Abstract
Aims
Different drivers are known to shape rhizosphere microbiome assembly. How soil texture (Texture) and presence or lack of root hairs (Root Hair) of plants affect the rhizosphere microbiome assembly and soil potential extracellular enzyme activities (EEA) at defined rooting depth (Depth) is still a knowledge gap. We investigated effects of these drivers on microbial assembly in rhizosphere and on potential EEA in root-affected soil of maize.
Methods
Samples were taken from three depths of root hair defective mutant rth3 and wild-type WT maize planted on loam and sand in soil columns after 22 days. Rhizosphere bacterial, archaeal, fungal and cercozoan communities were analysed by sequencing of 16S rRNA gene, ITS and 18S rRNA gene fragments. Soil potential EEA of ß-glucosidase, acid phosphatase and chitinase were estimated using fluorogenic substrates.
Results
The bacterial, archaeal and cercozoan alpha- and beta-diversities were significantly and strongly altered by Texture, followed by Depth and Root Hair. Texture and Depth had a small impact on fungal assembly, and only fungal beta-diversity was significantly affected. Significant impacts by Depth and Root Hair on beta-diversity and relative abundances at taxonomic levels of bacteria, archaea, fungi and cercozoa were dependent on Texture. Likewise, the patterns of potential EEA followed the trends of microbial communities, and the potential EEA correlated with the relative abundances of several taxa.
Conclusions
Texture was the strongest driver of rhizosphere microbiome and of soil potential EEA, followed by Depth and Root Hair, similarly to findings in maize root architecture and plant gene expression studies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The rhizosphere is defined as soil influenced by plant roots, it represents a zone with complex and dynamic interactions among plant roots, soil and biota (Hartmann et al. 2008, 2009). These interactions are modulated by physicochemical conditions such as pH, organic carbon and altered moisture. A wide variety of organisms inhabits the rhizosphere including bacteria, archaea, fungi, cercozoa, nematodes, algae, viruses and arthropods (Mendes et al. 2013). Rhizosphere microorganisms are attracted from the bulk soil to the plant roots via root exudates that consist of numerous primary and secondary metabolites serving as energy or carbon sources for their growth (Kawasaki et al. 2016; Sasse et al. 2018; Canarini et al. 2019; Cotton et al. 2019). In turn, rhizosphere microorganisms alter plant root exudations through systemic microbe-root signalling mechanisms (Korenblum et al. 2020), and contribute significantly to plant growth and health by enhancing nutrient acquisition, pathogen resistance and stress tolerance (Mendes et al. 2013; Berg et al. 2017; Mohanram and Kumar 2019). Rhizosphere inhabiting bacteria, archaea, fungi and cercozoa – hereafter termed rhizosphere microbiome - contribute to soil quality and health. Extracellular enzyme activities (EEA) produced by members of the soil microbiome accelerate the breakdown of complex organic substances for gaining energy and nutrients from soil (Nannipieri et al. 2002; Kompała-Bąba et al. 2021). Thus, they regulate the availability of nutrients for plants (Nannipieri et al. 2003; Dick 1997). Phosphomonoesterases or acid phosphatases are enzymes that catalyse the hydrolysis of organic monophosphoesters, releasing phosphate for plant and microbial uptake (Nannipieri et al. 2011). Other enzymes like β-glucosidases generally play a role in the last stage of cellulose degradation. They hydrolyze cellobiose residues, resulting in the release of sugar monomers, which serve as a source of energy for microbial growth and activity (Gil-Sotres et al. 2005; Merino et al. 2016). N-acetyl-β-d-glucosaminidases or chitinases are involved in the degradation of chitin (fungal cell wall component) and peptidoglycan (bacterial cell wall component) (Ekenler and Tabatabai 2004). Soil enzyme activities depend on the abundance and diversity of the microbiome and its metabolic activity (Zhang et al. 2017; Kompała-Bąba et al. 2021). However, our understanding of the relationship between different types of soil enzymes, their activities and corresponding microbial abundance and diversity is still limited.
In the rhizosphere microbial abundance and diversity is modulated by different drivers such as plant species, genotype and developmental stage (Smalla et al. 2001; Li et al. 2014; Walters et al. 2018; Brisson et al. 2019; Kong et al. 2020), metabolite composition of root exudates (Cotton et al. 2019; Murphy et al. 2021), root traits (Zai et al. 2021; Wei et al. 2021), soil type (Peiffer et al. 2013; Gomes et al. 2018; Seaton et al. 2020; Yim et al. 2015, 2020), soil texture (Hartmann et al. 2014), temperature and precipitation (Walters et al. 2018; Beirinckx et al. 2020) and microbial interactions (Jousset et al. 2008; Rosenberg et al. 2009). Association between bacterial, archaeal, fungal and cercozoan communities and diversity in the rhizosphere was largely ignored in previous works (Jousset et al. 2008; Rosenberg et al. 2009). In soil, most cercozoa are bacterivorous, and due to their selective feeding preferences they affect mainly bacterial community composition (Glücksman et al. 2010; Flues et al. 2017; Dumack et al. 2019). However, bacteria can also alter the cercozoan communities through the production of secondary metabolites (Jousset et al. 2008).
Soils differing in texture have differences in pore size distribution and connectivity and rooting space of plants as observed for the two soils loam and sand used as substrates to grow maize plants (Phalempin et al. 2021). Smaller pore sizes can increase bacterial diversity in soil due to reduced connectivity of microsites (Carson et al. 2010; Hemkemeyer et al. 2018; Seaton et al. 2020), and reduced access of predators (Rutherford and Juma 1992). Some microbial taxa show preference for different particle sizes of the soil, i.e., clay and silt were shown to be more preferred substrates for specific bacteria than sand particles due to their higher cation exchange capacity and nutrient content with consequences for the composition of the soil microbiome (Hemkemeyer et al. 2018; Seaton et al. 2020).
Most studies to date investigated rhizosphere microbiome assembly using composite samples of the whole root system of plants (Li et al. 2014; Silva et al. 2017; Gomes et al. 2018; Walters et al. 2018; Renoud et al. 2020; Kusstatscher et al. 2020) despite the great heterogeneity of root system morphology and architecture (Lippold et al. 2021). Although several studies reported the microbial assembly at different scales of the plant root, e.g., at different root zones along the primary root axis (Rüger et al. 2021), at different root types (DeAngelis et al. 2009; Kawasaki et al. 2016; Pervaiz et al. 2020), root sizes (Zai et al. 2021) and root age (Wei et al. 2021), there is still lack of information regarding the microbial assembly in the rhizosphere along defined soil and rooting depth.
Root hairs were shown to play a significant role in the uptake of water and nutrients, especially in soils with low phosphate (P) contents (Leitner et al. 2010; Klamer et al. 2019). Under limited P and water shortage of soil, root hairs were known to significantly enhance P uptake compared to a root hairless mutant of barley (Ruiz et al. 2020). Especially organic acids are important rhizodeposits of root hairs that help in the acquisition of P (Pantigoso et al. 2020), and may influence the rhizosphere microbiome (Robertson-Albertyn et al. 2017). Further, the presence of root hairs also increases formation of a rhizosheath (Burak et al. 2021). The role of root hairs in microbial colonization was previously examined in barley by Robertson-Albertyn et al. (2017) who reported that root hairless barley mutants had a less complex bacterial community with lower richness and diversity than the wild type in the rhizosphere. However, the impact of root hairs on the microbial assembly remains poorly understood. The two root hairless mutant lines of barley also showed significantly distinct microbial community composition and diversity (Robertson-Albertyn et al. 2017).
Maize (Z. mays) is a plant species with a wealth of data on root traits and rhizosphere microorganisms (reviewed recently by Bonkowski et al. 2021), and thus serves as an excellent model to address specific questions of rhizosphere biology (Vetterlein et al. 2021; Rüger et al. 2021).
In this study, we aimed to investigate spatial microbial colonization patterns along the roots of maize grown in soil columns (Fig. 1) in relation to 1) soil texture (Texture: loam and sand), 2) rooting depths (Depth: D1, D2 and D3) and 3) absence or presence of root hairs (Root Hair: root hair defective mutant rth3 and the corresponding wild-type WT). Further, we related soil potential EEA to the microbial abundance, diversity and dominant taxa in order to understand the mechanisms of plant–microbial interactions in the rhizosphere. Based on root gene expression (Ganther et al. 2021) and root system architecture (Lippold et al. 2021) analyses at D1, D2 and D3 of the maize plants, we also learned that increasing rooting depth means increasing share of young roots (≤ 7 days).
22-day-old maize plant cultivated in a soil column (WT in loam, left) under a control environment and maize root system architectures from different treatments obtained from X-ray computer tomography (right). Indicated depths were used for the rhizosphere and root-affected soil sampling. L, loam and S, sand
The following hypotheses were assessed: 1) loam as Texture with higher content of organic matter and sorption capacity but with lower porosity (Lippold et al. 2021) shows a higher microbial abundance and diversity and potential EEA (Vmax and Km) than sand; 2) particularly high differences between microbial community structure are expected between uppermost and lowest rooting depths. This stratification is dependent on porosity and water-holding capacity of the Texture, which means that a bigger difference is expected in sand; and 3) the effect of Root Hair on the microbial community assembly in the rhizosphere of maize, and potential EEA in the root-affected soil are more subtle compared to Texture and Depth.
Materials and methods
Soil column experiment and collection of samples
To address our hypotheses, we took samples from a soil column experiment (see Ganther et al. 2021) using a three-factorial design: Texture, Depth and Root Hair.
Briefly, soil columns (25 cm height and 7 cm inner diameter) were either packed with two soils with contrasting textures loam (33.2% sand, 47.7% silt, 19.1% clay, 0.84% C org, 0.084% Nt and pH 6.21) and sand. A mixture of 16.7% loam and 83.3% quartz sand was prepared for the Texture sand (88.6% sand, 8.1% silt, 3.3% clay, 0.14% C org, 0.014% Nt and pH 6.25). Loam and sand (≤ 1 mm mesh size) were fertilized differently to achieve a similar content of plant available nutrients for plant growth based on pre-trials (Vetterlein et al. 2021).
Two maize genotypes differing in root hair formation B73 wild-type (WT) and the root hair defective mutant (rth3) were used (Hochholdinger et al. 2008). Planted columns (six replicates per treatment) were grown in a growth chamber for 22 days under controlled conditions (12 h at 22 °C/12 h at 18 °C for day/night, 65% relative humidity and 350 μmol m−2 s−1 photo-synthetically active radiation). A volumetric water content of 22 and 18% (v/v) was maintained for loam and sand, respectively.
Rhizospheres and root-affected soil were sampled at three depths (D1: 4.5–6.1, D2: 9.0–10.6 and D3: 13.5–15.1 cm from soil surface, Fig. 1). A soil slice of a thickness of approximately 1.6 cm at each depth was taken. Root segments from each depth were gently shaken to remove excessive soil before being briefly submerged, and shaken in a 15 mL centrifuge tube containing sterile 0.3% NaCl. The soil obtained from shaking off the roots is determined as root-affected soil. While the rhizosphere was obtained by centrifugation of soil resuspension at 5000 x g for 30 min at 4 °C. The rhizosphere pellet was stored at −20 °C until the total community (TC-) DNA was extracted.
TC-DNA extraction and amplification of 16S rRNA gene fragments and ITS regions for real-time PCR analysis
The TC-DNA was isolated from about 0.52 ± 0.01 g per sample using the FastDNA Spin Kit and the Geneclean Spin Kit for soil following the manufacturer’s instructions (MP Biomedicals, Heidelberg, Germany).
Copy numbers of bacterial 16S rRNA gene fragments and ITS (internal transcribed spacer) regions were quantified in rhizosphere TC-DNAs by real-time quantitative PCR (qPCR). The qPCR for the 16S rRNA gene fragments was conducted as described in Suzuki et al. (2000) and Yim et al. (2015), using 1.2 μM BACT1369F as the forward primer (5′ -CGGTGAATACGTTCYCGG-3′) and 1 μM PROK1492R as the reverse primer (5’-GGWTACCTTGTTACGACTT-3′). The qPCR reaction mixture (50 μL) of ITS regions consisted of 1x PCR GoTaq buffer (Promega GmbH, Walldorf, Germany), 0.2 mM dNTPs, 2.5 mM MgCl2, 4% DMSO (dimethyl sulfoxide), 0.4 μM ITS1 as the forward primer (5-TCCGTAGGTGAACCTGCG G -3), 0.4 μM ITS4 as the reverse primer (5-TCCTCCGCTTATTGATATGC-3), 1x EvaGreen, 1.25 U GoTaq (Promega GmbH, Walldorf, Germany) and 1 μL TC-DNA (ca. 3 ng). The thermal cycling program was applied as previously described by Gschwendtner et al. (2010).
Library preparations for amplicon sequencing of 16S rRNA gene, ITS region and 18S rRNA gene fragments
Sequencing libraries of 16S rRNA gene fragments (bacteria and archaea) and ITS2 regions (fungi) were prepared as previously described (Ganther et al. 2020; Yim et al. 2020). Briefly, the primers Uni341F (5’-CCTAYGGGRBGCASCAG-3′) and Uni806R (5’-GGACTACNNGGGTATCTAAT-3′) were used for the first PCR step to flank approximately 460 bp of the V3-V4 regions of the 16S rRNA gene fragments, targeting bacterial (about 80–100%) and archaeal sequences (Sundberg et al. 2013). For fungi, the primers gITS7 (5’-GTGARTCATCGARTCTTTG-3′) and ITS4 (5’-TCCTCCGCTTATTGATATGC-3′) were applied to obtain the ITS2 regions (Ihrmark et al. 2012). Individual sample tags obtained from the second PCR step were purified using a HighPrep™ PCR Clean Up System (MagBio Genomics Inc., MD, USA) to remove DNA fragments below 100 bp in size. Normalized samples using SequalPrep Normalization Plate (96) Kit (Invitrogen, CA, USA), were pooled using 5 μL volume each, and were concentrated using the DNA Clean and Concentrator™-5 kit (Zymo Research, Irvine, USA). The pooled library concentration was adjusted to 4 nM, and the amplicon sequencing was performed on an Illumina MiSeq platform using Reagent Kit v2 [2 × 250 cycles] (Illumina Inc., CA, USA).
To identify cercozoan communities, amplicon sequencing of the 18S rRNA gene fragments covering about 350 bp of the V4 region was applied as described by Fiore-Donno et al. (2020). Briefly, in a first PCR the forward primers S615F_Cerco (5’-GTTAAAAAGCTCGTAGTTG-3′) and S615F_Phyt (5′- GTTAAAARGCTCGTAGTCG-3′) and the reverse primer S963R_Phyt (5’-CAACTTTCGTTCTTGATTAAA-3′) were used. Then in a semi-nested PCR, the forward primer S615F_Cer (5’GTTAAAARGCTCGTAGTYG-3′) and the reverse primer S947R_Cer (5‘-AAGARGAYATCCTTGGTG-3′), both tagged with barcodes, were used. To obtain a concentration of 1–2 ng μL−1 per sample, replicated PCR-products of each sample were pooled for purification and normalization with the SequalPrep Normalization Plate Kit (Invitrogen, CA, USA). Purified amplicons were pooled, concentrated, and finally sequenced on an Illumina MiSeq platform using Reagent Kit v3 [2 × 300 cycles] (Illumina Inc., San Diego, CA, USA) at the Cologne Centre for Genomics (Köln, Germany).
Data analysis of amplicon sequencing
Demultiplexed sequences of the 16S rRNA gene and ITS region fragments were processed as previously described (Ganther et al. 2020; Yim et al. 2020). Briefly, raw sequence reads of the bacterial and archaeal communities were trimmed for primers using cutadapt v.2.3 (Martin 2011). Primer-trimmed sequence reads were error-corrected and merged, and amplicon sequence variants (ASVs, 100% identity) were identified using DADA2 v.1.10.0 (Callahan et al. 2016) within QIIME2 (Bolyen et al. 2018). Each ASV was given a taxonomic annotation using q2-feature classifier classify-sklearn module trained with SILVA SSU rel. 132 database (Quast et al. 2013). For the fungal community, an automated ITS pipeline, so-called PIPITS (PIPITS_PREP, PIPITS_FUNITS and PIPITS_PROCESS) with default parameters for the ITS2 region was followed according to Gweon et al. (2015). Read-pairs from Illumina MiSeq sequencers of ITS files were merged into a single file, followed by quality filtered for the next step, generated by the PIPITS_PREP. Then, the PIPITS_FUNITS identified the ITS sub-regions using HMMER3 according to Mistry et al. (2013). The PIPITS_PROCESS generated ASVs, and calculated their read abundances as well as the RDP taxonomic assignments using the UNITE fungal ITS reference data set.
Regarding the cercozoan community, sequence reads of the 18S rRNA gene fragments were processed using the customized MOTHUR pipeline v.39.5 (Schloss et al. 2009). Paired-end reads were merged, not allowing any mismatches in primer or barcode sequences, maximum two mismatches and one ambiguity in the target sequence. Assembled sequences with an overlap lower than 200 bp were removed. Merged contigs were demultiplexed, and primers and tag sequences were trimmed. Remaining reads were clustered into operational taxonomic units (OTUs) using VSEARCH (Rognes et al. 2016) according to the abundance-based greedy algorithm (agc) with a similarity threshold of 97%. Clusters represented by less than 350 reads were removed as likely to represent amplification or sequencing noise (Fiore-Donno et al. 2018). OTUs were assigned to taxa using BLAST+ (Camacho et al. 2009) with an e-value of 1e-50 and the PR2 database (Guillou et al. 2013), keeping only the best hit. Sequences were aligned with the template provided by Fiore-Donno et al. (2018), allowing gaps of maximum five nucleotides, and cleaned from chimeras using UCHIME (Edgar et al. 2011) and non-cercozoan sequences.
Sequence contingency tables showing taxonomy identifications and read abundances (ASV or OTU table) were exported for subsequent analyses. For 14 samples, below 4,000 reads were obtained after removing any reads associated to plant materials such as chloroplast and mitochondrial DNA, and these samples were excluded from the analyses. Rarefaction analyses for 16S rRNA gene fragments (bacteria and archaea), ITS regions (fungi) and 18S rRNA gene fragments (cercozoa) were performed, and they showed that the sequences covered the diversity in the analysed samples (Fig. S1).
Soil potential extracellular enzyme activity (potential EEA)
Soil potential EEA (Vmax and Km) analyses in the root-affected soil were performed using fluorogenically labelled substrates (Marx et al. 2005; German et al. 2011) based on 4-methylumbelliferone – MUF: 4-MUF-D-glucoside for β-glucosidase (BG), 4-MUF-phosphate for acid phosphatase (AP) and 4-MUF-N-Acetyl-ß-D-glucosaminide for chitinase (NG). Resuspensions of root-affected soil (50 mL) were prepared using low energy sonication (40 J s−1 output energy) for 2 min. Thereafter, 50 μL of soil suspension, 100 μL substrate solution (2.5, 5, 10, 20, 50, 100 μM) and 50 μL of buffer (MES) were transferred into a 96-well microplate (Tian et al. 2020). Fluorescence was measured at 30, 60 and 120 min at 360 nm excitation and 465 nm emission wave-length and at a slit width of 35 nm with a plate reader (TECAN Infinite F200 Pro). Calibration curves were included in every series of enzyme measurements. Enzyme activities were expressed as MUF release in nM g−1 dry soil h−1. Michaelis-Menten equation was used to determine enzyme kinetic parameters Vmax, and Km (Eq. 1)
Where v is the rate of enzyme-mediated reaction, S is the substrate concentration and Km is an affinity constant equal to the substrate concentration at half of the maximum reaction rate Vmax.
Statistical analysis
Amplicon sequencing data, ASV or OTU species richness and the Shannon diversity index (Shannon et al. 1948) were evaluated using rarefied reads of the 16S rRNA gene fragments (4121), ITS regions (4993) and relative abundance reads of the 18S rRNA gene fragments. Microbial abundances (copy numbers of 16S rRNA gene and ITS region fragments) and soil potential EEA were fitted into bacterial, archaeal, fungal and cercozoan beta-diversities as environmental variables applying Redundancy Analysis (RDA) using scaling = 1 (vegan package, Oksanen et al. 2020). Permutations at 999 for the RDA under a reduced model were performed to examine an overall model fit relative to permuted matrix of data and to check effects of the individual environmental variable included in the model. The Permutational Analysis of Variance (PERMANOVA) test was applied to reveal effects of Texture, Depth and Root Hair on microbial beta-diversity (McArdle and Anderson 2001). The relative abundance of each phylum, ASV or OTU was calculated using non-rarefied data (avoiding exclusion of low read samples), and log transformed for all datasets 16S rRNA gene, ITS region and 18S rRNA gene fragments for statistical tests.
The 30 ASVs or OTUs with highest relative abundances (bacteria, archaea, fungi and cercozoa) were used to generate heatmaps (presenting the average relative abundance ASVs or OTUs per sample). Further, the Spearman’s rank correlation coefficient analysis was performed between the relative abundances of those 30 ASVs or OTUs and the soil potential EEA (BG, AP or NG), applying p value correction based on Benjamini and Hochberg (1995).
The Levene-Test was applied to check the variance homogeneity of the data. The three-way (Texture x Depth x Root Hair) ANOVA was performed to test effects of Texture, Depth and Root Hair on microbial assembly and potential EEA in the rhizosphere and root-affected soil of maize, respectively at p < 0.05. The two-way ANOVA (Depth x Root Hair) was applied per Texture (loam or sand). When the two-way ANOVA indicated significant differences between treatments, then multiple comparisons were followed in loam or sand using Tukey-Test, applying p value correction (at p < 0.05) based on Benjamini and Hochberg (1995).
All data analyses were performed using statistical software R-4.1.0 (R Core Team 2020).
Results
Microbial abundance and diversity in the rhizosphere of maize affected by Texture, Depth and Root Hair
Copy numbers of 16S rRNA gene determined by qPCR were more than three orders of magnitude higher than the copies of ITS fragments (Table S1). Three-way ANOVA of 16S rRNA gene copy numbers revealed a significant effect of Texture and Depth, while the ITS copy numbers did not differ significantly (Tables S1 & S2). The effects of interactions of Texture:Depth and Texture:Root Hair on 16S rRNA gene and ITS fragment copy numbers were significant, but they differed between loam and sand (Table S2).
After sequence processing, the reads clustered into 28,590 bacterial and archaeal, 2694 fungal ASVs and 409 cercozoan OTUs.
Texture significantly affected bacterial and archaeal ASV richness and Shannon indices, with a higher diversity in loam than in sand as shown in the box-plots and by three-way ANOVA (Table 1; Fig. 2A).
Alpha-diversity of A) bacterial and archaeal (16S rRNA gene fragments), B) fungal (ITS regions) and C) cercozoan (18S rRNA gene fragments) communities in the rhizosphere of maize. Letters indicate significant differences between treatments in loam or sand, Tukey test applying “BH” p value correction, p < 0.05 (Benjamini and Hochberg 1995), n = 6, except for L_rth3_D3, S_WT_D1 and S_rth3_D3 (n = 4) and S_WT_D3 (n = 5) for bacteria and archaea. For fungi, n = 6, except for S_WT_D2 and S_WT_D3 (n = 5), S_rth3_D3 (n = 4) and S_rth3_D2 (n = 3). For cercozoa, n = 6, except for S_WT_D3 (n = 5) and S_rth3_D3 (n = 3). L, loam and S, sand
Depth showed a significant effect on bacterial and archaeal Shannon indices, dependent on Texture (a significant interaction of Texture:Depth, Table 1). The effect of Root Hair on ASV richness and Shannon indices was dependent on Depth (a significant interaction of Depth:Root Hair, Table 1). Pairwise comparisons revealed that Shannon index in the rth3 rhizosphere was lower at D3 compared to D1 in both loam and sand (Fig. 2A). This pattern was not observed for the WT rhizosphere. Further, significantly less ASVs and a lower Shannon index were recorded in loam at D3 in rth3 compared to WT rhizosphere (Fig. 2A).
Fungal ASV richness and Shannon index were not affected by Texture, Depth and Root Hair (Table 1; Fig. 2B).
Similar to bacteria and archaea, cercozoan OTU richness and Shannon index were significantly affected by Texture (higher in loam than in sand), Depth and Root Hair (Table 1; Fig. 2C). The effect by Root Hair on OTU richness and Shannon index of cercozoa depended on Texture (a significant interaction of Texture:Root Hair, Table 1). At D2, the Shannon index of cercozoa was significantly lower in rth3 than in WT rhizosphere in sand (Fig. 2C).
The RDA showed that bacterial, archaeal, fungal and cercozoan beta-diversities differed between Texture (Fig. 3A, B & C) and Depth (Figs. S2, S3 & S4), and the significance was confirmed by the PERMANOVA (Table 2). The significant effect of Depth and Root Hair on bacterial, archaeal, fungal and cercozoan beta-diversity was dependent on Texture (significant interactions of Texture:Depth and Texture:Root Hair, Table 2).
Linkage the beta-diversity of A) bacteria and archaea (16S rRNA gene fragments), B) fungi (ITS regions) and C) cercozoa (18S rRNA gene fragments) to soil potential microbial extracellular enzyme activities (EEA) and to copy numbers of 16S rRNA gene fragments and ITS regions. n = 6, except for L_rth3_D3, S_WT_D1 and S_rth3_D3 (n = 4) and S_WT_D3 (n = 5) for bacteria and archaea. For fungi, n = 6, except for S_WT_D2 and S_WT_D3 (n = 5), S_rth3_D3 (n = 4) and S_rth3_D2 (n = 3). For cercozoa, n = 6, except for S_WT_D3 (n = 5) and S_rth3_D3 (n = 3). For soil potential EEA, n = 6, except for L_rth3_D3 (BG, AP and NG), L_WT_D3 (AP), S_rth3_D1 (BG and AP) and S_WT_D1/D2 (AP), n = 5; L_WT_D3 (NG), S_rth3_D2 (BG and AP) and S_WT_D3 (NG), n = 4; and S_rth3_D3 (BG) and S_WT_D3 (BG), n = 3. BG, ß-glucosidase; AP, acid phosphatase; NG, chitinase; L, loam; S, sand and arrows indicate association of the microbial beta-diversity to Km (in green) and to Vmax (in black). Significant association between the soil potential EEA and microbial beta-diversity: <0.0010 ‘***’, < 0.01 ‘**’ and < 0.05 ‘*’
Depth explained a higher variation in bacterial, archaeal, fungal and cercozoan beta-diversities in sand than in loam (PERMANOVA, Table 3). For bacteria and archaea, the greatest differences were observed between D1 vs. D3 for both rth3 and WT grown in loam or in sand (Table 4). Significant differences for fungal beta-diversity between D1 vs. D2 and D1 vs. D3 were only detected for rth3 grown in sand (Table 4). Further, for WT grown in loam significant differences of the fungal beta-diversity were found between D1 vs. D3. For cercozoa, the greatest differences in beta-diversity were shown between D2 vs. D3 in the rhizosphere of WT (in loam) and rth3 (in loam or sand) (Table 4).
Overall, the PERMANOVA tests revealed the Texture as the strongest driver, followed by Depth and Root Hair on bacterial, archaeal, fungal and cercozoan assembly (Table 2).
Relative abundance of bacterial, archaeal, fungal and cercozoan taxa affected by Texture, Depth and Root Hair
Texture had the greatest impact on bacterial, archaeal, fungal and cercozoan alpha- and beta-diversities; therefore, multiple comparisons were performed separately in loam or sand, to check the effects of Depth and Root Hair on relative abundances at different taxonomic levels (using Tukey test, with ‘BH’ p value correction at p < 0.05).
Bacterial and archaeal ASVs were affiliated to 35 phyla but 25 of them had a relative abundance below 1%, and thus they were grouped together and assigned as “Other”. The phylum Proteobacteria was most dominant in sand, while in loam the dominant phylum was Firmicutes (Fig. 4A). Acidobacteria and Thaumarchaeota were far more abundant in loam than in sand (Fig. 4A).
Effects of Texture, Depth and Root Hair on relative abundances of A) bacterial and archaeal (16S rRNA gene fragments), B) fungal (ITS regions) and C) cercozoan (18S rRNA gene fragments) communities at phylum or order levels in the rhizosphere of maize (> 1%). Letters indicate significant differences of each bacterial, archaeal, fungal and cercozoan phylum or order level in loam or sand, Tukey test applying “BH” p value correction, p < 0.05 (Benjamini and Hochberg 1995). n = 6, except for L_rth3_D3, S_WT_D1 and S_rth3_D3 (n = 4) and S_WT_D3 (n = 5) for bacteria and archaea; S_WT_D2 and S_WT_D3 (n = 5), S_rth3_D3 (n = 4) and S_rth3_D2 (n = 3) for fungi. n = 6 for cercozoa, except for S_WT_D3 (n = 5) and S_rth3_D3 (n = 3). L, loam and S, sand
Changes in relative abundances of bacteria and archaea at the phylum level along examined depths were also highly modulated by Texture. For instance, the relative abundance of Proteobacteria was significantly lower at D1 than D3 in the rhizosphere of both WT and rth3 in sand, and the opposite was observed in loam (Fig. 4A). The effects of Depth on Thaumarchaeota were only observed in the rhizosphere of maize grown in loam (Fig. 4A). Overall, Root Hair had a subtle effect on the relative abundances at the phylum level (Fig. 4A).
For fungi, all ASVs were affiliated to eight phyla (four of them were grouped together, and renamed as “Other” due to a low relative abundance ≤1%). The fungal phyla Ascomycota and Basidiomycota were dominant in loam or sand (Fig. 4B). Effects of Depth on fungal relative abundances at phylum level were only observed for Mortierellomycota that displayed significantly lower relative abundances at D3 than D1 for WT grown in loam (Fig. 4B). Overall, no effect of Root Hair on fungal relative abundances was detected at the phylum level in the rhizosphere of maize grown in loam or sand (Fig. 4B).
Cercozoan OTUs were affiliated to 19 orders, 11 of them were grouped together and re-named “Other” as each represented less than 1% of the total number of reads. The orders Glissomonadida and Cercomonadida dominated in the rhizosphere of maize grown in loam or sand and at all three depths (Fig. 4C). In loam, no clear effect of Depth or Root Hair was detected, while in sand an increase in the relative abundance of the order Glissomonadida was found with increasing depth, but only in the rth3 rhizosphere. Cryomonadida were mainly detected in sand, and their relative abundances were higher at D1 than at D2 or D3 in WT and rth3 rhizosphere (Fig. 4C).
To increase the level of resolution effects of Texture, Depth and Root Hair on the rhizosphere microbiome were also analysed at the ASV or OTU level.
The 30 bacterial or archaeal ASVs with the highest relative abundance can be depicted from Fig. 5A. ASVs belonging to archaeal family Nitrososphaeraceae (ASV_2 &_10) and the bacterial genera Terrimonas (ASV_19) and Bacillus (ASV_11, _12 & _13) were dominant in loam, while the ASVs affiliated to bacterial genera Dyella (ASV_41 & _45), Massilia (ASV_1, _4 & _14) and Streptomyces (ASV_30) were more abundant in sand (Fig. 5A). The relative abundance of Bacillus (ASV_13) was higher at D3 than D1 for the rth3 grown in loam (at p < 0.05). In sand, the relative abundance of Massilia (ASV_1 & _14) was significantly higher at D3 than D1 for the WT or rth3 rhizosphere. In contrast, the relative abundance of ASV_30 belonging to Streptomyces was significantly higher at D1 than D2 or D3 for the WT or rth3 rhizosphere in sand. Root Hair affected the distribution of several bacterial ASVs in the rhizosphere dependent on Texture and Depth. For instance, ASVs affiliated to bacterial Dyella (ASV_41 & _45) in the rhizosphere of rth3 grown in sand had significantly higher relative abundance at D1 than in the WT rhizosphere (Fig. 5A). It is important to note that not all ASVs affiliated to a certain genus or family showed identical Depth and Root Hair-dependent relative abundances.
Effects of Texture, Depth and Root Hair on 30 bacterial and archaeal (16S rRNA gene fragments) ASVs with highest relative abundances in the rhizosphere of maize (A) and their Spearman's rank correlation coefficient (r) to soil potential microbial extracellular enzyme activities (B). Different letters indicate significant differences between treatments of each ASV in loam or sand, Tukey test applying “BH” p value correction, p < 0.05 (Benjamini and Hochberg 1995). BG, ß-glucosidase; AP, acid phosphatase; NG, chitinase; V, Vmax and K, Km; Blank, no significant correlation and * significant correlation. n = 6, except for L_rth3_D3, S_WT_D1 and S_rth3_D3 (n = 4) and S_WT_D3 (n = 5). L, loam and S, sand
The majority of the 30 most abundant fungal ASVs did not show Texture, Depth and Root Hair-dependent differences in their relative abundances (Fig. 6A). But ASV_5 affiliated to the genus Trichophaea and the ASV_7 (unclassified Ascomycota) were dominant in loam and in sand, respectively.
Effects of Texture, Depth and Root Hair on 30 fungal (ITS regions) ASVs with highest relative abundances in the rhizosphere of maize (A) and their Spearman's rank correlation coefficient (r) to soil potential microbial extracellular enzyme activities (B). Different letters indicate significant differences between treatments of each ASV in loam or sand, Tukey test applying “BH” p value correction, p < 0.05 (Benjamini and Hochberg 1995). BG, ß-glucosidase; AP, acid phosphatase; NG, chitinase; V, Vmax and K, Km; Blank, no significant correlation and * significant correlation. n = 6, except for S_WT_D2 and S_WT_D3 (n = 5), S_rth3_D3 (n = 4) and S_rth3_D2 (n = 3). L, loam and S, sand
This similar pattern was also revealed for the 30 most abundant cercozoan OTUs (Fig. 7A). Sequences belonging to cercozoan families Allapsidae (OTU_15) and Sandonidae (OTU_8) were dominant in loam, while two other unclassified cercozoan OTUs again of the families Allapsidae (OTU_4) and Sandonidae (OTU_11) were more abundant in sand (Fig. 7A). The relative abundance of OTUs of the families Sandonidae (OTU_11) and Allapsida (OTU_4) was highest at D2 and D3 rhizosphere, respectively, for both WT or rth3 grown in sand. In loam, the OTU assigned to the Allapsidae (OTU_15) showed the lowest relative abundance at D2 for both WT and rth3 rhizosphere.
Effects of Texture, Depth and Root Hair on 30 cercozoan (18S rRNA gene fragments) OTUs with highest relative abundances in the rhizosphere of maize (A) and their Spearman's rank correlation coefficient (r) to soil potential microbial extracellular enzyme activities (B). Different letters indicate significant differences of each OUT in loam or sand, Tukey test applying “BH” p value correction, p < 0.05 (Benjamini and Hochberg 1995). BG, ß-glucosidase; AP, acid phosphatase; NG, chitinase; V, Vmax and K, Km; Blank, no significant correlation and * significant correlation. n = 6, except for S_WT_D3 (n = 5) and S_rth3_D3 (n = 3). L, loam and S, sand
Soil potential extracellular enzyme activities (EEA) affected by Texture, Depth and Root Hair
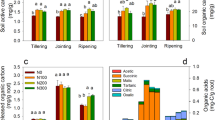
In general, the soil potential EEA (Vmax/ V) were approximately 2.5–7 times significantly higher in loam than sand (Table 5), and the significant effect of Texture was confirmed by three-way ANOVA (Table S3). Effects of Depth on the soil potential EEA were Texture-dependent, and a significant interaction of Texture:Depth was indicated (Table S3). The β-glucosidase activity (BG_V) was not affected by Depth in loam or in sand. Acid phosphatase (AP_V) showed lower activity at D3 compared to D1 or D2 for rth3 and WT in loam, but the opposite pattern was observed in sand (Table 5). Chitinase (NG_V) had a higher enzyme activity at D1 than D2 for rth3 (loam) and at D1 than D2 and D3 for WT (loam). Root Hair affected only the BG_V at D1 in sand as WT had a more than two-fold activity higher than rth3.
The soil potential EEA affinity (Km/ K) BG_K showed different enzyme affinities at D2 vs. D3 in loam of WT (Table 5). AP_K at D3 in loam and at D2 in sand were significantly higher than other depths of the rth3. NG_K indicated a similar enzyme affinity at all depths. Overall, enzyme kinetics also revealed Texture as the strongest driver, followed by Depth and Root Hair.
Data of soil potential EEA (Vmax and Km) were incorporated into the microbial beta-diversity in RDA plots to examine their linkage to the bacteria and archaea (Fig. 3A), fungi (Fig. 3B) and cercozoa (Fig. 3C) of both textures and to increase the resolution per Texture (Figs. S2, S3 & S4). The fitted vectors showed that bacterial and archaeal communities had significantly high activities of BG_V and AP_V in loam, while in sand significantly high enzyme affinity BG_K was observed (Fig. 3A). Spearman’s rank correlation coefficients were calculated to discover potential interactions between the relative abundances of the 30 dominant bacterial and archaeal ASVs and the soil potential EEA (Fig. 5B). We found that several bacterial ASVs affiliated to Flavisolibacter (ASV_32) in loam and Paenibacillus (ASV_47) and Massilia (ASV_1, _4 & _14) in sand were significantly and positively correlated to AP_V.
The fungal communities had higher AP_V, BG_V, NG_V and NG_K in loam than in sand but differences were not significant (Fig. 3B). Spearman’s rank correlation coefficients indicated several ASVs were positively correlated to AP_V, BG_V and NG_V, but they were not significant (Fig. 6B).
The cercozoan communities had significantly higher NG_V in loam than in sand (Fig. 3C). The relative abundance of OTUs affiliated to Allapsidae_OTU004 and Eocercomonas_OTU006 was significantly and positively correlated to AP_V, while the Paracercomonas_OTU007 was correlated to activity and affinity of NG_V and NG_K, respectively in sand (Fig. 7B).
Overall, the copy numbers of 16S rRNA gene and ITS region fragments were strongly linked with AP_V (Fig. 3A, B & 3C).
The RDA plot including the soil potential EEA and microbial communities per Texture (Figs. S2, S3 & S4) revealed that the bacterial, archaeal, fungal and cercozoan communities had higher AP_V at D2/D3 than D1 in sand. While in loam, the NG_V was higher at D1 for bacterial, archaeal and cercozoan communities.
Discussion
Effects of Texture on the maize rhizosphere microbiome
Substrates with different textures sand and loam used for growing maize in the present work differed per se in pore size distribution, and maize root-induced changes in porosity in the rhizosphere (Phalempin et al. 2021). Due to higher organic matter, silt and clay contents, loam has higher sorption sites than sand causing lower nutrient mobility and leaching substances in loam (Vetterlein et al. 2021). The dilution of loam with 83.3% quartz sand (for Texture sand) indeed resulted in a decrease in abundance (copy numbers of 16S rRNA gene fragments) and alpha-diversity (observed ASVs/OTUs and Shannon indices) of bacteria, archaea and cercozoa (Tables 1, S1 & S2; Fig. 2). Higher pore connectivity of sand was recently already reported in different studies to decrease bacterial diversity in soil due to easier migration of microbial cells between pores (Carson et al. 2010; Hemkemeyer et al. 2018; Seaton et al. 2020).
In the present study, changes in cercozoan richness and diversity displayed a similar pattern as those observed for bacteria and archaea (Table 1; Fig. 2). This is likely due to feeding traits of cercozoa. Most cercozoan taxa mainly feed on bacteria (Fiore-Donno et al. 2019), and these bacterivorous cercozoan taxa affect bacteria and archaea in soil due to their specificity in feeding patterns (Kreuzer et al. 2006; Rosenberg et al. 2009; Flues et al. 2017; Henkes et al. 2018). Subsequently, cercozoa are bottom up controlled by bacteria, as the cercozoan community assembly is affected by bacterial defense mechanisms (Jousset 2012), and their growth depends on the availability of their major food source. In addition, the differences in Texture of the two soils might have changed these predator-prey interactions. Loam with a higher content of fine soil pores, might have restricted the access of protists to their bacterial prey (Rutherford and Juma 1992).
Compared to bacteria, archaea and cercozoa, Texture revealed less effects on the fungal abundance (copy numbers of ITS regions, Tables S1 & S2), alpha- (Table 1) and beta-diversity (Table 2) as previously observed for other soils (Hartmann et al. 2014; Yim et al. 2017; Seaton et al. 2020). Fungal hyphae can easily bridge pore spaces (Ritz and Young 2004), while single cell organisms depend on the connectivity of the water film for dispersal.
Findings using maize roots obtained from the three depth layers of the soil column experiments of the present study, Ganther et al. (2021) found a higher expression level of genes for aquaporins involved in passive water transport, genes related to plant immunity function, e.g., gene related to pathogenesis-related protein 5 (PR5), chitinase, ethylene-intensive 3 transcription factor and genes related to secondary metabolite production in sand compared to loam. Genes coding for aquaporin (Marulanda et al. 2010), chitinases (Shoresh and Harman 2010), PR5 (Anisimova et al. 2021) and metabolites (Cotton et al. 2019; Murphy et al. 2021) that were differentially expressed between loam and sand, were previously shown to be involved in plant microbe interactions, and might be influenced by the Texture-dependent microbiome composition.
Effects of Depth on the maize rhizosphere microbiome
In an identical experiment, we also learned that share of young roots increased with increasing rooting depth (Lippold et al. 2021; Fig. 1). Young or fine roots play active roles in respiration, transport and absorption of water and nutrients, and they also have much more carbon release as exudates for microbial cells compared to the older roots (Nikolova et al. 2020). Thus, they might favor r-strategists (King et al. 2021; Wei et al. 2021) as observed in the present study for Massilia and Bacillus. The assembly of bacterial, archaeal and cercozoan communities in different root zones, i.e., distance to root tips of the primary root of nine-day-old WT maize plants grown in loam, was previously investigated by Rüger et al. (2021). They reported that along the primary root axis, the alpha-diversity was higher at root tips compared to older root regions. In our study, a mixture of roots per depth layer was used for obtaining the rhizosphere microbial pellet, but as the proportion of young roots increased with depth (Lippold et al. 2021), we observed a slightly increased alpha-diversity of bacteria and archaea in the rhizosphere of 22-day-old WT maize grown in loam at the D3 compared to D1 (Fig. 2A), confirming the findings by Rüger et al. (2021).
The soil water content showed a depth gradient due to gravity in the soil column (Vetterlein et al. 2021). As plants developed, differences in root length density with depth evolved, and might have introduced additional gradients in nutrient distribution. Therefore, differences in water, O2 availability and nutrient resources contributed to differences in microbial abundance and diversity at the three depth levels analysed as already reported in earlier studies (Schlüter et al. 2019; Schimel et al. 1999; Schlatter et al. 2018; Li et al. 2020). Further, microbe-microbe interactions in terms of competition and facilitation of niche occupancy (Sasse et al. 2018; Cotton et al. 2019), top-down control by microbial predators, i.e., protists (Bonkowski et al. 2021) - these factors also cause heterogeneity along rooting depth, and thus shape the rhizosphere microbiome.
Larger differences in bacterial and archaeal beta-diversity were observed between D1 vs. D3 than between D1 vs. D2 or D2 vs. D3 (Table 4), and similar findings were reported for plant root gene expression analysis using roots from the same experiment and depths (Ganther et al. 2021). Depth affected also the structure of acdS gene carrying bacteria in TC-DNAs of the same rhizosphere samples investigated in the present study (Gebauer et al. 2021).
Depth-related effects on fungal gene copy numbers and alpha-diversity were not observed in the present study (at 4.5–15.1 cm depth) likely due to relatively short soil columns being used.
Effects of Root Hair on the maize rhizosphere microbiome
Lack of Root Hair had little effects on microbial assembly compared to Texture and Depth in the present study, but was significant for bacteria, archaea and cercozoa (Table 2). Only in loam Root Hair had also an effect on fungal beta-diversity. This suggests that there might be a link to the findings by Lippold et al. (2021) that total P uptake was significantly lower for rth3 than WT in loam.
The small effect of Root Hair was in line with root gene expression (Ganther et al. 2021) and with bacterial community carrying acdS gene (Gebauer et al. 2021) observed for the same experiment.
Root hairs of the WT used in this study were relatively short, of a length of about 0.24 mm (Lippold et al. 2021; Phalempin et al. 2021). This resulted in small differences between the WT and rth3 in the rhizosphere zone and in their role in water and nutrient uptake. This might also explain the rather small effects on rhizosphere microbiome observed. Further, the subtle effects of Root Hair in microbial assembly might be attributed to fertilization of the soils and to well irrigation (Vetterlein et al. 2021).
In a previous work, the root hairless rth2 maize accumulated less biomass and P than WT plants, but only under water stress (Klamer et al. 2019), suggesting that testing the genotypes under different P and soil moisture levels could reveal stronger effects of Root Hair on the rhizosphere microbiome. Further, the effects of Root Hair on the rhizosphere microbiome might be larger for other plant species showing longer root hairs, i.e., for barley and growing under water deficiency or drought (Marin et al. 2021).
Relative abundance of microbial taxa affected by Texture, Depth and Root Hair
The dominant phyla in loam were Firmicutes followed by Proteobacteria (Fig. 4) as previously also reported by Ganther et al. (2020), for the same Texture and WT maize, using the same soil column experimental set-up to investigate the effects of X-ray computer tomography on soil bacterial communities. In contrast to loam, the relative abundance of Proteobacteria was dominant in the rhizosphere of WT and rth3 grown in sand. In particular, the genus Massilia was remarkably dominant in the Texture sand that was obtained by mixing loam with quartz sand. Texture-dependent changes in root exudation patterns or the soil physicochemical characteristics might have facilitated the successful rhizosphere assembly of the three ASVs affiliated to Massilia. Acidobacteria and also Thaumarchaeota were mainly detected in loam indicating a preference of these phyla to the soil conditions in loam, e.g., particle size preferences as recently reported by Hemkemeyer et al. (2018).
Wei et al. (2021) revealed that most of the enriched taxa at younger roots or at a lower depth were r-strategists, they proliferated, and responded quickly to nutrients available. Indeed, our findings revealed significantly higher relative abundances of ASVs affiliated to the genera Bacillus (in loam) and Massilia (in sand) in the rhizosphere of roots sampled from D3 compared to D1. Isolates affiliated to these genera are also known as r-strategists (Ofek et al. 2012; Wei et al. 2021). Further, strains from both genera were reported to have catalase activity (Yuan et al. 2021), and thus their increased abundance in the maize rhizosphere might reduce H2O2 concentration, and thus reduce H2O2 stress. Root gene expression analyses of maize showed that stress- or defense-related genes such as peroxidases were highly expressed at lower than upper depths (Ganther et al. 2021).
Slight decreases in oxygen availability, which coincide with the increase in water content with depth, might explain differences in relative abundances of ASVs affiliated to Bacillus. Bacillus isolates can adapt to a lower oxygen availability (Hartmann et al. 2014) that might support the present finding that their relative abundances were enriched at D3 in loam (Fig. 5A).
The fungal relative abundances were not affected by Depth and Root Hair at the phylum level and most of the 30 ASVs observed in the present work (Figs. 4B & 6A). This was likely due to our observation using a short soil column, and was under growth chamber conditions.
ASVs affiliated to the genus Dyella that were detected in significantly higher relative abundances in the rhizosphere of rth3 than WT grown in sand (Fig. 5A), are known to be involved in nitrogen fixation (Swarnalakshmi et al. 2020). Based on plant roots and rhizosphere microbiome feedback, strains belonging to Dyella might have been recruited to assist the rth3 plants for N-uptake in sand. Interestingly, root genes coding for nitrate transport were shown to be less expressed in the rth3 compared to WT (Ganther et al. 2021).
Linkage between bacteria, archaea, fungi, cercozoa and soil potential extracellular enzyme activities (EEA)
Both plant roots and microbial cells are able to release enzymes into soil (Cabugao et al. 2017) but we assume that the fraction of plant-originated enzymes decreases with the distance from the root, and as we used root-affected soil for the EEA analyses, the detected potential EEA were assumed to be of microbial origin.
In line with the bacterial and archaeal abundance and alpha-diversity, higher potential EEA (Vmax) were recorded in loam than in sand (Tables 5 & S1). This is explained by a greater soil organic carbon content in loam (Vetterlein et al. 2021; Feng et al. 2019; Ren et al. 2018).
The Km/K in soil studies is generally termed as an apparent affinity constant, which is surely affected to a certain extent by soil physical properties. The two-to-three-fold higher Km for acid phosphatase (AP_K) in sand compared to loam (Table 5), indicated the presence of different enzyme systems with a lower affinity resulting in a decline in the overall enzyme function under substrate limitation in loam (Stone et al. 2012; German et al. 2012). Further, as discussed above, due to the higher sorption capacity of loam, the affinity of AP_K was strongly reduced compared to sand (Table 5).
Variations in Km values at different depths were more pronounced for the AP, followed by NG (chitinase) and BG (ß-glucosidase). They revealed changes in functional traits of microorganisms, i.e., changes in the metabolic activity and microbial community composition (Blagodatskaya et al. 2021).
The enzyme activity and affinity were fitted as environmental variables of the bacterial, archaeal, fungal and cercozoan communities (Fig. 3). Higher activities of BG_V, AP_V and NG_V were shown in loam than in sand demonstrating different abilities of dominant bacteria, archaea, fungi or cercozoa to produce or release the enzymes investigated. The correlation of AP_V with multiple ASVs or OTUs indicated functional redundancy and possible mutualistic interactions in P acquisition and in cellulose degradation within the communities (Banerjee et al. 2016). Positive correlation of AP_V with ASVs affiliated to Massilia and the cercozoa Allapsidae and Eocercomonas in sand possibly indicate different specialized taxa in production of specific enzymes. The bacterial, archaeal and cercozoan communities showed significantly higher AP_V at lower depths in sand (Figs. S2 & S4), which is in line with higher relative abundances of Massilia, Allapsidae and Eocercomonas (Figs. 5A & 7A).
A higher enzyme affinity in sand (Fig. 3) to the decomposed compounds indicated different taxa produced distinctly different enzyme systems (Fontaine and Barot 2005; Blagodatskaya et al. 2009; Blagodatskaya and Kuzyakov 2013). The stronger linkage of enzyme parameters with bacteria and archaea than fungi observed in the present study might be due to the lower abundance of fungi but possibly indicated a broader functional diversity within bacterial and archaeal communities that are able to decompose plant and microbial residues as well as to hydrolyze organic P compounds, while fungi invested more resources in P acquisition (Smith et al. 2011; Chiba et al. 2021). Although the potential EAA were analysed in soil taken at a larger distance from root surface, we did find the correlation to the microbiome in the rhizosphere (Figs. 5, 6 & 7).
Conclusion
Our research on the relationship between Texture, Depth and Root Hair has three general implications. First, we have shown that Texture was the strongest driver of the rhizosphere microbial assembly and of potential EEA for both WT and rth3 plants. Second, we have demonstrated that Depth was another driver of the rhizosphere microbiome, suggesting that the abiotic environment may differ between the different layers of the column. Third, the small impact of Root Hair on the rhizosphere microbiome and potential EEA in root-affected soil of maize raises the question of the importance of the root hair length and the low plasticity of root hair defective mutant related to limited P availability as reported by Lippold et al. (2021).
Overall, our hypotheses were only partly confirmed which might be due to the resolution level of amplicon sequence analyses, but also due to the experimental design of the column experiment.
Most excitingly, the results of the present study as well as the previously published data on the plant side, e.g., plant gene expression and root system architecture from same and identical column experiments (Ganther et al. 2021; Lippold et al. 2021) showed the same drivers, and highlighted the close linkage between the plant and their rhizosphere microbiome and potential EEA.
Data availability
All raw sequences for bacteria, archaea and fungi were deposited at NCBI within the sequence read archive accession PRJNA677863. Raw sequences of cercozoa are available at the ENA under the accession number PRJEB49274.
References
Anisimova OK, Shchennikova AV, Kochieva EZ, Filyushin MA (2021) Pathogenesis-related genes of PR1, PR2, PR4 and PR5 families are involved in the response to fusarium infection in garlic (Allium sativum L.). Int J Mol Sci 22:6688. https://doi.org/10.3390/ijms22136688
Banerjee S, Kirkby CA, Schmutter D et al (2016) Network analysis reveals functional redundancy and keystone taxa amongst bacterial and fungal communities during organic matter decomposition in an arable soil. Soil Biol Biochem 97. https://doi.org/10.1016/j.soilbio.2016.03.017
Beirinckx S, Viaene T, Haegeman A et al (2020) Tapping into the maize root microbiome to identify bacteria that promote growth under chilling conditions. Microbiome 8:54. https://doi.org/10.1186/s40168-020-00833-w
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Series B 57:289–300. https://doi.org/10.1111/j.2517-6161.1995.tb02031.x
Berg G, Köberl M, Rybakova D et al (2017) Plant microbial diversity is suggested as the key to future biocontrol and health trends. FEMS Microbiol Ecol 93:5. https://doi.org/10.1093/femsec/fix050
Blagodatskaya E, Kuzyakov Y (2013) Active microorganisms in soil: critical review of estimation criteria and approaches. Soil Biol Biochem 67:192–211. https://doi.org/10.1016/j.soilbio.2013.08.024
Blagodatskaya EV, Blagodatsky SA, Anderson TH, Kuzyakov Y (2009) Contrasting effects of glucose, living roots and maize straw on microbial growth kinetics and substrate availability in soil. Eur J Soil Sci 60:186–197. https://doi.org/10.1111/j.1365-2389.2008.01103.x
Blagodatskaya E, Tarkka M, Knief C et al (2021) Bridging microbial functional traits with localized process rates at soil interfaces. F Microbiol 12:625697
Bolyen E, Rideout JR, Dillon et al (2018) QIIME 2: reproducible, interactive, scalable and extensible microbiome data science. Non Peer-Rev. https://doi.org/10.7287/peerj.preprints.27295v1
Bonkowski M, Tarkka M, Razavi BS et al (2021) Spatiotemporal dynamics of maize (Zea mays L.) root growth and its potential consequences for the assembly of the rhizosphere microbiota. F Microbiol 12:619499. https://doi.org/10.3389/fmicb.2021.619499
Brisson VL, Schmidt JE, Northen TR et al (2019) Impacts of maize domestication and breeding on rhizosphere microbial community recruitment from a nutrient depleted agricultural soil. Sci Rep 9:15611. https://doi.org/10.1038/s41598-019-52148-y
Burak E, Quinton JN, Dodd IC (2021) Root hairs are the most important root trait for rhizosheath formation of barley (Hordeum vulgare), maize (Zea mays) and Lotus japonicus (Gifu). Ann Bot 128(1):45–57. https://doi.org/10.1093/aob/mcab029
Cabugao KG, Timm CM, Carrell AA et al (2017) Root and rhizosphere bacterial phosphatase activity varies with tree species and soil phosphorus availability in Puerto Rico tropical forest. F Plant Sci 8:1834. https://doi.org/10.3389/fpls.2017.01834
Callahan BJ, McMurdie PJ, Rosen MJ et al (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583
Camacho C, Coulouris G, Avagyan V et al (2009) BLAST+: architecture and applications. BMC Bioinform 10:421. https://doi.org/10.1186/1471-2105-10-421
Canarini A, Kaiser C, Merchant A et al (2019) Root exudation of primary metabolites: mechanisms and their roles in plant responses to environmental stimuli. F Plant Sci 10:157. https://doi.org/10.3389/fpls.2019.00157
Carson JK, Gonzalez-Quiñones V, Murphy DV et al (2010) Low pore connectivity increases bacterial diversity in soil. Appl Environ Microbiol 76:3936–3942
Chiba A, Uchida Y, Kublik S et al (2021) Soil bacterial diversity is positively correlated with decomposition rates during early phases of maize litter decomposition. Microorganisms 9(2):357. https://doi.org/10.3390/microorganisms9020357
Cotton TEA, Pétriacq P, Cameron DD et al (2019) Metabolic regulation of the maize rhizobiome by benzoxazinoids. ISME J 13:1647–1658. https://doi.org/10.1038/s41396-019-0375-2
DeAngelis KM, Brodie EL, DeSantis TZ et al (2009) Selective progressive response of soil microbial community to wild oat roots. ISME J 3:168–178
Dick RP (1997) Soil enzyme activities as integrative indicators of soil health. In: Pankhurst CE, Doube BM, Gupta VVSR (eds) Biological indicators of soil health. CAB Int, Wellingford, pp 121–156
Dumack K, Fiore-Donno AM, Bass D, Bonkowski M (2019) Making sense of environmental sequencing data: ecologically important functional traits of the protistan groups Cercozoa and Endomyxa (Rhizaria). Mol Ecol Resour 20:398–403. https://doi.org/10.1111/1755-0998.13112
Edgar RC, Brian JH, Jose CC et al (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27(16):2194–2200
Ekenler M, Tabatabai MA (2004) β-Glucosaminidase activity as an index of nitrogen mineralization in soils. Commun Soil Sci Plant Anal 35(7–8):1081–1094. https://doi.org/10.1081/CSS-120030588
Feng X, Zhang L, Zhao F et al (2019) Effects of mixing feldspathic sandstone and sand on soil microbial biomass and extracellular enzyme activities - a case study in mu us sandy land in China. Appl Sci 9:3963. https://doi.org/10.3390/app9193963
Fiore-Donno AM, Rixen C, Rippin M et al (2018) New barcoded primers for efficient retrieval of cercozoan sequences in high-throughput environmental diversity surveys, with emphasis on worldwide biological soil crusts. Mol Ecol Resour 18(2):229–239
Fiore-Donno AM, Richter-Heitmann T, Degrune F et al (2019) Functional traits and spatio-temporal structure of a major group of soil protists (Rhizaria: Cercozoa) in a temperate grassland. F Microbiol 10:1–12. https://doi.org/10.3389/fmicb.2019.01332
Fiore-Donno AM, Richter-Heitmann T, Bonkowski M (2020) Contrasting responses of protistan plant parasites and phagotrophs to ecosystems, land management and soil properties. F Microbiol 11:1–33
Flues S, Bass D, Bonkowski M (2017) Grazing of leaf-associated Cercomonads (Protists: Rhizaria: Cercozoa) structures bacterial community composition and function. Environ Microbiol 19(8):3297–3309. https://doi.org/10.1111/1462-2920.13824
Fontaine S, Barot S (2005) Size and functional diversity of microbe populations control plant persistence and long-term soil carbon accumulation. Ecol Lett 8:1075–1087. https://doi.org/10.1111/j.1461-0248.2005.00813.x
Ganther M, Yim B, Ibrahim Z et al (2020) Compatibility of x-ray computed tomography with plant gene expression, rhizosphere bacterial communities and enzyme activities. J Exp Bot. https://doi.org/10.1093/jxb/eraa262
Ganther M, Vetterlein D, Heintz-Buschart A, Tarkka MT (2021) Transcriptome sequencing analysis of maize roots reveals the effects of substrate and root hair formation in a spatial context. Plant Soil. https://doi.org/10.1007/s11104-021-04921-0
Gebauer L, Bouffaud M-L, Ganther M et al (2021) Soil texture, sampling depth and root hairs shape the structure of ACC deaminase bacterial community composition in maize rhizosphere. F Microbiol 12:616828. https://doi.org/10.3389/fmicb.2021.616828
German DP, Weintraub MN, Grandy AS et al (2011) Optimization of hydrolytic and oxidative enzyme methods for ecosystem studies. Soil Biol Biochem 43(7):1387–1397. https://doi.org/10.1016/j.soilbio.2011.03.017
German DP, Marcelo KRB, Stone MM, Allison SD (2012) The Michaelis-Menten kinetics of soil extracellular enzymes in response to temperature: a cross latitudinal study. Glob Chang Biol 18:1468–1479
Gil-Sotres F, Trasar-Cepeda C, Leirós M, Seoane S (2005) Different approaches to evaluating soil quality using biochemical properties. Soil Biol Biochem 37:877–887. https://doi.org/10.1016/j.soilbio.2004.10.003
Glücksman E, Bell T, Griffiths RI, Bass D (2010) Closely related protist strains have different grazing impacts on natural bacterial communities. Environ Microbiol 12:3105–3113. https://doi.org/10.1111/j.1462-2920.2010.02283.x
Gomes EA, Lana UGP, Quensen JF et al (2018) Root-associated microbiome of maize genotypes with contrasting phosphorus use efficiency. Phytobiomes 2:129–137
Gschwendtner S, Reichmann M, Müller et al (2010) Effects of genetically modified amylopectin-accumulating potato plants on the abundance of beneficial and pathogenic microorganisms in the rhizosphere. Plant Soil 335:413–422
Guillou L, Bachar D, Audic S et al (2013) The protist ribosomal reference database (PR2): a catalog of unicellular eukaryote small subunit rRNA sequences with curated taxonomy. Nucleic Acids Res 41:D597–D604. https://doi.org/10.1093/nar/gks1160
Gweon HS, Oliver A, Taylor J et al (2015) PIPITS: an automated pipeline for analyses of fungal internal transcribed spacer sequences from the Illumina sequencing platform. Methods Ecol Evol 6:973–980
Hartmann A, Rothballer M, Schmid M (2008) Lorenz Hiltner, a pioneer in rhizosphere microbial ecology and soil bacteriology research. Plant Soil 312:7–14
Hartmann A, Schmid M, Tuinen DV, Berg G (2009) Plant-driven selection of microbes. Plant Soil 321:235–257
Hartmann M, Niklaus PA, Zimmermann S et al (2014) Resistance and resilience of the forest soil microbiome to logging-associated compaction. ISME J 8:226–244
Hemkemeyer M, Dohrmann AB, Christensen BT, Tebbe CC (2018) Bacterial preferences for specific soil particle size fractions revealed by community analyses. F Microbiol 9:149. https://doi.org/10.3389/fmicb.2018.00149
Henkes GJ, Kandeler E, Marhan S, Scheu S, Bonkowski M (2018) Interactions of mycorrhiza and protists in the rhizosphere systemically alter microbial community composition, plant shoot-to-root ratio and within-root system nitrogen allocation. F Environ Sci 6:117. https://doi.org/10.3389/fenvs.2018.00117
Hochholdinger F, Wen T-J, Zimmermann R et al (2008) The maize (Zea mays L.) root hairless 3 gene encodes a putative GPI-anchored, monocot-specific, COBRA-like protein that significantly affects grain yield. Plant J 54:888–898
Ihrmark K, Bödeker ITM, Cruz-Martinez K et al (2012) New primers to amplify the fungal ITS2 region – evaluation by 454-sequencing of artificial and natural communities. FEMS Microbiol Ecol 82:666–677
Jousset A (2012) Ecological and evolutive implications of bacterial defences against predators. Environ Microbiol 14(8):1830–1843. https://doi.org/10.1111/j.1462-2920.2011.02627.x
Jousset A, Scheu S, Bonkowski M (2008) Secondary metabolite production facilitates establishment of rhizobacteria by reducing both protozoan predation and the competitive effects of indigenous bacteria. Funct Ecol 22:714–719. https://doi.org/10.1111/j.1365-2435.2008.01411.x
Kawasaki A, Donn S, Ryan PR et al (2016) Microbiome and exudates of the root and rhizosphere of Brachypodium distachyon, a model for wheat. PLoS One 11(10):e0164533. https://doi.org/10.1371/journal.pone.0164533
King WL, Yates CF, Guo J et al (2021) The hierarchy of root branching order determines bacterial composition, microbial carrying capacity and microbial filtering. Commun Biol 4:483. https://doi.org/10.1038/s42003-021-01988-4
Klamer F, Vogel F, Li X et al (2019) Estimating the importance of maize root hairs in low phosphorus conditions and under drought. Ann Bot 124:961–968
Kompała-Bąba A, Bierza W, Sierka E et al (2021) The role of plants and soil properties in the enzyme activities of substrates on hard coal mine spoil heaps. Sci Rep 11(1):5155. https://doi.org/10.1038/s41598-021-84673-0
Kong X, Han Z, Tai X et al (2020) Maize (Zea mays L. sp.) varieties significantly influence bacterial and fungal community in bulk soil, rhizosphere soil and phyllosphere. FEMS Microbiol Ecol 96:fiaa020. https://doi.org/10.1093/femsec/fiaa020
Korenblum E, Dong Y, Szymanski J et al (2020) Rhizosphere microbiome mediates systemic root metabolite exudation by root-to-root signalling. PNAS 117(7):3874–3883
Kreuzer K, Adamczyk J, Iijima M et al (2006) Grazing of a common species of soil Protozoa (Acanthamoeba castellanii) affects rhizosphere bacterial community composition and root architecture of rice (Oryza sativa L.). Soil Biol Biochem 38:1665–1672. https://doi.org/10.1016/j.soilbio.2005.11.027
Kusstatscher P, Wicaksono WA, Thenappan DP et al (2020) Microbiome management by biological and chemical treatments in maize is linked to plant health. Microorganisms 8:1506. https://doi.org/10.3390/microorganisms8101506
Leitner D, Klepsch S, Ptashnyk M et al (2010) A dynamic model of nutrient uptake by root hairs. New Phytol 185:792–802
Li X, Rui J, Mao Y, Yannarell A, Mackie R (2014) Dynamics of the bacterial community structure in the rhizosphere of a maize cultivar. Soil Biol Biochem 68:392–401
Li X, Wang H, Li X et al (2020) Distribution characteristics of fungal communities with depth in paddy fields of three soil types in China. J Microbiol 58(4):279–287
Lippold E, Phalempin M, Schlüter S, Vetterlein D (2021) Does the lack of root hairs alter system architecture of Zea mays? Plant Soil 467:267–286
Marin M, Feeney DS, Brown LK et al (2021) Significance of root hairs for plant performance under contrasting field conditions and water deficit. Ann Bot 128:1–16. https://doi.org/10.1093/aob/mcaa181
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17:10–12. https://doi.org/10.14806/ej.17.1.200
Marulanda A, Azcón R, Chaumont F et al (2010) Regulation of plasma membrane aquaporins by inoculation with a Bacillus megaterium strain in maize (Zea mays L.) plant under unstressed and salt-stressed conditions. Planta 232:533–543. https://doi.org/10.1007/s00425-010-1196-8
Marx MC, Kandeler E, Wood M et al (2005) Exploring the enzymatic landscape: distribution and kinetics of hydrolytic enzymes in soil particle-size fractions. Soil Biol Biochem 37(1):35–48. https://doi.org/10.1016/j.soilbio.2004.05.024
McArdle BH, Anderson MJ (2001) Fitting multivariate models to community data: a comment on distance-based redundancy analysis. Ecol 82:290–297
Mendes R, Garbeva P, Raaijmakers JM (2013) The rhizosphere microbiome: significance of plant beneficial, plant pathogenic and human pathogenic microorganisms. FEMS Microbiol Rev 37:634–663
Merino C, Godoy R, Matus F (2016) Soil enzymes and biological activity at different levels of organic matter stability. J Soil Sci Plant Nutr 16:14–30. https://doi.org/10.4067/S0718-95162016005000002
Mistry J, Finn RD, Eddy SR et al (2013) Challenges in homology search: HMMER3 and convergent evolution of coiled-coil regions. Nucleic Acids Res 41(12):e121. https://doi.org/10.1093/nar/gkt263
Mohanram S, Kumar P (2019) Rhizosphere microbiome: revisiting the synergy of plant-microbe interactions. Ann Microbiol 69:307–320. https://doi.org/10.1007/s13213-019-01448-9
Murphy KM, Edwards J, Louie KB et al (2021) Bioactive diterpenoids impact the composition of the root-associated microbiome in maize (Zea mays). Sci Rep 11:333. https://doi.org/10.1038/s41598-020-79320-z
Nannipieri P, Kandeler E, Ruggiero P (2002) Enzyme activities and microbiological and biochemical processes in soil. In: Burns RG, Dick RP (eds) Enzymes in the environment. Marcel Dekker, New York, pp 1–33
Nannipieri P, Ascher J, Ceccherini MT et al (2003) Microbial diversity and soil functions. Eur J Soil Sci 54:655–670. https://doi.org/10.1111/ejss.2_12398
Nannipieri P, Giagnoni L, Landi L, Renella G (2011) Role of phosphatase enzymes in soil. In: Bunemann EK, Obreson A, Frossard E (eds) Phosphorus in action. Springer, Berlin, pp 215–243
Nikolova PS, Bauerle TL, Häberle KH et al (2020) Fine-root traits reveal contrasting ecological strategies in European beech and Norway spruce during extreme drought. F Plant Sci 11:1211. https://doi.org/10.3389/fpls.2020.01211
Ofek M, Hadar Y, Minz D (2012) Ecology of root colonizing Massilia (Oxalobacteraceae). PLoS One 7(7):e40117. https://doi.org/10.1371/journal.pone.0040117
Oksanen J, Blanchet FG, Friendly M et al (2020) Vegan: community ecology package. R package Version 2. 5–7. https://cran.r-project.org, https://github.com/vegandevs/vegan. Accessed 1 Dec 2020
Pantigoso HA, Yuan J, He Y et al (2020) Role of root exudates on assimilation of phosphorus in young and old Arabidopsis thaliana plants. PLoS One 15(6):e0234216. https://doi.org/10.1371/journal.pone.0234216
Peiffer JA, Spor A, Koren O et al (2013) Diversity and heritability of the maize rhizosphere microbiome under field conditions. PNAS 110(16):6548–6553
Pervaiz ZH, Contreras J, Hupp BM et al (2020) Root microbiome changes with root branching order and root chemistry in peach rhizosphere soil. Rhizosphere 16:100249. https://doi.org/10.1016/j.rhisph.2020.100249
Phalempin M, Lippold E, Vetterlein D, Schlüter S (2021) Soil texture and structure heterogeneity predominantly governs bulk density gradients around roots. Vadose Zoen J 20:e20147. https://doi.org/10.1002/vzj2.20147
Quast C, Pruesse E, Yilmaz P et al (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596
R Core Team (2020) R: a language and environment for statistical computing. R foundation for statistical computing, Vienna. https://www.R-project.org. Accessed 15.10.2021
Ren C, Zhang W, Zhong Z et al (2018) Differential responses of soil microbial biomass, diversity and compositions to altitudinal gradients depend on plant and soil characteristics. Sci Total Environ 610:750–758. https://doi.org/10.1016/j.scitotenv.2017.08.110
Renoud S, Bouffaud M-L, Dubost A et al (2020) Co-occurrence of rhizobacteria with nitrogen fixation and/or 1-aminocyclopropane-1carboxylate deamination abilities in the maize rhizosphere. FEMS Microbiol Ecol 96:fiaa062. https://doi.org/10.1093/femsec/fiaa062
Ritz K, Young IM (2004) Interactions between soil structure and fungi. Mycologist 18:52–59. https://doi.org/10.1017/S0269915XO4002010
Robertson-Albertyn S, Alegria Terrazas R, Balbirnie K et al (2017) Root hair mutations displace the barley rhizosphere microbiota. F Plant Sci 8:094. https://doi.org/10.3389/fpls.2017.01094
Rognes T, Flouri T, Nichols B et al (2016) VSEARCH : a versatile open source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Rosenberg K, Bertaux J, Krome K et al (2009) Soil amoebae rapidly change bacterial community composition in the rhizosphere of Arabidopsis thaliana. ISME J 3:675–684. https://doi.org/10.1038/ismej.2009.11
Rüger L, Feng K, Dumack K et al (2021) Assembly patterns of the rhizosphere microbiome along the longitudinal root axis of maize (Zea mays L.). F Microbiol 12:614501. https://doi.org/10.3389/fmicb.2021.614501
Ruiz S, Koebernick N, Duncan S et al (2020) Significance of root hairs at the field scale – modelling root water and phosphorus uptake under different field conditions. Plant Soil 447:281–304
Rutherford PM, Juma NG (1992) Influence of texture on habitable pore space and bacterial-protozoan populations in soil. Biol Fertil Soil 12:221–227
Sasse J, Martinoia E, Northen T (2018) Feed your friends: do plant exudates shape the root microbiome? Trends Plant Sci 23:1. https://doi.org/10.1016/j.tplants.2017.09.003
Schimel JP, Gulledge JM, Clein-Curley JS et al (1999) Moisture effects on microbial activity and community structure in decomposing birch litter in the Alaskan taiga. Soil Biol Biochem 31:831–838
Schlatter DC, Kahl K, Carlson B et al (2018) Fungal community composition and diversity vary with soil depth and landscape position in a no-till wheat-based cropping system. F Microbiol Ecol 94:fiy098. https://doi.org/10.1093/femsec/fiy098
Schloss PD, Sarah LW, Ryabin T et al (2009) Introducing Mothur : open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75(23):7537–7541
Schlüter S, Zawallich J, Vogel H-J, Dörsch P (2019) Physical constraints for respiration in microbial hotspots in soil and their importance for denitrification. Biogeosci 16:3665–3678. https://doi.org/10.5194/bg-16-3665-2019
Seaton FM, George PBL, Paul BL et al (2020) Soil textural heterogeneity impacts bacterial but not fungal diversity. Soil Biol Biochem 144:107766. https://doi.org/10.1016/j.soilbio.2020.107766
Shannon C, Petigara N, Seshasai S (1948) A mathematical theory of communication. Bell Syst Tech J 27(3):379–423
Shoresh M, Harman GE (2010) Differential expression of maize chitinases in the presence or absence of Trichoderma harzianum strain T22 and indications of a novel exo- endo-heterodimeric chitinase activity. BMC Plant Biol 10:136
Silva UC, Medeiros JD, Leite LR et al (2017) Long-term rock phosphate fertilization impacts the microbial communities of maize rhizosphere. F Microbiol 8:1266. https://doi.org/10.3389/fmicb.2017.01266
Smalla K, Wieland G, Buchner A et al (2001) Bulk and rhizosphere soil bacterial communities studied by denaturing gradient gel electrophoresis: plant-dependent enrichment and seasonal shifts revealed. Appl Environ Microbiol 67(10):4742–4751
Smith SE, Jakobsen I, Grønlund M, Smith FA (2011) Roles of arbuscular mycorrhizas in plant phosphorus nutrition: interactions between pathways of phosphorus uptake in arbuscular mycorrhizal roots have important implications for understanding and manipulating plant phosphorus acquisition. Plant Physiol 156(3):1050–1057. https://doi.org/10.1104/pp.111.174581
Stone MM, Weiss M, Goodale C et al (2012) Temperature sensitivity of soil enzyme kinetics under N fertilization in two temperate forests. Glob Chang Biol 18:1173–1184
Sundberg C, Al-Soud WA, Larsson M et al (2013) 454 pyrosequencing analyses of bacterial and archaeal richness in 21 full-scale biogas digesters. FEMS Microbiol Ecol 85:612–626
Suzuki MT, Taylor LT, DeLong EF (2000) Quantitative analysis of small-subunit rRNA genes in mixed microbial populations via 5′- nuclease assays. Appl Environ Microbiol 66:4605–4614
Swarnalakshmi K, Yadav V, Tyagi D et al (2020) Significance of plant growth promoting rhizobacteria in grain legumes: growth promotion and crop production. Plants 9:1596. https://doi.org/10.3390/plants9111596
Tian P, Razavi BS, Zhang X et al (2020) Microbial growth and 214 enzyme kinetics in rhizosphere hotspots are modulated by soil organics and nutrient 215 availability. Soil Biol Biochem 141:107662. https://doi.org/10.1016/j.soilbio.2019.107662
Vetterlein D, Lippold E, Schreiter S et al (2021) Experimental platforms for the investigation of spatiotemporal patterns in the rhizosphere-laboratory and field scale. J Plant Nutr Soil Sci 184:35–55. https://doi.org/10.1002/jpln.202000079
Walters WA, Jin Z, Youngblut N et al (2018) Large-scale replicated field study of maize rhizosphere identifies heritable microbes. PNAS 115(28):7368–7373
Wei S, Jacquiod S, Philippot L et al (2021) Spatial analysis of the root system coupled to microbial community inoculation shed light on rhizosphere bacterial community assembly. Biol Fertil Soil 57:973–989. https://doi.org/10.1007/s00374-021-01590-0
Yim B, Winkelmann T, Ding G-C, Smalla K (2015) Different bacterial communities in heat and gamma irradiation treated replant disease soils revealed by 16S rRNA gene analysis – contribution to improved aboveground apple plant growth? F Microbiol 6:1224. https://doi.org/10.3389/fmicb.2015.01224
Yim B, Nitt H, Wrede A et al (2017) Effects of soil pre-treatment with Basamid® granules, Brassica juncea, Raphanus sativus and Tagetes patula on bacterial and fungal communities at two apple replant disease sites. F Microbiol 8:1604. https://doi.org/10.3389/fmicb.2017.01604
Yim B, Baumann A, Grunewaldt-Stöcker G et al (2020) Rhizosphere microbial communities associated to rose replant disease: links to plant growth and root metabolites. Hort Res 7:144. https://doi.org/10.1038/s41438-020-00365-2
Yuan F, Yin S, Xu Y et al (2021) The richness and diversity of catalases in bacteria. F Microbiol 12:645477. https://doi.org/10.3389/fmicb.2021.645477
Zai X, Luo W, Bai W et al (2021) Effect of root diameter on the selection and network interactions of root-associated bacterial microbiomes in Robinia pseudoacacia L. Microb Ecol 82:391–402
Zhang Y, Dong S, Gao Q (2017) Soil bacterial and fungal diversity differently correlated with soil biochemistry in alpine grassland ecosystems in response to environmental changes. Sci Rep 7:43077. https://doi.org/10.1038/srep43077
Acknowledgements
This project was carried out in the framework of the priority program 2089 “Rhizosphere spatiotemporal organization - a key to rhizosphere functions” funded by DFG (Deutsche Forschungsgemeinschaft), project numbers 403637238, 403640293, 403641192, 403664478 and 403635931. Seeds of the maize WT and rth3 were provided by Caroline Marcon and Frank Hochholdinger (University of Bonn). We also would like to acknowledge all participants involved in the sampling at UFZ Halle, Dr. Doreen Babin and Ilse-Marie Jungkurth for reading the MS.
Funding
Open Access funding enabled and organized by Projekt DEAL. This research was conducted within the research program “Rhizosphere Spatiotemporal Organization – a Key to Rhizosphere Functions” of the German Science Foundation, funded by the Deutsche Forschungsgemeinschaft (project numbers 403637238, 403640293, 403641192, 403664478 and 403635931). AHB was supported by the German Centre for Integrative Biodiversity Research (iDiv) Halle–Jena–Leipzig, funded by the Deutsche Forschungsgemeinschaft (DFG-FZT 118, 202548816).
Author information
Authors and Affiliations
Contributions
MG and MT designed and carried out experiments. ZI, LR, MG and BY collected the samples. BY prepared rhizosphere DNA and amplicon sequencing analyses of 16S rRNA gene fragments and ITS regions. LM prepared amplicon libraries of 16S rRNA gene and ITS region fragments, sequenced and produced ASV Tables. LR performed amplicon-sequencing of cercozoa. ZI analysed soil microbial extracellular enzyme activities. AHB contributed data analyses. BY, ZI, LR, MT, DV, MB, EB and KS wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Responsible Editor: Hans Lambers.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 1.35 mb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yim, B., Ibrahim, Z., Rüger, L. et al. Soil texture is a stronger driver of the maize rhizosphere microbiome and extracellular enzyme activities than soil depth or the presence of root hairs. Plant Soil 478, 229–251 (2022). https://doi.org/10.1007/s11104-022-05618-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-022-05618-8