Abstract
Background and aims
Plant breeding activities shape the rhizosphere microbiome but less is known about the relationship of both with the seed microbiome. We analyzed the composition of bacterial communities of seeds and rhizospheres of Styrian oil pumpkin genotypes in comparison to bulk soil to elucidate specific microbial signatures to support a concept involving plant-microbe interactions in breeding strategies.
Methods
The seed and rhizosphere microbiomes of 14 genotypes of oilseed pumpkin and relatives were analyzed using a 16S rRNA gene amplicon sequencing approach, which was assessed by bioinformatics and statistical methods.
Results
All analyzed microhabitats were characterized by diverse bacterial communities, but the relative proportions of phyla and the overall diversity was different. Seed microbiomes were characterized by the lowest diversity and dominant members of Enterobacteriaceae including potential pathogens (Erwinia, Pectobacterium). Potential plant-beneficial bacteria like Lysobacter, Paenibacillus and Lactococcus contributed to the microbial communities in significant abundances. Interestingly, strong genotype-specific microbiomes were detected for seeds but not for the rhizospheres.
Conclusions
Our study indicates a strong impact of the Cucurbita pepo genotype on the composition of the seed microbiome. This should be considered in breeding of new cultivars that are more capable of exploiting beneficial indigenous microbial communities.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microbial communities have central roles for plant development and health throughout the entire life cycle (Mendes et al. 2011; Philippot et al. 2013). This knowledge has been a revolutionary advance in biological sciences, also directing plant research towards a more holistic view (Berg et al. 2016). Since more than a century, diversity and function of the rhizosphere microbiome was intensively studied and the impact of the plant genotype and soil quality determined (Hiltner 1904; Smalla et al. 2001; Berg and Smalla 2009; Lundberg et al. 2012). Recent studies on crop cultivars revealed that breeding shapes the composition of the root-associated bacterial communities including the antagonistic potential towards pathogens (Peiffer and Ley 2013; Bouffaud et al. 2014; Cardinale et al. 2015). In the past, mainly seed-borne pathogens were studied but recently the influence of the whole seed microbiome on plant health has gained more interest (Aleklett and Hart 2013; Barret et al. 2015). Seeds are of particular interest as microbial carriers because they are involved in the transmission of both potential beneficial and pathogenic microorganisms from one generation to another (Johnston-Monje et al. 2016). Breeding plants for beneficial plant-microbe interactions is an emerging field mainly focusing on below ground interactions in the rhizosphere (reviewed in Wissuwa et al. 2009; Bakker et al. 2012); however the impact of the seed microbiome is completely unclear.
Styrian oil pumpkin (Cucurbita pepo L. subsp. pepo var. styriaca Greb.) is a cultural heritage of the Austro-Hungarian Empire and of international importance today. The seeds are used for the production of a unique, dark-green seed oil, which is traditionally consumed in Austria, and is increasingly popular in gourmet cuisines worldwide. In 2016, the acreage of the Styrian oil pumpkin reached a peak level with 39,450 ha in Austria, and the market demand for this high-value crop continues to rise. New growing areas have been mainly established in China but also in the US. The morphological structure of C. pepo seeds comprises a root-hypocotyl embryo, two distinct photosynthetic cotyledons, a thin endosperm, remains of the nucellus and a seed coat with five layers. Styrian oil pumpkin emerged from a natural mutation and is lacking a lignification of the seed coat resulting in a high susceptibility to various fungal and bacterial pathogens during seed germination (Heinisch and Ruthenberg 1950). Thus, commercially available Styrian oil pumpkin seeds are by default treated with chemical strippers; mainly synthetic fungicides or copper-based products. Sowing of untreated seeds generally results in a drastic reduction of germination rates or germination fails totally, if weather or soil conditions are unfavorable after sowing. A disease responsible for high yield losses is the fruit rot caused by the consortium of Didymella bryoniae and Pectobacterium carotovorum (syn. Erwinia carotovora) (Grube et al. 2011) or Erwinia atrosepticum. Leaves of adult plants can be infested by fungi such as Didymella bryoniae or Phyllosticta cucurbitacearum (Bedlan 2012) and by bacterial pathogens like Xanthomonas campestris pv. cucurbitae, Pseudomonas syringae or P. viridiflava. Strong cultivar-specificity regarding the susceptibility to fruit rot was shown in evaluations of registered oil pumpkin cultivars (AGES 2016; Winkler et al. 2008). A high genotype-specificity has been widely shown for the interactions of plants with pathogens (Neupane et al. 2015; Bruns et al. 2012; Rubiales and Niks 1996) and thus breeding for resistances against pathogens is common practice (Niks et al. 2011; Pachner et al. 2015; Roane 1973; Ashkani et al. 2015). Conversely, genotype-specific beneficial plant-microbe associations have not been considered in breeding strategies thus far. Our hypothesis was that pumpkin seeds contain a genotype-specific microbiome, which consists of a core subset of the plant-associated microbiome with mainly plant-beneficial traits.
The objective of this study was a comprehensive microbiome analysis based on a 16S rRNA gene sequencing approach targeting the bacterial and parts of the archaeal diversity of 14 C. pepo genotypes. Those included one open-pollinated cultivar, three hybrids and their pedigree components (four inbred lines) and five segregating lines of the Styrian oil pumpkin as well as a zucchini hybrid. The study should be rated as a first step towards gaining a deeper understanding of genotype-dependent differences in plant-microbe interactions of the Styrian oil pumpkin. The overall aim is the development of a concept to (re-)integrate beneficial plant-microbe interactions into the plant breeding activities of the Styrian oil pumpkin.
Materials & methods
Pumpkin genotypes
Upon selection of the C. pepo genotypes for analysis, the focus was toward coverage of cultivars with a high market share (three-way cross hybrids ‘GL Opal’ and ‘GL Rustikal’) including their pedigree components (inbred Line A – D and the single cross hybrid ‘Gleisdorfer Diamant’ (‘Gl. Diamant’)). An open-pollinated cultivar frequently used in organic agricultural systems, ‘GL Classic’, and six other cultigens bred in countries other than Austria were included to broaden the spectrum of genotypes (Table 1). The geographic origin records the country in which a genotype was selected or bred. With the exception of the single cross zucchini hybrid Naxos, the seeds used for amplicon sequencing approaches were harvested from plants grown on three different field sites near Gleisdorf (province of Styria, Austria). Post-harvest processing of those seeds was performed according to the standard procedures of the Saatzucht Gleisdorf GmbH breeding station.
Sampling of seed, rhizosphere and soil replicates
For the seed microbiome analysis, 40 seeds of each genotype were washed five times for one minute with 50 mL sterile deionized water and soaked in 25 mL sterile deionized water for 4 h on a rotary shaker at 100 rpm. Subsequently, the seeds were divided into four replicates of 10 seeds each and were ground with a pestle in 10 mL 0.85% NaCl in sterile bags (Nasco Whirl-Pak®). For each replicate, a 3 mL suspension was pelleted by 20 min centrifugation at 4°C and 13,500 g.
For the rhizosphere microbiome analysis, 40 seeds per genotype of the same seed lots used for seed microbiome analysis were coated with 0.3 g of the fungicide Maxim® XL (Syngenta) and split into four replicates. Seeds were sown at a field site near the breeding station of Saatzucht Gleisdorf GmbH in randomized plots (47°06′57.3″N, 15°42′31.3″ E). The soil of the field site is described as gleyed loose brown earth, loamy silt, and cover loams on a quaternary terrace deficient in lime, with a pH-value of 6.5. In parallel 40 seeds per genotype were sown without fungicide coating on the same field site. As only 2.7% of those plants emerged, the rhizosphere samples had to be taken from plants grown from fungicide treated seeds (93.1% germination rate): one month after sowing, rhizosphere material from four randomly chosen plants per plot was sampled and pooled. Additionally, four bulk soil samples were taken from random places at the field site. Five to 7 g of each rhizosphere and soil replicate were suspended in 50 mL 0.85% NaCl and homogenized by a 3 min bag mixer (stomacher) treatment, then 4 mL of the homogenized solution were pelleted as described above.
DNA extraction and amplicon sequencing
The DNA extraction was performed using a modified protocol of the FastDNA™ SPIN Kit for Soil (MP Biomedicals). The variable region 4 of the 16S rRNA gene was amplified with 515f (5´-GTGCCAGCMGCCGCGGTAA-3´) and 806r (5´-GGACTACHVGGGTWTCTAAT-3´) primers extended by individual barcodes. For PCR amplification, a modified protocol of Lundberg et al. (2013) including synthetic peptide nucleic acid PCR clamps (PNAs) for blocking the amplification of mitochondrial and plastid 16S rRNA gene sequences of plants was applied. Three independent PCR amplifications were performed per replicate sample. The triplicate amplification products were pooled and purified using the Wizard® SV Gel and PCR Clean-Up System (Promega) protocol. The PCR products were adjusted volumetrically to reach equimolarity of each sample in one common pool for 16S rDNA sequencing. Amplicon sequencing was performed with the Illumina MiSeq V2 sequencing platform (2 × 150 bp paired-end) by GATC Biotech (Germany).
Bioinformatic and statistical analyses
Following de-multiplexing of raw reads and clipping of sequencing adapters, read pairs were joined and sorted according to sample-specific barcodes. Joint reads were further analyzed with the open-source bioinformatics pipeline QIIME 1.9.1 (Caporaso et al. 2010a). First, reads were quality (Phred score ≥ 20) and length (290–300 bp) filtered, and primer and barcodes flanking the reads were clipped. Chimeric sequences were removed by means of the de novo UCHIME method (Edgar 2010; Edgar et al. 2011). Remaining sequences were clustered into operational taxonomic units (OTUs) at a 97% similarity level using the de novo UCLUST clustering method with default parameters (Edgar 2010). The most abundant representative read per OTU was picked and taxonomically assigned using RDP Classifier 2.2. (Wang et al. 2007) based on the reference database Greengenes release gg_13_8_99 (DeSantis et al. 2006). The alignment of reads was performed using PyNAST (Caporaso et al. 2010b). Additionally, an approximately-maximum-likelihood phylogenetic tree using FastTree 2.1.3. (Price et al. 2010) was generated.
Each replicate was comprised of 11,245 to 276,132 sequences after initial data processing. Prior to statistical analyses, the mitochondrial (ranging from 0.6% to 32.9% per replicate) and plastid (0.3% to 6.8%) sequences with plant origin and unassigned OTUs were excluded by filtering the OTU table. Additionally, out of four replicates per sample, the replicate with the lowest read number was discarded. As the remaining replicates comprised of 3758 to 256,248 sequences, the subsequent analyses were performed after normalizing the sequence number per replicate to 3758.
Statistical analyses to calculate significance of differences in diversity indices were performed using the non-parametric Kruskal-Wallis Rank Sum Test (Hollander and Wolfe 1973) and the Pairwise Test for Multiple Comparisons of Mean Rank Sums (Nemenyi-Test) (Sachs 1997) implemented in the open source data analysis software RStudio (RStudio Team 2015). Non-parametric analyses of similarities (ANOSIM) were calculated according to Fierer et al. (2010) and Clarke (1993). A non-metric multidimensional scaling (NMDS) analysis was performed using the open source data analysis software RStudio and the function metaMDS {vegan} (Faith et al. 1987; Minchin 1987) with calculation of the distance matrix based on a Bray-Curtis algorithm.
Results
Microbial communities associated with seed, rhizosphere and soil
All analyzed microhabitats were characterized by a high diversity of certain phyla but the relative proportions were different (Fig. 1). Proteobacteria predominated seed (83%), rhizosphere (41%) and soil (24%) microbiomes and considerable proportions of the phyla Firmicutes (11%, 8% and 6%) and Actinobacteria (2%, 17% and 15%) were found in all habitats. Thaumarchaeota as well as Acidobacteria, Bacteroidetes, Chloroflexi, Nitrospirae, Gemmatimonadetes, Planctomycetes, and Verrucomicrobia contributed to the microbiomes of the rhizosphere and soil and only to minor degree to the seed microbiome.
Circular representation of the proportional structure of bacterial communities at phylum level associated with rhizosphere and seeds of C. pepo genotypes as well as soil (outer circle). Taxa with a proportion lower than 1.5% in all three habitats are summarized as ‘Others’. Values within the inner circle indicate the number of reads of a phylum within the normalized dataset. The graphic was built using the open-source software Circos (Krzywinski et al. 2009)
OTUs representing the core microbiome were calculated separately for each of the habitats seed, rhizosphere and soil and then summarized (Fig. 2). Differences in the relative abundance of those OTUs between the core microbiomes exist for several Nitrososphaeria, Acidobacteria, Chloracidobacteria and Chloroflexi OTUs, which occurred in a higher proportion in the soil microbiome compared to the rhizosphere and seed microbiomes, whereas several Bacilli, Actinobacteria, Saprospirae, Alpha-, Beta- and parts of the Gammaproteobacteria OTUs occurred to a greater degree in the rhizosphere microbiome in comparison to the soil and seed microbiomes. The seed core microbiome was dominated by high abundances of eight Gammaproteobacteria (seven Enterobacteriaceae and one Pseudomonadaceae) and two Bacilli (one Lactococcus and one Exiguobacterium) OTUs.
Summary of the core microbial communities represented by 16S rRNA gene sequences in the seed and rhizosphere of C. pepo genotypes as well as in the bulk soil. Relative abundance values of taxa belonging to the core of a habitat are framed, while other abundance values are not belonging to the core in the respective habitat. OTUs with abundance lower than 0.5% in all core microbiomes are summarized as ‘Others’. Taxa marked with asterisks were complemented with additional taxonomic information from NCBI database, while the other denominations are from Greengenes database. The phylogenetic tree using the representative sequences of the OTUs was calculated with the NCBI tree method fast minimum evolution (max sequence difference 0.75) and illustrated in MEGA7
OTU distribution and diversity analyses
The comparison of the seed and rhizosphere core microbiomes with the bulk soil microbiome shows that seeds and rhizosphere shared only 10.5% of the total OTUs including OTUs from soil, whereas the rhizosphere and the bulk soil shared 32.6% of those OTUs (Fig. 3a). Within the rhizosphere OTUs 16.8% were conserved in seeds and soil as well, 4.5% were derived solely from the seeds and 49.6% solely from the soil. Apparently, the seed microbiome has a smaller influence on the rhizosphere communities than the soil microbiome. 29.0% of the rhizosphere OTUs were unique. Some of those OTUs could have been derived from the seed testa, but were removed due to the washing and soaking procedure for seed microbiome analysis. This probably released a number of microorganisms to the washing suspension. Another source of inoculum could have been rare soil bacteria that were below the detection level in the soil microbiome analysis but have been enriched in the rhizosphere due to the rhizosphere effect. A detailed analysis of the contribution of Enterobacteriaceae to the communities revealed that major proportions of those OTUs observed were unique for seeds, whereas in rhizosphere and soil no unique Enterobacteriaceae were detected (Fig. 3b). The same trends were observed when the analyses were calculated on C. pepo genotype level: the family of Enterobacteriaceae was strongly associated with the seed as habitat.
Alpha diversity measures Chao1 and Shannon indices revealed that the species richness in the rhizosphere was significantly higher than in the seeds (calculated with the QIIME script alpha_diversity.py; paired t-test, significance level α = 0.01, p values = 0.003), whereas the richness in the soil was significantly higher than in the rhizosphere. The Heip index (calculated with the QIIME script alpha_diversity.py) indicated that the evenness in the seeds was considerably lower than in the rhizosphere and soil, meaning that the relative abundance of taxa was not evenly distributed. The seed microbiomes of the genotypes Line E, ‘Gleisdorfer Diamant’ and Line G showed a higher alpha diversity (Shannon diversity index H´ of 8.6, 7.9 and 7.1, respectively and Heip evenness index E´ of 0.29, 0.17 and 0.16, respectively) than the other investigated genotypes. In contrast to the rhizosphere, differences in alpha diversity among the seeds of several genotypes were significant (Table 2).
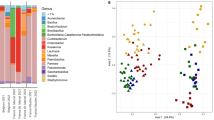
Genotype specific colonization patterns of seeds
The microbial seed communities of all C. pepo genotypes were dominated by Proteobacteria with 83% on average, 64% belonged to the family of Enterobacteriaceae. Genotype-specific colonization patterns of the seeds were evident in the open-pollinated cultivar ‘GL Classic’ and in the inbred Line D (Fig. 4). In those genotypes, the genus Erwinia was part of the microbiome with a relative abundance of 38% and 33% respectively. A detailed analysis of the sequences clustered in Erwinia genera revealed that they contained sequences of the important pathogen Pectobacterium carotovorum (syn. Erwinia carotovora). The genus Pseudomonas was present in all genotypes but was enriched in four out of six cultivars bred in other countries than Austria. Firmicutes were enriched in the two three-way cross hybrids ‘GL Opal’ (19%) and ‘GL Rustikal’ (36%), as well as in the two inbred Lines B and D and the segregating Line F. Within the Firmicutes, the genus Lactococcus was more abundant in four out of eight cultivars bred in the province of Styria (Austria) and only low abundant in cultivars bred in other countries. The genus Acinetobacter was more abundant in the segregating lines, especially in Line H with 11%. Actinobacteria were observed to a higher extent in Line E, Line G and ‘Gleisdorfer Diamant’. These three genotypes had similar communities which were more diverse than those of the other genotypes, congruent with the results of the calculated diversity indices.
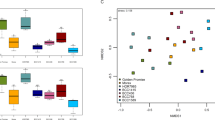
No specific patterns concerning the field site origin of studied seeds were observed. According to the non-parametric analysis of similarities the genotype (R = 0.527, p value = 0.001) has greater influence on the bacterial community composition than the field origin of the seeds (R = 0.181, p value = 0.008). While only 21 OTUs showed significantly different abundances among the rhizosphere (Kruskal-Wallis test, α = 0.01), 121 OTUs differed significantly among the seed microbiomes of the 14 genotypes. Six OTUs showed significantly different degrees of abundancy in both habitats. These OTUs were assigned to the taxa of Exiguobacterium, Chthoniobacteraceae, Nitrospirales, Xanthomonadaceae and Bacillus, according to the Greengenes database. In order to visualize the beta diversity and the relationships of seed associated bacterial taxa with significant different abundances among the genotypes, a NMDS analysis of the 14 C. pepo genotypes was performed based on a community-by-species matrix comprising of 121 significantly different OTUs (Fig. 5).
Non-metric multidimensional scaling of 121 significantly different bacterial OTUs in seed microbiomes of 14 C. pepo genotypes. Genotypes bred in Austria are written in bold, genotypes bred in the Netherlands (Naxos), Germany (Line E), Slovenia (Line F and G) or China (Line H and I) in italics. The stress (i.e. the discrepancy between 2D configuration and predicted values from regression) was 0.159
Altogether, 53 of the 121 different abundant seed-associated OTUs referred to the phylum of Proteobacteria, wherein 12 belonged to the family of Enterobacteriaceae, of which three were further assigned to the genus Erwinia. Higher abundances of Pseudomonas viridiflava were observed in the seed microbiomes of Line F and Line I as well as in the open-pollinated ‘GL Classic’. Furthermore, the abundances of Lysobacter and Paenibacillus were significantly different among the seed microbiomes of the genotypes investigated. Within this analysis, the phylum Firmicutes comprised of 23 OTUs, wherein 11 OTUs were assigned to the family of Leuconostocaceae and four to Paenibacillaceae. A large proportion of Leuconostocaceae is located at similar coordinates as ‘GL Rustikal’ and ‘GL Opal’ within the NMDS plot. A weak clustering is distinguishable for the Lines A, B, C, D, ‘Gleisdorfer Diamant’, ‘GL Classic’, ‘GL Opal’ and ‘GL Rustikal’ with the geographic origin in Austria. The three cultivars with a highly diverse seed microbiome, ‘Gleisdorfer Diamant’, Line E and Line G, together with Line I are located at similar coordinates as the group of ‘Other’ bacterial taxa with significantly different abundances. According to the Shannon diversity index shown in Table 2, Line I is the fourth diverse C. pepo cultivar.
Specific analyses of seed microbiomes of agronomically important cultivars and of the ‘GL Rustikal’ pedigree
When comparing the three agronomically most important cultivars ‘GL Classic’, ‘GL Rustikal’ and ‘GL Opal’ with the most important component in the hybrid seed production, ‘Gleisdorfer Diamant’, 14% of the OTUs comprised the core microbiome (Fig. 6a). The ‘Gleisdorfer Diamant’ seeds were colonized with 90 unique OTUs compared to the other three genotypes investigated. Concerning the Enterobacteriaceae (Fig. 6b), 13 of the observed OTUs were shared within all genotypes investigated, 13 OTUs were unique to ‘GL Opal’, three to ‘GL Rustikal’ and five to ‘GL Classic’. ‘Gleisdorfer Diamant’ seeds did not harbor unique Enterobacteriaceae OTUs. Three Erwinia OTUs were common among all four genotypes, one and three were unique in ‘GL Opal’ and ‘GL Classic’ and none were exclusively observed in ‘GL Rustikal’ and ‘Gleisdorfer Diamant’.
OTU distribution within the seed microbiomes of (a) total OTUs and (b) Enterobacteriaceae OTUs of the three-way cross hybrids ‘GL Rustikal’ and ‘GL Opal’, the single cross hybrid ‘Gleisdorfer Diamant’ and the open-pollinated cultivar ‘GL Classic’. OTU numbers of the genus Erwinia are in superscript. Fraction of samples that OTU was observed in to be considered as ‘core’: 100%
To visualize the relationships of the seed associated bacterial taxa among the different genotypes a taxonomic interaction network (illustrated in Fig. 7a) of the three-way cross hybrid ‘GL Rustikal’ and its pedigree components (relationships shown in Fig. 7b) was created. The seed microbiome of the highly diverse ‘Gleisdorfer Diamant’ was comprised of 117 taxa of which 65 were unique, whereas the seed microbiomes of the other genotypes showed no (Line A, Line D, ‘GL Rustikal’) or just one unique taxa (Line B). The family of Enterobacteriaceae dominated the seed associated communities of the ‘GL Rustikal’ pedigree. The analysis of the OTU distribution within the ‘GL Rustikal’ pedigree components (Fig. 7c) revealed that ‘Gleisdorfer Diamant’ and Line D as parental components shared 22% and 20% of OTUs with ‘GL Rustikal’, whereas the genetically more distant components ‘Gleisdorfer Diamant’ and Line D shared 29%. The core microbiome of all five genotypes investigated was comprised of 14% of the OTUs. 14 (30%) of the observed Enterobacteriaceae OTUs, including three Erwinia OTUs, were shared within all five genotypes, which could indicate their having an essential function or that the inheritance of the microbiome is focused on certain taxa. There may be a connection of the higher bacterial diversity in ‘Gleisdorfer Diamant’ with the displacement and competition of Enterobacteriaceae in its seeds.
a Taxonomic interaction network (created with the open source bioinformatics software Cytoscape (Shannon et al. 2003)) at genus level; seed associated bacterial communities of the ‘GL Rustikal’ pedigree and b the pedigree component relationships. The outer squares in the network illustrate the C. pepo genotypes. Seed associated bacterial taxa with a relative abundance of at least 0.1% are connected with the corresponding genotype by a grey line. The line width correlates with the relative abundance of each taxon connected with the respective genotype. The size of the shape next to the bacterial taxa corresponds to the mean of the relative abundance of all 118 bacterial taxa analyzed in this network. The 20 taxa belonging to the core microbiome are shown as triangles. Taxa that are shared by four, three or two genotypes, or that are unique in a genotype are illustrated as circles. Taxa of the same phylum are depicted in the same shape color. c Distribution of total OTUs within the seed microbiomes of the three-way cross hybrid ‘GL Rustikal’ and its pedigree components and of Enterobacteriaceae OTUs (fraction of samples that OTU was observed in to be considered ‘core’: 100%) The proportions are not drawn to scale
A Blast analysis against the NCBI nucleotide database for Enterobacteriaceae OTUs with an observation count of more than 10 sequences per OTU of the ‘GL Rustikal’ pedigree component’s seed and rhizosphere samples revealed that the 16S rRNA gene sequences of most of the 54 Enterobacteriaceae OTUs had the highest similarity to Klebsiella sp. Other OTUs were assigned to Pantoea sp., Salmonella sp., Enterobacter sp., Trabulsiella sp., Yersinia sp., Erwinia sp., Kluyvera sp. and Cedecea sp. Pectobacterium carotovorum was part of the seed microbiome of all genotypes, except ‘GL Rustikal’, but was not detected in rhizosphere and bulk soil.
Discussion
Pumpkin seeds and those from related breeding lines are associated with a unique and genotype-specific microbiome. In comparison to the rhizosphere and bulk soil, microbial seed communities were characterized by a lower bacterial diversity, dominantly comprising members of Enterobacteriaceae including potential pathogens (Erwinia, Pectobacterium), but also beneficial bacteria like Lysobacter, Paenibacillus, and Lactococcus. In general, the data confirmed our hypothesis of a genotype-specific microbiome consisting of mainly plant-beneficial traits but several interesting and different findings will be additionally discussed.
The rhizosphere as crucial soil-plant interface was described for the first time by Lorenz Hiltner in 1904. The rhizosphere effect triggered by root exudates is also well studied (Rovira 1956) and the phenomenon that rhizosphere communities harbor less diversity than the surrounding soil was confirmed by deep sequencing technologies (Lundberg et al. 2012; Bulgarelli et al. 2012). In our study, we showed that several taxa such as Bacilli, Actinobacteria, Saprospirae, Alpha-, Beta- and members of the Gammaproteobacteria were enriched in the rhizosphere microbiome, while Thaumarchaeota, Acidobacteria, and Chloroflexi occurred in higher proportions in the soil. These results fit to the conclusion that Gammaproteobacteria, Betaproteobacteria and Firmicutes contribute to disease suppressive microbiome and that plants selectively select beneficial microorganisms (Berendsen et al. 2012). In addition to the rhizosphere effect, we can define a spermosphere, or better a seed effect, because the spermosphere only comprises the microenvironment surrounding the seeds. This seed effect is also characterized by a selective enrichment of specific microorganisms of which we assume that they are useful for germination and plant growth and health.
Traditionally, seeds were considered as carrier for pathogens only. In our study, we found potential pathogens as well as potential plant-beneficial microbes. First of all, we have to consider that 16S rDNA analyses, as performed in our study, allow only a limited prediction of the functional role. However, our analyses revealed several findings, which have to be interpreted carefully. For example, Pectobacterium carotovorum, an important pathogen causing fruit rot on the Styrian oil pumpkin, was part of the seed microbiome in all genotypes, with the exception of ‘GL Rustikal’. The genus Erwinia was enriched in two genotypes. Moreover, the genus Pseudomonas was present in the seed microbiome of all genotypes, which comprises a number of beneficial species (Avis et al. 2008), but also includes species which can cause leaf necrosis in Cucurbita pepo, such as P. vridiflava (Grube et al. 2011; Huss and Mavridis 2007). Representatives of X. campestris and P. syringae, for example, have been shown to be seed-borne pathogens in the Cucurbitaceae family (Zitter et al. 1996; Robinson and Decker-Walters 1997; Babadoost and Zitter 2009; Blancard et al. 1994).
Possible beneficials like Lysobacter and Paenibacillus were also part of the seed microbiomes. Strains of both genera, Lysobacter gummosus L101 and Paenibacillus polymyxa PB71, were already used for seed treatment and resulted in significant effects on harvest yields (Fürnkranz et al. 2012). The genus Serratia includes potential biocontrol species as well, which led to considerable increases in germination rates of chemically untreated seeds in field experiments of the same study, but was detected in extremely low abundances in 16 out of 56 seed samples and in 6 out of 56 rhizosphere samples. In contrast, no Serratia signature was identified in analyzed soil samples, which indicates a specific plant-associated occurrence. Based on the results of this study, Lactococcus species can also be suggested as biological control agents for the Styrian oil pumpkin. This genus was highly abundant in some genotypes and Shrestha et al. (2014) reported good effects of lactic acid bacteria against Pectobacterium carotovorum as well as Xanthomonas campestris.
Due to, compared to other crops, a relatively short breeding history, the genetic differences between the Cucurbita pepo L. subsp. pepo var. styriaca genotypes are small, as this thin-coated seed segregant first appeared in the late nineteenth century (Teppner 2004). The commercial breeding program for the Styrian oil pumpkin in Austria at Saatzucht Gleisdorf GmbH started in 1960 and was intensified 20 years ago. Thus, it is remarkable that cultivar dependent differences in the seed microbiomes were found within this narrow gene pool. Our results contrast with the findings of Klaedtke et al. (2016), which revealed that the microbial assemblages of bean seeds were shaped by the seed production site rather than by the genotype. Within the components of the ‘GL Rustikal’ pedigree, the seeds of ‘Gleisdorfer Diamant’ were colonized by a significantly more diverse microbiome than the other genotypes, whereas for example ‘GL Rustikal’ seeds were characterized by a significantly greater level of colonization of Leuconostocaceae than other genotypes. The seed core microbiome of the cultivars analyzed in this study is dominated by high abundances of seven Enterobacteriaceae, one Pseudomonadaceae, one Lactococcus and one Exiguobacterium OTU.
The results of the seed analyses are of particular interest for the seed production industry, as the Styrian oil pumpkin is highly susceptible to various fungal and bacterial pathogens during germination making chemical or complex seed treatments inevitable. It remains to be investigated to which extent naturally occurring seed-borne bacteria influence germination and plant development. The dominance of the seed-associated microbiomes by Proteobacteria, and in particular Enterobacteriaceae, may contribute to disease susceptibility as the microbial richness and evenness of the microbial taxa are important for the maintenance of plant health (Bakker et al. 2012). The interactions of seed-borne microorganisms with the indigenous soil populations may influence the expression of biological control traits or the subsequent colonization of the rhizosphere (Nelson 2004). Therefore, the results of our study could direct the design of tailored biological seed treatments or influence seed disinfection strategies that might replace fungicide treatments in future. A possible implication for breeding programs could be the selection of genotypes enriching less enterobacteriaceal pathogens and/or expressing a higher microbial diversity in their seeds. The hologenome theory of evolution postulates that the host and its associated beneficial microbiome (holobiont) co-evolve as one unit to provide benefits to one another including defense mechanisms (Zilber-Rosenberg and Rosenberg 2008). When postulating that breeding plants is some form of directed evolution, it may be assumed that breeding under conventional conditions (for example with the use of chemical strippers and fungicides) leads to a loss of natural defense mechanisms originally provided by the holobiont system. The results of studies like this could contribute to a paradigm shift towards ecological breeding programs.
References
AGES, editor (2016) Österreichische Beschreibende Sortenliste 2016 Landwirtschaftliche Pflanzenarten Schriftenreihe 21/2016
Aleklett K, Hart M (2013) The root microbiota - a fingerprint in the soil? Plant Soil 370:671–686. doi:10.1007/s11104-013-1647-7
Ashkani S, Rafii MY, Shabanimofrad M, Miah G, Sahebi M, Azizi P, Tanweer FA, Akhtar MS, Nasehi A (2015) Molecular breeding strategy and challenges toward improvement of blast disease resistance in rice crops. Front Plant Sci 6:1–14. doi:10.3389/fpls.2015.00886
Avis TJ, Gravel V, Antoun H, Tweddell RJ (2008) Multifaceted beneficial effects of rhizosphere microorganisms on plant health and productivity. Soil Biol Biochem 40:1733–1740. doi:10.1016/j.soilbio.2008.02.013
Babadoost M, Zitter T (2009) Fruit rots of pumpkin - a serious threat to the pumpkin industry. Plant Dis 93(8):772–782. doi:10.1094/PDIS-93-8-0772
Bakker MG, Manter DK, Sheflin AM, Weit TL, Vivanco JM (2012) Harnessing the rhizosphere microbiome through plant breeding and agricultural management. Plant Soil 360:1–13. doi:10.1007/s11104-012-1361-x
Barret M, Briand M, Bonneau S, Préveaux A, Valière S, Bouchez O, Hunault G, Simoneau P, Jacques MA (2015) Emergence shapes the structure of the seed microbiota. AEM 81(4):1257–1266. doi:10.1128/AEM.03722-14
Bedlan G (2012) First report of Phyllosticta cucurbitacearum on Cucurbita pepo Var. styriaca in Austria. J Kult 64(5):171–172
Berendsen RL, Pieterse CMJ, Bakker PAHM (2012) The rhizosphere microbiome and plant health. Trends Plant Sci 17:478–486. doi:10.1016/j.tplants.2012.04.001
Berg G, Smalla K (2009) Plant species and soil type cooperatively shape the structure and function of microbial communities in the rhizosphere. FEMS Microbiol Ecol 68:1–13. doi:10.1111/j.1574-6941.2009.00654.x
Berg G, Rybakova D, Grube M, Köberl M (2016) The plant microbiome explored: implications for experimental botany. J Exp Bot 67(4):995–1002. doi:10.1093/jxb/erv466
Blancard D, Lecoq H, Pitrat M (1994) Cucurbit diseases: observation, identification and control (a colour atlas). Halstead Press, New York
Bouffaud ML, Poirier MA, Muller D, Moënne-Loccoz Y (2014) Root microbiome relates to plant host evolution in maize and other Poaceae. Environ Microbiol 16(9):2804–2814. doi:10.1111/1462-2920.12442
Bruns E, Carson M, May G (2012) Pathogen and host genotype differently affect pathogen fitness through their effects on different life-history stages. BMC Evol Biol 12(135):1–13. doi:10.1186/1471-2148-12-135
Bulgarelli D, Rott M, Schlaeppi K, van Ver L, Themaat E, Ahmadinejad N, Assenza F, Rauf P, Huettel B, Reinhardt R, Schmelzer E, Peplies J, Gloeckner FO, Amann R, Eickhorst T, Schulze-Lefert P (2012) Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488(7409):91–95. doi:10.1038/nature11336
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010a) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. doi:10.1038/nmeth.f.303
Caporaso JG, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R (2010b) PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26:266–267. doi:10.1093/bioinformatics/btp636
Cardinale M, Grube M, Erlacher A, Quehenberger J, Berg G (2015) Bacterial networks and co-occurrence relationships in the lettuce root microbiota. Environ Microbiol 17(1):239–252. doi:10.1111/1462-2920.12686
Clarke KR (1993) Non-parametric multivariate analysis of changes in community structure. J Aust Ecol 18:117–143. doi:10.1111/j.1442-9993.1993.tb00438.x
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. AEM 72:5069–5072. doi:10.1128/AEM.03006-05
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. doi:10.1093/bioinformatics/btq461
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. doi:10.1093/bioinformatics/btr381
Faith DP, Minchin PR, Belbin L (1987) Compositional dissimilarity as a robust measure of ecological distance. Vegetatio 69(1):57–68. doi:10.1007/BF00038687
Fierer N, Lauber CL, Zhou N, McDonald D, Costello EK, Knight R (2010) Forensic identification using skin bacterial communities. Proc Natl Acad Sci 107:6477–6481. doi:10.1073/pnas.1000162107
Fürnkranz M, Adam E, Müller H, Grube M, Huss H, Berg G (2012) Promotion of growth, health and stress tolerance of Styrian oil pumpkins by bacterial endophytes. Eur J Plant Pathol 134(3):509–519. doi:10.1007/s10658-012-0033-2
Grube M, Fürnkranz M, Zitzenbacher S, Huss H (2011) Emerging multi-pathogen disease caused by Didymella bryoniae and pathogenic bacteria on Styrian oil pumpkin. Eur J Plant Pathol 131:539–548. doi:10.1007/s10658-011-9829-8
Heinisch O, Ruthenberg M (1950) Die Bedeutung der Samenschale für die Züchtung des Ölkürbis. Z Pflanzenzüchtung 29:159–174
Hiltner L (1904) Über neuere Erfahrungen und Probleme auf dem Gebiete der Bodenbakteriologie und unter besonderer Berücksichtigung der Gründungung und Brache. Arb Deut Landw Gesell 98:59–78
Hollander M, Wolfe DA (1973) Nonparametric statistical methods. John Wiley & Sons, New York, pp. 115–120
Huss H, Mavridis A (2007) Bakterium Pseudomonas viridiflava: Neue Blattfleckenkrankheit am Steirischen Ölkürbis. Der Pflanzenarzt 60:8–9
Johnston-Monje D, Lundberg DS, Lazarovits G, Reis VM, Raizada MN (2016) Bacterial populations in juvenile maize rhizospheres originate from both seed and soil. Plant Soil 405:337–355. doi:10.1007/s11104-016-2826-0
Klaedtke S, Jacques MA, Raggi L, Préveaux A, Bonneau S, Negri V, Chable V, Barret M (2016) Terroir is a key driver of seed-associated microbial assemblages. Environ Microbiol 18(6):1792–1804. doi:10.1111/1462-2920.12977
Krzywinski MI, Schein JE, Birol I, Connors J, Gascoyne R, Horsman D, Jones SJ, Marra MA (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19:1639–1645. doi:10.1101/gr.092759.109
Lundberg DS, Lebeis SL, Paredes SH, Yourstone S, Gehring J, Malfatti S, Tremblay J, Engelbrektson A, Kunin V, del Rio TG, Edgar RC, Eickhorst T, Ley RE, Hugenholtz P, Tringe SG, Dangl JL (2012) Defining the core Arabidopsis thaliana root microbiome. Nature 488(7409):86–90. doi:10.1038/nature11237
Lundberg DS, Yourstone S, Mieczkowski P, Jones CD, Dangl JL (2013) Practical innovations for high-throughput amplicon sequencing. Nat Methods 10:999–1002. doi:10.1038/nmeth.2634
Mendes R, Kruijt M, de Bruijn I, Dekkers E, van der Voort M, Schneider JHM, Piceno YM, DeSantis TZ, Andersen GL, Bakker PAHM, Raaijmakers JM (2011) Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332(6033):1097–1100. doi:10.1126/science.1203980
Minchin PR (1987) An evaluation of relative robustness of techniques for ecological ordinations. Vegetatio 69(1):89–107. doi:10.1007/BF00038690
Nelson EB (2004) Microbial dynamics and interactions in the spermosphere. Annu Rev Phytopathol 42:271–309. doi:10.1146/annurev.phyto.42.121603.131041
Neupane A, Tamang P, Brueggeman RS, Friesen TL (2015) Evaluation of a barley core collection for spot form net blotch reaction reveals distinct genotype-specific pathogen virulence and host susceptibility. Phytopathology 105:509–517. doi:10.1094/PHYTO-04-14-0107-R
Niks RE, Parlevliet JE, Lindhout P, Bai Y (2011) Breeding crops with resistance to diseases and pests. Wageningen Academic Publishers. doi:10.3920/978-90-8686-171-2
Pachner M, Paris HS, Winkler J, Lelley T (2015) Phenotypic and marker-assisted pyramiding of genes for resistance to zucchini yellow mosaic virus in oilseed pumpkin (Cucurbita pepo). Plant Breed 134(1):121–128. doi:10.1111/pbr.12219
Peiffer JA, Ley RE (2013) Exploring the maize rhizosphere microbiome in the field: a glimpse into a highly complex system. Commun Integr Biol 6(5):e25177. doi:10.4161/cib.25177
Philippot L, Raaijmakers JM, Lemanceau P, van der Putten WH (2013) Going back to the roots: the microbial ecology of the rhizosphere. Nat Rev Microbiol 11:789–799. doi:10.1038/nrmicro3109
Price MN, Dehal PS, Arkin AP (2010) FastTree 2-approximately maximum-likelihood trees for large alignments. PLoS One 5:e9490. doi:10.1371/journal.pone.0009490
Roane WC (1973) Trends in breeding for disease resistance in crops. Annu Rev Phytopathol 11:463–486
Robinson RW, Decker-Walters DS (1997) Cucurbits. CAB International, New York
Rovira AD (1956) Plant root excretions in relation to the rhizosphere effect. Plant Soil 7:178–194. doi:10.1007/BF01343726
RStudio Team (2015) RStudio: integrated development for R. RStudio, Inc., Boston, MA. http://www.rstudio.com/. Accessed 26 April 2016
Rubiales D, Niks RE (1996) Avoidance of rust infection by some genotypes of Hordeum chilense due to their relative inability to induce the formation of appressoria. Physiol Mol Plant Pathol 49:89–101
Sachs L (1997) Angewandte Statistik. Springer, Berlin 395-397:662–664
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. doi:10.1101/gr.1239303
Shrestha A, Kim BS, Parka DH (2014) Biological control of bacterial spot disease and plant growth-promoting effects of lactic acid bacteria on pepper. Biocontrol Sci Tech 24(7):763–779. doi:10.1080/09583157.2014.894495
Smalla K, Wieland G, Buchner A, Zock A, Parzy J, Kaiser S, Roskot N, Heuer H, Berg G (2001) Bulk and rhizosphere soil bacterial communities studied by denaturing gradient gel electrophoresis: plant-dependent enrichment and seasonal shifts revealed. Appl Environ Microbiol 67:4742–4751. doi:10.1128/AEM.67.10.4742-4751.2001
Teppner H (2004) Notes on Lagenaria and Cucurbita (Cucurbitaceae) – review and new contributions. Phyton (Horn, Austria) 44(2):245–308
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. AEM 73:5261–5267. doi:10.1128/AEM.00062-07
Winkler J, Freistetter B, Huss H (2008) Sortenabhängige und zeitliche Entwicklung von Fruchtfäule bei Ölkürbis (C. pepo var. styriaca). 59. Tagung der Vereinigung der Pflanzenzüchter und Saatgutkaufleute Österreichs. 137–138
Wissuwa M, Mazzola M, Picard C (2009) Novel approaches in plant breeding for rhizosphere related traits. Plant Soil 321(1):409–430. doi:10.1007/s11104-008-9693-2
Zilber-Rosenberg I, Rosenberg E (2008) Role of microorganisms in the evolution of animals and plants: the hologenome theory of evolution. FEMS Microbiol Rev 32:723–735. doi:10.1111/j.1574-6976.2008.00123.x
Zitter T, Hopkins D, Thomas C (1996) Compendium of cucurbit diseases. The American Phytopathological Society, Minnesota/USA
Acknowledgments
Open access funding provided by Graz University of Technology. The authors gratefully acknowledge financial support from funding of the Austrian Research Promotion Agency FFG (project no. 836466). We thank Saatzucht Gleisdorf GmbH and Alwera AG for sharing their expertise, providing seeds and the field site, as well as Barbara Fetz, Monika Schneider-Trampitsch and Alexander Fuchs for their help with sampling and Timothy Mark for English revision.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Philippe Simoneau.
Eveline Adam and Maria Bernhart contributed equally to this work.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Adam, E., Bernhart, M., Müller, H. et al. The Cucurbita pepo seed microbiome: genotype-specific composition and implications for breeding. Plant Soil 422, 35–49 (2018). https://doi.org/10.1007/s11104-016-3113-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-016-3113-9