hotspot of ARGs (Cadena et al. 2018; Tan et al. 2018; Forsberg et al. 2012; Zhang et al. 2015). Plants and their associated bacteria can absorb ARGs from soil and threaten human health (Zhang et al. 2011; Buchholz et al. 2011). Despite this, little research has been performed of the ARGs present in PGPB used in agriculture, and no description yet exists of the ARGs in endophytes inhabiting green parts of metallophytes.
It has been proposed that a regulatory framework is needed for new bacterial-based biofertilizers (Mahdi et al. 2022). This should include inter alia better characterization of new biofertilizers (genome mining) regarding their antibiotic resistance (AR) profile, ARG content and ARG transfer potential. Moreover, multidrug resistant strains or human pathogens should be excluded. It has also been suggested that standard criteria, regulations and quality control procedures for biofertilizer candidates should be established, so as to guarantee environmental and public health protection (Mahdi et al. 2022).
The present study describes the isolation and characterization of Armeria maritima subsp. halleri (Wallr.) Rothm. endophytes. It demonstrates that isolated Pseudomonas spp. endophytes were resistant to antibiotics and metal ions, and they harboured potential resistance genes. It also explores the possible resistance mechanisms present in the bacteria and attempts to explain the origin of the ARGs present in the isolated endophytes.
Materials and methods
Study site, sampling and soil physicochemical analysis
The studied area was located near the ZGH “Bolesław” mining and metallurgical plant in Bukowno village, in the south of Poland (50°16’40.7"N 19°28’13.8"E). ZGH “Bolesław” S.A. is a Polish company that has been operating since 1955 in Bukowno village, near Olkusz. Today, it is a modern mining and metallurgical complex, the main producer of zinc in Poland and a supplier of zinc to neighboring countries, mainly the Czech Republic, Slovakia, Austria and Hungary. In this plant zinc and lead ores are extracted and processed to produce electrolytic zinc, zinc alloys, sulfuric acid and zinc and lead concentrates.
Samples were taken during May 2015, during the flowering stage of the plants. The plant species selected for investigations was Armeria maritima subsp. halleri (Wallr.) Rothm. All collected plants were placed in polyethylene bags and transported to the laboratory in an ice cooler at 4 °C; all testing was performed within two days.
The total organic carbon, pH, calcium, magnesium, and metal content (Cr, Cu, Cd, Ni, Pb, Zn, Hg) were determined. Hg content was determined as described in DIN ISO 16,772. The other metals were tested according to the following: (ICP-OES/ICP-MS) – DIN EN ISO 11,885/DIN EN ISO 17294-2. pH was determined according to DIN EN ISO 10,390 and Total Organic Carbon (TOC) according to DIN EN ISO 15,936.
Isolation and purification of metal-tolerant bacteria
Any metal-tolerant endophytic bacteria were isolated using the Luria Bertani agar (LB) medium supplemented with filter-sterilized soluble salts of lead (CH3COO)2Pb (Pb2+) or zinc ZnSO4 (Zn2+) at a concentration of 20 mg/dm3. To isolate endophytic bacteria, the green parts of plants were separated and subjected to surface sterilization in sterile conditions under a laminar chamber (Goryluk et al. 2009). Before starting the procedure, the ends of the stem sections were secured against the inflow of sterilization agents. The first stage of sterilization was to rinse the plant fragments in 70% ethanol for about 60 s; these were then transferred to 2% mercury (II) chloride solution for 10 s and rinsed three times in distilled water. After surface sterilization, the plant material was homogenized. The obtained homogenates were diluted 10-fold and 100-fold, and 0.1 cm3 aliquots were plated on culture media. All plates were incubated at 30° C for 24–48 h. In order to determine the dry weight of the tested plants, each homogenate was poured onto a filter paper and weighed after complete drying. Based on these results, the numbers of colony forming units were then calculated per one gram of dry plant matter.
Individual bacterial colonies with different morphological characteristics were randomly selected and streaked on the LB agar medium supplemented with metal salts until pure cultures were obtained. A total of 100 bacterial isolates were selected for further studies and stored in 20% glycerol stock at -80 ºC.
Characterization of metal-tolerant bacteria
Identification
The morphological features of bacterial isolates (Gram staining) were recorded using light microscopy. Following this, biochemical analyses were performed, involved to determined oxydase and catalase activity, gelatin hydrolysis, citrate utilization, glucose fermentation and urease and fluoresceine production. All tests were prepared according to Bergey’s Manual of Systematic Bacteriology and isolates were identified to genus level (Bergey 1994).
Five out of 100 Gram-negative bacterial isolates with different morphologies were selected for further analysis. To identify the species, isolates were plated on LB agar and MALDI-TOF MS analysis was conducted by a commercial service (ALAB laboratory, Warsaw Poland). The standard Bruker interpretative criteria were applied. A score > 2.300 was used for certain species identification (Suppl. Tab. S1).
The toxic metal MIC assay
The Minimum Inhibitory Concentration (MIC) values were determined by the plate dilution method as adopted by Malik and Jaiswal (2000) with modifications. Luria Bertani LB medium supplemented with filter-sterilized soluble salts of (CH3COO)2Pb (Pb2+) and ZnSO4 (Zn2+) was used. The starting concentration for each metal was 10 mM. The inoculation was performed using 0.1 ml of bacterial suspension with a density of 106 CFU/ml. The MIC was taken as the lowest metal concentration that prevented the growth of the bacteria (Haroun et al. 2017). In this experiment E. coli 1655 strain was used as control (Spain and Alm 2003).
The antibiotic susceptibility test
Antibiotic susceptibility was determined by the disk diffusion method according to the European Committee on Antimicrobial Susceptibility Testing EUCAST version 11.0, valid from 2023-01-01. All 13 antibiotics recommended for Pseudomonas spp. were tested. Two additional antibiotics not included in EUCAST breakpoints were tested, viz. fosfomycin (50 µg) and streptomycin (25 µg), based on the presence of resistance genes detected by genome sequencing (see below). The diameter of bacterial growth inhibition zone around each of the antibiotic discs was interpreted according to the EUCAST criteria for Pseudomonas spp. If the antibiotic was not included in the standard, then a lack of any inhibition zone was interpreted as no susceptibility to the given antibiotic.
Isolation of resistance genes
The genomic DNA of the selected bacterial isolates was extracted according to Kpoda et al. (2018), and then stored at -20 °C for subsequent use. The genes coding for the efflux pump were identified using PCR amplification, while bla genes were isolated using multiplex PCR.
The efflux pump genes mexA and mex B of the Mex AB-OprM pump were amplified by PCR as described by Ugwuanyi et al. (2021). MexD, mexF and mexY genes of the MexCD-OprJ, MexEF-OprN, MexXY-OprM efflux pumps were amplified according to Poonsuk and Chuanchuen (2014). In addition, the czcA and czcR genes encoding components of the CzcCBA efflux pump were amplified using primers proposed by Perron et al. (2004). The types of ß-lactamase coding genes present were determined by multiplex PCR (Colom et al. 2003; Dallenne et al. 2010; Piotrowska et al. 2019). Four multiplex PCR assays were performed for the detection of bla genes: blaTEM, blaSHV and blaOXA genes (Multiplex I); blaCTXM genes (Multiplex II); blaVER, blaPES and blaGES genes (Multiplex III) and blaKPC, blaIMP and blaVIM genes (Multiplex IV).
All the PCR amplicons were gel-purified (Gel-Out kit, AA Biotechnology) and submitted for sequencing by a commercial service (Institute of Biochemistry and Biophysics, Polish Academy of Sciences) using ABI 3730 Genetic Analyzer, Applied Biosystems (BigDye v3.1 sequencing chemistry). Sequences of obtained gene fragments were searched against the National Center of Biotechnology Information (NCBI) using a local BLASTX program. A gene was designated as a resistanc gene if it shared at least 98% identity with other resistance gene in the database.
The whole genome sequencing and bioinformatic analysis
Genomic DNA from five selected bacteria was extracted using a Genomic Mini® kit (A&A Biotechnology) as described by the manufacturer. DNA concentration and quality were checked with the QubitTMfluorometer (Invitrogen) and bacterial genomes were sequenced by a commercial service (Institute of Biochemistry and Biophysics, Polish Academy of Sciences).
The whole genome sequencing WGS was performed on MiSeq platform (Illumina) with 300 bp paired-end reads (Supp. Tab. S4). Only high quality reads after filtering using fastp (https://github.com/OpenGene/fast) were taken for assembly step. The Unicycler version 0.4.8 assembly method was used. This Whole Genome Shotgun BioProject was deposited at DDBJ/ENA/GenBank under the accession number PRJNA886618.
Phylogenetic affiliation analysis
The genome sequences of each strain were uploaded to the Type Strain Genome Server (TYGS), i.e. a bioinformatic platform for digital, highly-reliable estimation of the relatedness of genomes based on DNA-DNA hybridization (DDH) (available at the website: http://tygs.dsmz.de) (Meier-Kolthoff et al. 2013). Additionally, a phylogenetic tree was constructed based on the RNA polymerase sigma factor RpoD (rpoD) gene (Banasiewicz et al. 2021; Girard et al. 2020). Briefly, 37 environmental-type Pseudomonas spp. strain rpoD genes were uploaded from the NCBI database (accession numbers listed in Supp. Table 2). The rpoD sequences (650 bp) were aligned using Clustal W software. The multiple sequence alignments were then used to create phylogenetic trees by the Neighbor Joining method with complete deletion of gaps, implemented in MEGA7 software (Kumar et al. 2016; Saitou and Nei 1987; Tamura et al. 2004). The evolutionary distances between sequences were computed using the Maximum Composite Likelihood method (Tamura et al. 2004), represented as the units of the number of base substitutions per site. The tree itself was drawn to scale, with branch lengths given in the same units as those of the evolutionary distances used to infer the phylogenetic tree.
Resistance genes screening
Genomes were annotated in the Rapid Annotation using Subsystem Technology (RAST) server (available at the website: http://rast.nmpdr.org) (Brettin et al. 2015). The annotation process enables the prediction of protein-coding genes, like ARG and HMRG, as well as other important elements, like direct and inverted repeats, insertion sequences, transposons and plasmids. ARGs were predicted using the Resistance Gene Identifier (RGI) application, available at the Comprehensive Antibiotic Resistance Database (CARD) (available at the website: http://card.mcmaster.ca/analyzer/rgi). In addition, Antibacterial Biocide and Metal Resistance genes Database (BacMet) was used to find resistance genes (Pal et al. 2014; available at the website: http://bacmet.biomedicine.gu.se). Finally, manual annotation was performed.
BLASTX analysis
The obtained gene fragments were searched against the National Center of Biotechnology Information (NCBI) using a local BLASTX program. A gene was designated as an ARG or MRG if it shared at least 98% identity with the best hit in the database.
Results
Physico-chemical properties of soil
At the test site, the total concentrations of Pb and Zn were 1100 mg/kg soil and 3620 mg/kg soil, respectively (Table 1). Hence, the tested soil was classified as highly contaminated (Trafas et al. 2006).
Biochemical characterization of metal tolerant bacteria
The mean total count of metal-tolerant bacteria isolated from A. maritima endosphere varied from 5.73 log CFU/g of plant material on lead supplemented medium to 5.46 log CFU/g on the zinc supplemented material. All 100 isolates selected for further studies were Gram-negative, catalase positive and glucose fermentation negative. Some differences in urease and fluoresceine production were noted between selected isolates (Table 2). According to Bergey’s Manual of Determinative Bacteriology, the isolates were identified as Pseudomonas spp. MALDI-TOF-MS analysis failed to identify the isolates down to the species level (Supp. Tab. S1).
The antibiotic susceptibility test, performed according to EUCAST, revealed different resistance profiles among selected isolates (Table 3, Supp. Tab. S3). All of the isolates were resistant to aztreonam (ATM), and meropenem (MEM) while four out of five isolates were resistant to ceftazidime (CAZ), cefepime (FEP) and streptomycin (S). Only two isolates showed resistance to imipenem (IPM). Regarding metal resistance, three isolates demonstrated a maximum MIC of 60 mM for Pb (II) (AM4, AM8, AM14) and one isolate a maximum MIC of 220 mM for Zn (II) (AM14; Table 3). The lowest MIC (30 mM) was observed for isolate Z18, for both metals tested.
Resistance determinants detection
The tested bacteria were screened for genes encoding multidrug efflux pumps known to be common in various Pseudomonas strains, such as the Resistance Nodulation Cell Division family pump genes (RND). However, none of the Mex-type pumps genes were detected and only one of two tested CzcCBA system genes were detected in any tested strain. The PCR amplicons of the czcR gene were shorter (315 bp) than the expected czcR gene (880 bp) and sequence analysis did not confirm membership of any known resistance gene.
Regarding antibiotic resistance multiplex PCR amplify any selected variants of the bla genes.
Genome characterization of endophytic Pseudomonas sp. strains
Multidrug-resistant endophytic Pseudomonas sp. strains were sequenced on Illumina platform (AM4, AM8, AM14, Z13, Z18). The draft whole genome sequence length varied from 6.1 Mb to 7.4 Mb (Table 4, Supp. Tab. S4). Mean G + C content ranged from 60 to 61%. Annotation performed using RAST server predicted between 5,700 and 7,059 coding sequences. The analysis revealed the presence of between 374 subsystems with 65 RNA genes and 395 subsystems with 63 RNA genes. The entire Genome Shotgun project was deposited at DDBJ/ENA/GenBank under the following accessions: SAMN31135831, SAMN31136163, SAMN31136268, SAMN31137609, SAMN31137624 (Table 4). The version described in this paper is the first version of the WGS project.
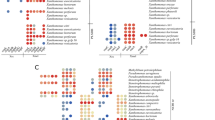
A whole-genome based taxonomic analysis based on DNA-DNA hybridization (in TYGS) found that isolate AM8 shared high homology with Pseudomonas marginalis species (dDDH above 80%; Supp. Tab. S5). In addition, the species demonstrating the greatest homology to the rest of the tested bacteria were, as follows: P. paracarnis (strain AM4, dDDH = 78.6%); P. koreensis (AM14, dDDH = 78.7%) and P. yamanorum (Z13, dDDH = 72.2%; Z18, dDDH = 70.7%) (Supp. Tab. S5). The rpoD phylogeny confirmed the species identification of three isolates, viz. AM4, Z13, and Z18; however, the species of AM8 and AM14 remain unclear (Fig. 1).
Neighbor-Joining phylogeny of rpoD partial gene sequences (650 bp), comprising type strains of 30 Pseudomonas species and 5 unknown Pseudomonas spp. (AM4, AM8, AM14, Z13, Z18). Escherichia coli ATCC11775 was used as an outgroup. The optimal tree with a sum of branch length = 1.73015111 is shown. The analysis involved 43 nucleotide sequences. Codon positions included were 1st + 2nd + 3rd + Noncoding. All positions containing gaps and missing data were eliminated. There were a total of 617 positions in the final dataset
Prediction of metal resistance genes
The annotated draft genomes confirmed the presence of genes conferring resistance to arsenic, cadmium, chromium, cobalt, copper, lead, tellurium and zinc (Table 5). All tested endophytic bacteria genomes demonstrated different ars, cus and teh genes. Moreover, all the genomes carried copC, copD, copG and cueO. The Pseudomonas sp. AM8 genome lacked the cadA, cadR and czcD genes. The czcB, czcR and czcS genes were absent from the P. paracarnis AM4 and Pseudomonas sp. AM14 genomes, while chrA and copB genes were not detected in P. yamanorum species (Z13, Z18).
The detected genes encode components of various metal resistance mechanisms (Table 6). Gene arsB encodes an inner membrane polypeptide of the ArsAB efflux pump. ArsB confer resistance to As(III) and it provides sufficient arsenic resistance, even if the bacterium lacks ArsA (Bhattacharjee and Rosen 2007). Gene arsC encodes arsenate reductase, which reduces As(V) to As(III), and gene arsR encodes transcriptional regulator of the ars operon. Moreover, the arsH gene encodes a product that strengthens bacterial resistance to arsenate and arsenite; however, its function remains unclear (Rosen 2002).
Metal-translocating P-type ATPase genes, such as cadA, were present in the genomes of Armeria maritima endophytes. The CadA protein catalyzes the active efflux of zinc, cadmium and lead ions (Rossbach et al. 2000). Additionally, genes coding transcriptional regulators were detected in our studies, like cadC and cadR. Other P-type ATPase genes conferring resistance to copper were detected, such as copB, copC, copD and copG genes, which encode periplasmic proteins that bind and/or transport Cu ions. Genes confering other copper resistance mechanisms were also observed, such as cueO encoding multicopper oxidase, and the cusR and cusS genes encoding regulatory proteins of the CusCFBA system, involved in periplasmic copper detoxification. CusS is a sensor histidine kinase while CusR is a regulatory protein (Nies 1999).
Another resistance mechanism detected in bacteria was the chemiosmotic pump protein ChrA, a membrane transporter protein responsible for the efflux of chromium out of the cell cytoplasm. It is encoded by the chromium resistance gene chrA (Branco et al. 2008).
The CzcCBA system functions as a cation-proton antiporter by transferring Cd2+, Co2+ and Zn2+ ions out from the bacterial cell (Wang et al. 2017). The czcD gene encodes cation diffusion facilitator transporter CzcD, while czcB encodes one of the CzcCBA efflux transporter system as genes czcR and czcS encode the two-component regulatory system CzcRS. Those genes form part of the Czc system, comprising the CzcCBA transporter lying across the inner and outer membrane, regulated by the CzcRS two-component system, and the CzcD cation diffusion facilitator protein (CDF).
Additionally, the cusR and cusS genes encoding regulatory proteins of the CusCFBA system involved in periplasmic copper detoxification were detected. CusS is a sensor histidine kinase while CusR is a regulatory protein (Bondarczuk and Piotrowska-Seget 2013). The proteins of the CusCFBA efflux pump, CusA and CusB, were also detected (Bondarczuk and Piotrowska-Seget 2013).
Finally, the tehA and tehB genes, which may conferr resistance to tellurium, were detected. TehA encodes an internal membrane protein, while tehB encodes the membrane associated protein (Turner et al. 1995).
Prediction of antibiotic resistance genes
Tested Pseudomonas genomes carried genes conferring different antibiotic resistance mechanisms (Table 7). All five tested genomes contained genes believed to encode components of efflux pumps, such as: acrA, acrB, macA, macB, mexT, pmpM, lysR, soxR, tolC. All sequenced genomes were found to contain genes encoding components of the antibiotic inactivation system, such as ampC, ampR, and those coding for MBL (metallo-ß-lactamase encoding gene) and the GNAT enzyme family. In addition, all tested genomes demonstrated point mutations in the gyrA and gyrB gene. The Pseudomonas yamanorum genomes lacked fosA and oprM genes, while those of the rest of the tested Pseudomonas spp. lacked oprN. The presence of genes believed to encode components of various Mex pumps varied among the tested Pseudomonas strains (Table 7). The following genes were detected: mexA, mexB, mexE, mexH, mexI and mexX.
Genes coding the components of three antibiotic resistance mechanism types, viz. antibiotic inactivation, antibiotic target alteration and antibiotic efflux pumps, were identified in the tested genomes (Table 8). Five genes responsible for antibiotic inactivation were found in the tested genomes. Metallo-ß-lactamase genes (MBL), ampC and ampR genes confer resistance to ß-lactam antibiotics. Resistance to ß-lactams is largely mediated by enzymes called ß-lactamases encoded by bla genes. These enzymes are divided into four classes (A, B, C, D) and form two groups: metallo-ß-lactamases (MBL) and serine ß-lactamases. Metallo-ß-lactamases belong to Class B. Class B is divided in three subclasses: chromosomally-encoded genes of B2 or B3 subclasses, and MGE-encoded genes of the B1 subclass (Behzadi et al. 2020). In the present study, all tested genomes contained bla genes (Table 7). This result was consistent with the antibiotic resistance profile, as the tested bacteria were resistant to ß-lactam antibiotics (Table 3). A comparison of the sequences of identified genes with some other bla genes in the NCBI database confirmed that they coded for metallo-ß-lactamases. There is therefore a high probability that the bla genes belong to the chromosomally-encoded B2 or B3 subclass ß-lactamases.
Another antibiotic inactivation gene detected in the tested genomes was fosA. FosA encodes the FosA protein, and is the most common fosfomycin resistance mechanism in Gram-negative bacteria. FosA is a Mn2+, K+ dependent metalloenzyme that catalyzes the addition of glutathione to fosfomycin, thus resulting in antibiotic inactivation. In the tested genomes, the fosA gene was detected in Pseudomonas spp. AM4, AM8 and AM14 strains (Table 7). These results were in accordance with antibiotic resistance profile, as these three strains demonstrated a much narrower inhibition zone around the antibiotic disk (< 22 mm) compared to the other two strains (> 40 mm) (Supp. Table 3).
Finally, the GNAT-family gene was detected in tested genomes The gene encodes GCN5-related N-acetyltransferase (GNAT) responsible for acetylation and inactivation of aminoglycoside antibiotics (Burckhardt and Escalante-Semerena 2019).
Two genes responsible for antibiotic target alteration, gyrA and gyrB, were detected in all tested strains. Antibiotic target alteration is driven by single point mutations in gyrA and gyrB, which reduce the affinity between an antibiotic and its target. This is a very common mechanism of fluoroquinolone resistance detected in Gram-negative bacteria. Mutations in quinolone-resistance determining regions, such as gyrA or gyrB in DNA gyrase, are chromosomally encoded. Research indicates that high-level resistance to fluoroquinolones requires mutations in at least two genes with quinolone-resistance determining regions (Zhang et al. 2015). In the present study, all tested genomes contained single point mutations in the gyrA and gyrB genes. Our phenotypic antibiotic susceptibility test found the inhibition zone around fluoroquinolones (CIP, LEV) to be narrower for all strains (< 30 mm) than the susceptibility zone defined by EUCAST (> 50 mm) (Supp. Tab. S3).
The tested Pseudomonas genomes were found to encode numerous efflux pump components and regulatory protein genes. They included all genes of three efflux pumps, viz. MacAB-TolC, AcrAB-TolC and MexAB-OprM. Moreover, genes encoding components of other efflux pumps were detected: mexE and oprN of the MexEF-OprN pump, mexX and oprM of the MexXY-OprM pump, mexH and mexI of the MexGHI-OpmM pump.
The redox-sensitive protein SoxR is a global regulator of various efflux pump genes. It belongs to the MerR-family transcriptional regulators and it is common among both Gram-negative and Gram-positive bacteria. In enteric bacteria, SoxR mediates resistance to oxidative stress caused by nitric oxide or superoxide, and induces the expression of the soxS gene. SoxS activates the transcription of more than 100 genes encoding products that repair cellular damage. In nonenteric bacteria like Pseudomonas spp., SoxR directly activates the transcription of several multidrug efflux pump genes known to confer resistance to antibiotics (Park et al. 2006).
Discussion
Since the beginning of the XXI century, endophytes inhabiting metallophytic plants have gained increasing attention (El-Deeb et al. 2006; Idris et al. 2004; Stepanauskas et al. 2005; Ma et al. 2015); indeed, by the end of 2022, about 25 metallophytic plants had been tested for bacterial endophytes (Alves et al. 2022; Goryluk-Salmonowicz and Popowska 2019). All tested plants were found to harbor such endophytes, and all bacteria were resistant to high metal concentrations (He et al. 2013; Idris et al. 2004; Ma et al. 2015). Therefore, it was proposed that all metallophytic plants harbor endophytes resistant to metals. Even so, the presence and origin of ARGs in the bacteria, and their antibiotic resistance mechanisms, remain unclear. Seeing that endophytes isolated from hyperaccumulators have recently been used as beneficial biofertilizers, it is important to monitor the presence of ARGs in the genomes to prevent uncontrolled spread of ARGs in the environment.
Several studies have found endophytes inhabiting metallophytes to demonstrate antibiotic resistance. In 2005, Pseudomonas fluorescens and Microbacterium sp. endophytes isolated from Brassica napus (Stepanauskas et al. 2005) were found to be resistant to the antibiotics ampicilin, kanamycin and spectinomycin. In 2006, Enterobacter sp. endophytes resistant to ampicilin, kanamycin and tetracycline were isolated from Eichhornia crassipes (El-Deeb et al. 2006), while in 2015, a Stenotrophomonas sp. strain inhabiting Sedum plumbizincicola resistant to ampicilin, kanamycin and chloramphenicol was detected (Ma et al. 2015). While all these bacteria were found to be resistant to lead, zinc and cadmium, their antibiotic resistance genes were not researched.
In the present study, Pseudomonas spp. endophytes were isolated from the green parts of the hyperaccumulator plant Armeria maritima. The bacteria were resistant to ß-lactam antibiotics, fosfomycin, streptomycin and toxic metals, and demonstrated genes conferring possible resistance to arsenic, cadmium, chromium, cobalt, copper, lead, tellurium and zinc (Tables 5 and 6). Additionally, genes responsible for antibiotic inactivation, antibiotic target alteration, and genes coding efflux pumps were identified (Tables 7 and 8).
Bacteria are known to employ various mechanisms to provide metal resistance (Bruins et al. 2000; Ji and Silver 1995; Niño-Martínez et al. 2019), some of which were observed in the present study; for instance, some strains were found to harbor genes of the CzcCBA system, which are believed to confer resistance to cadmium, cobalt and zinc. Genes of the CzcCBA system can be detected in other Pseudomonas spp. endophytic genomes (Supp. Tab. S6). Previously, these genes were reported in the P. poae A2-S9 genome isolated from Panicum virgatum plant, the Pseudomonas sp.382 genome isolated from Paullinia cupana seeds (Xia et al. 2013, 2019; de Siqueira et al. 2018; Liotti et al. 2018) and the P. putida GM4FR genome isolated from the green parts of Festuca rubra L. (Wemheuer et al. 2016, 2017) (Supp. Tab. S6).
Interestingly, it has been proposed that the CzcRS two-component regulatory system may also be responsible for carbapenem antibiotic resistance (Wang et al. 2017). In the presence of Zn(II) ions, CzcS autophosphorylates and transmits a signal to the response regulator CzcR. CzcR up-regulates the expression of the CzcCBA efflux pump and represses the expression of the OprD porin responsible for the entry of carbapenem antibiotics (Perron et al. 2004). This is an example of a co-regulation system that act as a co-selection mechanism between the metal and antibiotic resistance mechanisms (Baker-Austin et al. 2006; Goryluk-Salmonowicz and Popowska 2019). It is possible that a co-regulation system also operates in the endophytes tested in our present study, as these were found to be resistant to both zinc ions and carbapenem antibiotics (Table 3).
Further, possible chromium resistance genes were detected, one of which codes for ChrA, a membrane potential dependent transporter. The gene has previously been detected in a variety of bacterial genera, including Arthrobacter spp., Bacillus spp., Lysinibacillus spp. and Pseudomonas spp. (He et al. 2010, 2011; Henne et al. 2009; Mondaca et al. 1998). The ChrA gene has also been found in environmental Pseudomonas genomes, like P. chlororaphis GP72 isolated from green pepper rhizosphere, P. viridiflava CDRTc14 isolated from the roots of Lepidium draba and P. fluorescens UM270 isolated from the roots of Medicago truncatula (Supp. Tab. S6) (Hernández-León et al. 2015; Samad et al. 2016; Liu et al. 2006; Shen et al. 2012). Interestingly, the chrA gene is commonly detected on plasmid or Tn DNA. In 1990, it was detected on the Pseudomonas aeruginosa plasmid pUM505, and on the Alcaligenes eutrophus pMOL28 (Cervantes et al. 1990; Nies et al. 1990). The ChrA protein was also found to be encoded by a gene detected on transposon TnOtChr from Ochrobactrum tritici (Branco et al. 2008). Similarly, genes encoding CzcCBA components can be located on mobile elements; these genes have been detected on the plasmid pMOL30 of C. metallidurans (Nies et al. 1990).
Finally, we identified genes associated with three copper resistance mechanisms (CusCFBA efflux system, P-type ATPases and multicopper oxidase CueO) and one zinc/cadmium/lead transporter (P-type ATPase CadA).Arsenic (ArsAB efflux pump), chromium (ChrA transporter) and tellurium (TehAB transporter) metalloid resistance genes were detected in all tested bacterial genomes (Table 5). These genes were also detected in other genomes of environmental Pseudomonas strains (Supp. Tab. S6).
The broad spectrum of metal tolerance demonstrated by the isolated endophytes makes them interesting candidates as bioremediation enhancing agents. However, this raises the important question of whether the presence of metal resistance genes promotes antibiotic resistance. Numerous studies have confirmed that metal-resistant environmental bacteria isolated from soil and water environments harbor antibiotic resistance genes (Barker-Reid et al. 2010; Cycoń et al. 2019; Forsberg et al. 2014; Su et al. 2020). There is a growing concern that metal contamination acts as selective agent in the spread of antibiotic resistance, especially when the metal resistance genes are located on mobile elements (Goryluk-Salmonowicz and Popowska 2019, 2022; Seiler and Berendonk 2012; Zhang et al. 2020; Baker-Austin et al. 2006; Perron et al. 2004). Therefore, the present study examined the locations of any potential resistance genes in the genomes of the tested bacteria.
The Pseudomonas spp. genomes were searched for genes encoding components of mobile genetic elements (MGE). Interestingly, numerous transposase genes were detected in the tested genomes. Transposases are needed for efficient transposition of insertion sequences (IS) or transposon DNA (Tn) (Beuzón et al. 2004). Transposases encoding genes of different Insertion Sequence (IS) families were detected, including IS5, IS66, IS110, IS200like, IS630, InsE, InsO and ISL3. However, it is unlikely that potential ARGs detected in the sequenced Pseudomonas genomes are a part of their mobilome, as the detected ARGs are commonly located on the chromosomes of other Pseudomonas spp. (CARD database). Further research is needed to confirm the presence of the detected ARGs on bacterial chromosomes. If so, the isolated endophytes can be used in laboratory experiments to evaluate their potential to promote plant growth and increase metal pollution remediation capacity. An interesting question is whether all detected potential metal resistance genes encode functional metal resistance proteins.
Conclusion
The present study demonstrated that Armeria maritima subsp. halleri (Wallr.) Rothm. is inhabited by Pseudomonas endophytes that are resistant to metals and antibiotics. Genome analysis confirmed the presence of genes conferring resistance to arsenic, cadmium, chromium, cobalt, copper, lead, tellurium and zinc. Genes encoding components of efflux pumps, extracellular sequestration proteins, P-type ATPases or metal detoxification proteins were detected. Moreover, the bacteria were resistant to antibiotics: streptomycin, fosfomycin and ß-lactams. Genes that may confer resistance to macrolides (MacAB-TolC efflux pump) and multidrug efflux pumps genes (AcrAB-TolC and MexAB-OprM) were identified, as well as some genes that may promote antibiotic inactivation and antibiotic target alteration.
The spread of antibiotic resistance genes (ARGs) in the environment is a global problem, and their main reservoir is considered to be soil. The bacteria inhabiting soil and plants are recipients of ARGs and hence form part of their transmission routes. Therefore, while the bacterial endophytes inhabiting hyperaccumulators may be beneficial for the host plant, they can also hasten the rise of antibiotic resistance.
In the available literature, little attention has been given to the problem of antibiotic resistance in bacteria used to promote plant growth in agriculture, and the resistance profiles of endophytes used in phytoremediation have not been addressed at al. Noteworthily, such biological control agents and biofertilizers form important parts of new agricultural management systems. Such growth in the number of biopreparations available on the market, and the consequent large-scale and long-term usage of biopreparations containing ARGs may further increase the dissemination of antibiotic resistance.
Data Availability
The datasets generated during the current study are available from the corresponding author on reasonable request.
References
Ahemad M (2015) Enhancing phytoremediation of chromium-stressed soils through plant-growth-promoting bacteria. J Genet Eng Biotechnol 13:51–58. https://doi.org/10.1016/j.jgeb.2015.02.001
Alves AR, Yin Q, Oliveira RS, Silva EF, Novo LA (2022) Plant growth-promoting bacteria in phytoremediation of metal-polluted soils: current knowledge and future directions. Sci Total Environ. https://doi.org/10.1016/j.scitotenv.2022.156435
Arber W (2014) Horizontal gene transfer among bacteria and its role in biological evolution. Life 4:217–224. https://doi.org/10.3390/life4020217
Baker-Austin C, Wright MS, Stepanauskas R, McArthur JV (2006) Co-selection of antibiotic and metal resistance. Trends Microbiol 14:176–182. https://doi.org/10.1016/j.tim.2006.02.006
Banasiewicz J, Granada CE, Lisboa BB, Grzesiuk M, Matuśkiewicz W, Bałka M, Stępkowski T, Banasiewicz J, Granada CE, Lisboa BB, Grzesiuk M, Matuśkiewicz W, Bałka M, Schlindwein G, Vargas LK, Passaglia LMP, Stępkowski T (2021) Diversity and phylogenetic affinities of Bradyrhizobium isolates from Pampa and Atlantic Forest Biomes. Syst Appl Microbiol 44:126203. https://doi.org/10.1016/j.syapm.2021.126203
Bargaz A, Lyamlouli K, Chtouki M, Zeroual Y, Dhiba D (2018) Soil microbial resources for improving fertilizers efficiency in an integrated plant nutrient management system. Front Microbiol 9:1606. https://doi.org/10.3389/fmicb.2018.01606
Barker-Reid F, Fox EM, Faggian R (2010) Occurrence of antibiotic resistance genes in reclaimed water and river water in the Werribee Basin, Australia. J Water Health 8:521–531. https://doi.org/10.2166/wh.2010.102
Behzadi P, García-Perdomo HA, Karpiński TM, Issakhanian L (2020) Metallo-ß-lactamases: a review. Mol Biol Rep 47:6281–6294. https://doi.org/10.1007/s11033-020-05651-9
Bergey DH (1994) Bergey’s manual of determinative bacteriology. WR Hensyl. (ed) Lippincott Williams & Wilkins, Philadelphia, USA
Beuzón CR, Chessa D, Casadesús J (2004) IS200: an old and still bacterial transposon. Int Microbiol 7:3–12
Bhattacharjee H, Rosen BP (2007) Arsenic metabolism in prokaryotic and eukaryotic microbes. In: Nies D, Silver S (eds) Molecular microbiology of heavy metals. Springer, Berlin Heidelberg, pp 371–406
Bondarczuk K, Piotrowska-Seget Z (2013) Molecular basis of active copper resistance mechanisms in Gram-negative bacteria. Cell Biol Toxicol 29:397–405. https://doi.org/10.1007/s10565-013-9262-1
Branco R, Chung AP, Johnston T, Gurel V, Morais P, Zhitkovich A (2008) The chromate-inducible chrBACF operon from the transposable element Tn OtChr confers resistance to chromium (VI) and superoxide. JBacteriol 190:6996–7003. https://doi.org/10.1128/JB.00289-08
Brettin T, Davis JJ, Disz T, Edwards RA, Gerdes S et al (2015) RASTtk: a modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci Rep 5:8365. https://doi.org/10.1038/srep08365
Bruins MR, Kapil S, Oehme FW (2000) Microbial resistance to metals in the environment. Ecotoxicol Environ Saf 45:198–207. https://doi.org/10.1006/eesa.1999.1860
Buchholz U, Bernard H, Werber D, Böhmer MM, Remschmidt C, Wilking H, Kühne M, Buchholz U, Bernard H, Werber D, Böhmer MM, Remschmidt C, Wilking H, Deleré Y, an der Heiden M, Adlhoch C, Dreesman J, Ehlers J, Ethelberg S, Faber M, Frank C, Fricke G, Greiner M, Höhle M, Ivarsson S, Jark U, Kirchner M, Koch J, Krause G, Luber P, Rosner B, Stark K, Kühne M (2011) German outbreak of Escherichia coli O104: H4 associated with sprouts. N Engl J Med 365:1763–1770. https://doi.org/10.1056/NEJMoa1106482
Burckhardt RM, Escalante-Semerena JC (2019) Insights into the function of the N-acetyltransferase SatA that detoxifies streptothricin in Bacillus subtilis and Bacillus anthracis. Appl Environ Microbiol 85:e03029-18. https://doi.org/10.1128/AEM.03029-18
Cadena M, Durso LM, Miller DN, Waldrip HM, Castleberry BL, Drijber RA, Wortmann C (2018) Tetracycline and sulfonamide antibiotic resistance genes in soils from Nebraska organic farming operations. Front Microbiol 9:1283. https://doi.org/10.3389/fmicb.2018.01283
Cervantes C, Ohtake H, Chu L, Misra TK, Silver S (1990) Cloning, nucleotide sequence, and expression of the chromate resistance determinant of Pseudomonas aeruginosa plasmid pUM505. J Bacteriol 172:287–291. https://doi.org/10.1128/jb.172.1.287-291.1990
Chen QL, Cui HL, Su JQ, Penuelas J, Zhu YG (2019) Antibiotic resistomes in plant microbiomes. Trends Plant Sci 24:530–541. https://doi.org/10.1016/j.tplants.2019.02.010
Colom K, Pérez J, Alonso R, Fernández-Aranguiz A, Lariño E, Cisterna R (2003) Simple and reliable multiplex PCR assay for detection of bla TEM, bla SHV and bla OXA–1 genes in Enterobacteriaceae. FEMS Microbiol Lett 223:147–151. https://doi.org/10.1016/S0378-1097(03)00306-9
Cycoń M, Mrozik A, Piotrowska-Seget Z (2019) Antibiotics in the soil environment-degradation and their impact on microbial activity and diversity. Front Microbiol 10:338. https://doi.org/10.3389/fmicb.2019.00338
Dallenne C, Da Costa A, Decré D, Favier C, Arlet G (2010) Development of a set of multiplex PCR assays for the detection of genes encoding important β-lactamases in Enterobacteriaceae. J Antimicrob Chemother 65:490–495. https://doi.org/10.1093/jac/dkp498
De Siqueira KA, Liotti RG, Mendes TADO, Soares MA (2018) Draft genome sequences of Pseudomonas sp. strain 382 and Pantoea coffeiphila 342, endophytic bacteria isolated from Brazilian Guarana [Paullinia cupana (Mart.) Ducke]. Genome Announc. https://doi.org/10.1128/genomeA.00287-18
El-Deeb BA, El-Sharouny HM, Fahmy N (2006) Plasmids incidence, antibiotic and heavy metal resistance patterns of endophytic bacteria isolated from aquatic plant, Eichhornia crassipes. J Bot 33:151–171
Forsberg KJ, Patel S, Gibson MK, Lauber CL, Knight R, Fierer N, Dantas G (2014) Bacterial phylogeny structures soil resistomes across habitats. Nature 509:612–616. https://doi.org/10.1038/nature13377
Forsberg KJ, Reyes A, Wang B, Selleck EM, Sommer MO, Dantas G (2012) The shared antibiotic resistome of soil bacteria and human pathogens. Science 337:1107–1111. https://doi.org/10.1126/science.1220761
Gamalero E, Glick BR (2011) Mechanisms used by plant growth-promoting bacteria. In: Maheshwari D (ed) Bacteria in agrobiology: plant nutrient management, 3rd edn. Springer, pp 17–46
Girard L, Lood C, Rokni-Zadeh H, van Noort V, Lavigne R, De Mot R (2020) Reliable identification of environmental Pseudomonas isolates using the rpoD gene. Microorganisms 8:1166. https://doi.org/10.3390/microorganisms8081166
Glick BR (2012) Plant growth-promoting bacteria: mechanisms and applications. Scientifica. https://doi.org/10.6064/2012/963401
Goryluk A, Rekosz-Burlaga H, Blaszczyk M (2009) Isolation and characterization of bacterial endophytes of Chelidonium majus L. Pol J Microbiol 58:355–361
Goryluk-Salmonowicz A, Popowska M (2019) Występowanie zjawiska koselekcji w środowiskach pozaklinicznych. Postep Mikrobiol. https://doi.org/10.21307/PM-2019.58.4.433. (In Polish)
Goryluk-Salmonowicz A, Popowska M (2022) Factors promoting and limiting antimicrobial resistance in the environment–existing knowledge gaps. Front Microbiol 13:992268. https://doi.org/10.3389/fmicb.2022.992268
Haroun AA, Kamaluddeen KK, Alhaji I, Magaji Y, Oaikhena EE (2017) Evaluation of heavy metal tolerance level (MIC) and bioremediation potentials of Pseudomonas aeruginosa isolated from Makera-Kakuri industrial drain in Kaduna, Nigeria. Eur J Exp Biol. https://doi.org/10.21767/2248-9215.100028
He M, Li X, Guo L, Miller SJ, Rensing C, Wang G (2010) Characterization and genomic analysis of chromate resistant and reducing Bacillus cereus strain SJ1. BMC Microbiol 10:1–10. https://doi.org/10.1186/1471-2180-10-221
He M, Li X, Liu H, Miller SJ, Wang G, Rensing C (2011) Characterization and genomic analysis of a highly chromate resistant and reducing bacterial strain Lysinibacillus fusiformis ZC1. J Hazard Mater 185:682–688. https://doi.org/10.1016/j.jhazmat.2010.09.072
He X, Lu F, Yuan F, Jiang D, Zhao P, Zhu J, Lu G, He X, Lu F, Yuan F, Jiang D, Zhao P, Zhu J, Cheng H, Cao J, Lu G (2015) Biofilm formation caused by clinical Acinetobacter baumannii isolates is associated with overexpression of the AdeFGH efflux pump. Antimicrob Agents Chemother 59:4817–4825. https://doi.org/10.1128/AAC.00877-15
He H, Ye Z, Yang D, Yan J, Xiao L, Zhong T, Jing Y, He H, Ye Z, Yang D, Yan J, Xiao Li, Zhong T, Yuan M, Cai X, Fang Z, Jing Y (2013) Characterization of endophytic Rahnella sp. JN6 from Polygonum pubescens and its potential in promoting growth and cd, pb, zn uptake by Brassica napus. Chemosphere 90:1960–1965. https://doi.org/10.1016/j.chemosphere.2012.10.057
Henne KL, Nakatsu CH, Thompson DK, Konopka AE (2009) High-level chromate resistance in Arthrobacter sp. strain FB24 requires previously uncharacterized accessory genes. BMC Microbiol 9:1–14. https://doi.org/10.1186/1471-2180-9-199
Hernández-León R, Rojas-Solís D, Contreras-Pérez M, del Carmen Orozco-Mosqueda M, Macías-Rodríguez LI, Reyes-de la Cruz H, Santoyo G (2015) Characterization of the antifungal and plant growth-promoting effects of diffusible and volatile organic compounds produced by Pseudomonas fluorescens strains. Biol Control 81:83–92. https://doi.org/10.1016/j.biocontrol.2014.11.011
Idris R, Trifonova R, Puschenreiter M, Wenzel WW, Sessitsch A (2004) Bacterial communities associated with flowering plants of the Ni hyperaccumulator Thlaspi goesingense. Appl Environ Microbiol 70:2667–2677. https://doi.org/10.1128/AEM.70.5.2667-2677.2004
Ji G, Silver S (1995) Bacterial resistance mechanisms for heavy metals of environmental concern. J Ind Microbiol 14:61–75
Khan AG, Kuek C, Chaudhry TM, Khoo CS, Hayes WJ (2000) Role of plants, mycorrhizae and phytochelators in heavy metal contaminated land remediation. Chemosphere 41:197–207. https://doi.org/10.1016/S0045-6535(99)00412-9
Kloepper JW, Leong J, Teintze M, Schroth MN (1980) Enhanced plant growth by siderophores produced by plant growth-promoting rhizobacteria. Nature 286:885–886. https://doi.org/10.1038/286885a0
Kong Z, Glick BR (2017) The role of plant growth-promoting bacteria in metal phytoremediation. Adv Microb Physiol 71:97–132. https://doi.org/10.1016/bs.ampbs.2017.04.001
Kpoda DS, Ajayi A, Somda M, Traore O, Guessennd N, Ouattara AS, Dosso M, Kpoda DS, Ajayi A, Somda M, Traore O, Guessennd N, Ouattara AS, Sangare L, Traore AS, Dosso M (2018) Distribution of resistance genes encoding ESBLs in Enterobacteriaceae isolated from biological samples in health centers in Ouagadougou, Burkina Faso. BMC Res Notes 11:1–5. https://doi.org/10.1186/s13104-018-3581-5
Kumar PN, Dushenkov V, Motto H, Raskin I (1995) Phytoextraction: the use of plants to remove heavy metals from soils. Environ Sci Technol 29:1232–1238. https://doi.org/10.1021/es00005a014
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Liotti RG, da Silva Figueiredo MI, da Silva GF, de Mendonça EAF, Soares MA (2018) Diversity of cultivable bacterial endophytes in Paullinia cupana and their potential for plant growth promotion and phytopathogen control. Microbiol Res 207:8–18. https://doi.org/10.1016/j.micres.2017.10.011
Liu H, Dong D, Peng H, Zhang X, Xu Y (2006) Genetic diversity of phenazine-and pyoluteorin-producing pseudomonads isolated from green pepper rhizosphere. Arch Microbiol 185:91–98. https://doi.org/10.1007/s00203-005-0072-6
Liu S, Yang B, Liang Y, Xiao Y, Fang J (2020) Prospect of phytoremediation combined with other approaches for remediation of heavy metal-polluted soils. Environ Sci Pollut Res Int 27:16069–16085. https://doi.org/10.1007/s11356-020-08282-6
Ma Y, Oliveira RS, Nai F, Rajkumar M, Luo Y, Rocha I, Freitas H (2015) The hyperaccumulator Sedum plumbizincicola harbors metal-resistant endophytic bacteria that improve its phytoextraction capacity in multi-metal contaminated soil. J Environ Manage 156:62–69. https://doi.org/10.1016/j.envman.2015.03.024
Mahdi I, Fahsi N, Hijri M, Sobeh M (2022) Antibiotic resistance in plant growth promoting bacteria: a comprehensive review and future perspectives to mitigate potential gene invasion risks. Front Microbiol 13:999988. https://doi.org/10.3389/fmicb.2022.999988
Malik A, Jaiswal R (2000) Metal resistance in Pseudomonas strains isolated from soil treated with industrial wastewater. World J Microbiol Biotechnol 16:177–182. https://doi.org/10.1023/A:1008905902282
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:1–14. https://doi.org/10.1186/1471-2105-14-60
Mondaca M, González G, Zaror Z (1998) Isolation, characterization and expression of a plasmid encoding chromate resistance in Pseudomonas putida KT2441. Lett Appl Microbiol 26:367–371. https://doi.org/10.1046/j.1472-765X.1998.00349.x
Nies DH (1999) Microbial heavy-metal resistance. Appl Microbiol 51:730–750
Nies A, Nies DH, Silver S (1990) Nucleotide sequence and expression of a plasmid-encoded chromate resistance determinant from Alcaligenes eutrophus. J Biol Chem 265:5648–5653. https://doi.org/10.1016/S0021-9258(19)39411-6
Niño-Martínez N, Salas Orozco MF, Martínez-Castañón GA, Torres Méndez F, Ruiz F (2019) Molecular mechanisms of bacterial resistance to metal and metal oxide nanoparticles. Int J Mol Sci 20:2808. https://doi.org/10.3390/ijms20112808
Olanrewaju OS, Glick BR, Babalola OO (2017) Mechanisms of action of plant growth promoting bacteria. World J Microbiol Biotechnol 33:1–16. https://doi.org/10.1007/s11274-017-2364-9
Pal C, Bengtsson-Palme J, Rensing C, Kristiansson E, Larsson DJ (2014) BacMet: antibacterial biocide and metal resistance genes database. Nucleic Acids Res 42:D737–D743. https://doi.org/10.1093/nar/gkt1252
Park W, Peña-Llopis S, Lee Y, Demple B (2006) Regulation of superoxide stress in Pseudomonas putida KT2440 is different from the SoxR paradigm in Escherichia coli. Biochem Biophys Res Commun 341:51–56. https://doi.org/10.1016/j.bbrc.2005.12.142
Perron K, Caille O, Rossier C, Van Delden C, Dumas JL, Köhler T (2004) CzcR-CzcS, a two-component system involved in heavy metal and carbapenem resistance in Pseudomonas aeruginosa. J Biol Chem 279:8761–8768. https://doi.org/10.1074/jbc.M312080200
Piotrowska M, Kowalska S, Popowska M (2019) Diversity of β-lactam resistance genes in gram-negative rods isolated from a municipal wastewater treatment plant. Ann Microbiol 69:591–601. https://doi.org/10.1007/s13213-019-01450-1
Poonsuk K, Chuanchuen R (2014) Detection of the mex efflux pumps in Pseudomonas aeruginosa by using a combined resistance-phenotypic markers and multiplex RT-PCR. Open J Med Microbiol. https://doi.org/10.4236/ojmm.2014.43018
Rosen BP (2002) Transport and detoxification systems for transition metals, heavy metals and metalloids in eukaryotic and prokaryotic microbes. Comp Biochem Phys A 133:689–693. https://doi.org/10.1016/S1095-6433(02)00201-5
Rossbach S, Wilson TL, Kukuk ML, Carty HA (2000) Elevated zinc induces siderophore biosynthesis genes and a znta-like gene in Pseudomonas fluorescens. FEMS Microbiol Lett 191:61–70. https://doi.org/10.1111/j.1574-6968.2000.tb09320.x
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Samad A, Trognitz F, Antonielli L, Compant S, Sessitsch A (2016) High-quality draft genome sequence of an endophytic Pseudomonas viridiflava strain with herbicidal properties against its host, the weed Lepidium draba L. Genome Announc 4:e01170-16. https://doi.org/10.1128/genomeA.01170-16
Seiler C, Berendonk TU (2012) Heavy metal driven co-selection of antibiotic resistance in soil and water bodies impacted by agriculture and aquaculture. Front Microbiol. https://doi.org/10.3389/fmicb.2012.00399
Sharma RK, Agrawal M, Marshall F (2007) Heavy metal contamination of soil and vegetables in suburban areas of Varanasi, India. Ecotoxicol Environ Saf Ecotoxicol Environ Saf 66:258–266. https://doi.org/10.1016/j.ecoenv.2005.11.007
Shen X, Chen M, Hu H, Wang W, Peng H, Xu P, Zhang X (2012) Genome sequence of Pseudomonas chlororaphis GP72, a root-colonizing biocontrol strain. J Bacteriol 194:1269–1270. https://doi.org/10.1128/JB.06713-11
Silambarasan S, Logeswari P, Valentine A, Cornejo P, Kannan VR (2020) Pseudomonas citronellolis strain SLP6 enhances the phytoremediation efficiency of Helianthus annuus in copper contaminated soils under salinity stress. Plant Soil 457:241–253. https://doi.org/10.1007/s11104-020-04734-7
Spain A, Alm E (2003) Implications of microbial heavy metal tolerance in the environment. Rev Undergraduate Res 2:1–6
Stepanauskas R, Glenn TC, Jagoe CH, Tuckfield RC, Lindell AH, McArthur JV (2005) Elevated microbial tolerance to metals and antibiotics in metal-contaminated industrial environments. Environ Sci Technol 39:3671–3678
Su S, Li C, Yang J, Xu Q, Qiu Z, Xue B, Su S, Li C, Yang J, Xu Q, Qiu Z, Xue B, Wang S, Zhao C, Xiao Z, Wang J, Shen Z (2020) Distribution of antibiotic resistance genes in three different natural water bodies-a lake, river and sea. Int J Environ Res Public Health 17:552. https://doi.org/10.3390/ijerph17020552
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci 101:11030–11035. https://doi.org/10.1073/pnas.0404206101
Tan L, Li L, Ashbolt N, Wang X, Cui Y, Zhu X, Luo Y, Tan Lu, Li L, Ashbolt N, Wang X, Cui Y, Zhu X, Xu Y, Yang Y, Mao D, Luo Yi (2018) Arctic antibiotic resistance gene contamination, a result of anthropogenic activities and natural origin. Sci Total Environ 621:1176–1184. https://doi.org/10.1016/j.scitotenv.2017.10.110
Trafas M, Eckes T, Gołda T (2006) Lokalna zmienność zawartości metali ciężkich w glebach okolicy Olkusza. Inżynieria Środowiska/Akademia Górniczo-Hutnicza im. S Staszica w Krakowie 11:127–144 (In Polish)
Turner RJ, Weiner JH, Taylor DE (1995) The tellurite-resistance determinants tehAtehB and klaAklaBtelB have different biochemical requirements. Microbiology 141:3133–3140. https://doi.org/10.1099/13500872-141-12-3133
Ugwuanyi FC, Ajayi A, Ojo DA, Adeleye AI, Smith SI (2021) Evaluation of efflux pump activity and biofilm formation in multidrug resistant clinical isolates of Pseudomonas aeruginosa isolated from a Federal Medical Center in Nigeria. Ann Clin Microbiol Antimicrob 20:1–7. https://doi.org/10.1186/s12941-021-00417-y
Walker DJ, Clemente R, Roig A, Bernal MP (2003) The effects of soil amendments on heavy metal bioavailability in two contaminated Mediterranean soils. Environ Pollut 122:303–312. https://doi.org/10.1016/S0269-7491(02)00287-7
Wang D, Chen W, Huang S, He Y, Liu X, Hu Q, Chen H (2017) Structural basis of zn (II) induced metal detoxification and antibiotic resistance by histidine kinase CzcS in Pseudomonas aeruginosa. PLoS Pathog. https://doi.org/10.1371/journal.ppat.1006533
Wang X, Luo S, Chen Y, Zhang R, Lei L, Lin K, Xu H, Wang X, Luo S, Chen Y, Zhang R, Lei L, Lin K, Qiu C, Xu H (2023) Potential of Miscanthus floridulus associated with endophytic bacterium Bacillus cereus BL4 to remediate cadmium contaminated soil. Sci Total Environ 857:159384. https://doi.org/10.1016/j.scitotenv.2022.159384
Wemheuer F, Hollensteiner J, Poehlein A, Granzow S, Daniel R, Vidal S, Wemheuer B (2017) Draft genome sequence of Pseudomonas putida strain GM4FR, an endophytic bacterium isolated from Festuca rubra L. Genome Announc 5:e00086-17. https://doi.org/10.1128/genomeA.00086-17
Wemheuer F, Wemheuer B, Kretzschmar D, Pfeiffer B, Herzog S, Daniel R, Vidal S (2016) Impact of grassland management regimes on bacterial endophyte diversity differs with grass species. Lett Appl Microbiol 62:323–329. https://doi.org/10.1111/lam.12551
Wu K, Luo J, Li J, An Q, Yang X, Liang Y, Li T (2018) Endophytic bacterium Buttiauxella sp. SaSR13 improves plant growth and cadmium accumulation of hyperaccumulator Sedum alfredii. Environ Sci Pollut Res Int 25:21844–21854. https://doi.org/10.1007/s11356-018-2322-6
Xia Y, DeBolt S, Ma Q, McDermaid A, Wang C, Shapiro N, Kyrpides NC, Xia Ye, DeBolt S, Ma Q, McDermaid A, Wang C, Shapiro N, Woyke T, Kyrpides NC (2019) Improved draft genome sequence of Pseudomonas poae A2-S9, a strain with plant growth-promoting activity. Microbiol Resour Announc 8:e00275-19. https://doi.org/10.1128/MRA.00275-19
Xia Y, Greissworth E, Mucci C, Williams MA, De Bolt S (2013) Characterization of culturable bacterial endophytes of switchgrass (Panicum virgatum L.) and their capacity to influence plant growth. Gcb Bioenergy 5:674–682. https://doi.org/10.1111/j.1757-1707.2012.01208.x
Zhang S, Abbas M, Rehman MU, Huang Y, Zhou R, Gong S, Cheng A, Zhang S, Abbas M, Rehman MU, Huang Y, Zhou R, Gong S, Yang H, Chen S, Wang M, Cheng A (2020) Dissemination of antibiotic resistance genes (ARGs) via integrons in Escherichia coli: a risk to human health. Environ Pollut 266:115260. https://doi.org/10.1016/j.envpol.2020.115260
Zhang L, Kinkelaar D, Huang Y, Li Y, Li X, Wang HH (2011) Acquired antibiotic resistance: are we born with it? Appl Environ Microbiol 77:7134–7141. https://doi.org/10.1128/AEM.05087-11
Zhang X, Lin L, Zhu Z, Yang X, Wang Y, An Q (2013) Colonization and modulation of host growth and metal uptake by endophytic bacteria of Sedum alfredii. Int J Phytoremediat 15:51–64. https://doi.org/10.1080/15226514.2012.670315
Zhang QQ, Ying GG, Pan CG, Liu YS, Zhao JL (2015) Comprehensive evaluation of antibiotics emission and fate in the river basins of China: source analysis, multimedia modeling, and linkage to bacterial resistance. Environ Sci Technol 49:6772–6782. https://doi.org/10.1021/acs.est.5b00729
Acknowledgements
We are grateful to Professor Mieczysław Błaszczyk for his assistance in the fieldwork. This work was supported by the National Science Centre and the national Ministry of Education and Science.
Funding
This work was supported by the National Science Centre (NCN), Poland (UMO-2019/32/Z/NZ8/0011), international project in the frame of the Biodiversity and its influence on human, animal and plant health (BiodivERsA Call 2018): project “ANTIVERSA – Biodiversity as an ecological barrier for the spread of clinically relevant antibiotic resistance in the environment” to MP and partially in the frame of the ‘Excellence Initiative-Research University (2020–2026)’ Program at the University of Warsaw. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Author information
Authors and Affiliations
Contributions
A.G-S. – Conceptualization, Investigation, Formal Analysis, Writing, Funding acquisition, Original Draft, Visualization; AW.M. – Investigation; M.P. – Writing, Review & Editing, Supervision, Funding Acquisition, Project Administration.
All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationship that could be construed as a potential conflict of interest.
Additional information
Responsible Editor: Antony Van der Ent.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
ESM 1
(DOC 192 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Goryluk-Salmonowicz, A., Myczka, A.W. & Popowska, M. Antibiotic- and metal-resistant endophytes inhabit Armeria maritima hyperaccumulator. Plant Soil 495, 57–76 (2024). https://doi.org/10.1007/s11104-023-06320-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-023-06320-z