Abstract
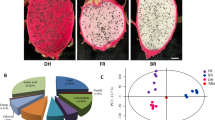
Pomegranate (Punica granatum L.) which belongs to family Lythraceae, is one of the most important fruit crops of many tropical and subtropical regions. A high variability in fruit color is observed among different pomegranate accessions, which arises from the qualitative and quantitative differences in anthocyanins. However, the mechanism of fruit color variation is still not fully elucidated. In the present study, we investigated the red color mutation between a red-skinned pomegranate ‘Hongbaoshi’ and a purple-red-skinned cultivar ‘Moshiliu’, by using transcriptomic and metabolomic approaches. A total of 51 anthocyanins were identified from fruit peels, among which 3-glucoside and 3,5-diglucoside of cyanidin (Cy), delphinidin (Dp), and pelargonidin (Pg) were dominant. High proportion of Pg in early stages of ‘Hongbaoshi’ but high Dp in late stages of ‘Moshiliu’ were characterized. The unique high levels of Cy and Dp anthocyanins accumulating from early developmental stages accounted for the purple-red phenotype of ‘Moshiliu’. Transcriptomic analysis revealed an early down-regulated and late up-regulated of anthocyanin-related structure genes in ‘Moshiliu’ compared with ‘Hongbaoshi’. Alao, ANR was specially expressed in ‘Hongbaoshi’, with extremely low expression levels in ‘Moshiliu’. For transcription factors R2R3-MYB, the profiles demonstrated a much higher transcription levels of three subgroup (SG) 5 MYBs and a sharp decrease in expression of SG6 MYB LOC116202527 in high-anthocyanin ‘Moshiliu’. SG4 MYBs exhibited two entirely different patterns, LOC116203744 and LOC116212505 were down-regulated whereas LOC116205515 and LOC116212778 were up-regulated in ‘Moshiliu’ pomegranate. The results indicate that specific SG members of the MYB family might promote the peel coloration in different manners and play important roles in color mutation in pomegranate.
Key message
Integrating metabolomic and transcriptomic data identified differentially anthocyanins, expressed structural genes and R2R3-MYB genes from SG4, SG5, and SG6, revealing the color variation in red and purple-red pomegranate.







Similar content being viewed by others
Data availability
The RNA-seq data in this study have been deposited into the NCBI Sequnece Read Archive (SRA) database under the BioProject with accession number PRJNA952822 for ‘Hongbaoshi’ and PRJNA990934 for ‘Moshiliu’.
References
Allan AC, Espley RV (2018) MYBs drive novel consumer traits in fruits and vegetables. Trends Plant Sci 23:693–705. https://doi.org/10.1016/j.tplants.2018.06.001
Anders S, Pyl PT, Huber W (2015) HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31:166–169. https://doi.org/10.1093/bioinformatics/btu638
Arlotta C, Puglia GD, Genovese C, Toscano V, Karlova R, Beekwilder J, De Vos RCH, Raccuia SA (2020) MYB5-like and bHLH influence flavonoid composition in pomegranate. Plant Sci 298:110563. https://doi.org/10.1016/j.plantsci.2020.110563
Attanayake R, Rajapaksha R, Weerakkody P, Bandaranayake PG (2019) The effect of maturity status on biochemical composition, antioxidant activity, and anthocyanin biosynthesis gene expression in a pomegranate (Punica granatum L.) cultivar with red flowers, yellow peel, and pinkish arils. J Plant Growth Regul 38:992–1006. https://doi.org/10.1007/s00344-018-09909-2
Balík J, Kumšta M, Rop O (2013) Comparison of anthocyanins present in grapes of Vitis vinifera L. varieties and interspecific hybrids grown in the Czech Republic. Chem Zvesti 67:1285–1292. https://doi.org/10.2478/s11696-013-0378-9
Ban Y, Honda C, Hatsuyama Y, Igarashi M, Bessho H, Moriguchi T (2007) Isolation and functional analysis of a MYB transcription factor gene that is a key regulator for the development of red coloration in apple skin. Plant Cell Physiol 48:958–970. https://doi.org/10.1093/pcp/pcm066
Bar-Ya’akov I, Tian L, Amir R, Holland D (2019) Primary metabolites, anthocyanins, and hydrolyzable tannins in the pomegranate fruit. Front Plant Sci 10:620. https://doi.org/10.3389/fpls.2019.00620
Barnes JS, Nguyen HP, Shen S, Schug KA (2009) General method for extraction of blueberry anthocyanins and identification using high performance liquid chromatography-electrospray ionization-ion trap-time of flight-mass spectrometry. J Chromatogr A 1216:4728–4735. https://doi.org/10.1016/j.chroma.2009.04.032
Ben-Simhon Z, Judeinstein S, Nadler-Hassar T, Trainin T, Bar-Ya’akov I, Borochov-Neori H, Holland D (2011) A pomegranate (Punica granatum L.) WD40-repeat gene is a functional homologue of Arabidopsis TTG1 and is involved in the regulation of anthocyanin biosynthesis during pomegranate fruit development. Planta 234:865–881. https://doi.org/10.1007/s00425-011-1438-4
Ben-Simhon Z, Judeinstein S, Trainin T, Harel-Beja R, Bar-Ya’akov I, Borochov-Neori H, Holland D (2015) A “White” anthocyanin-less pomegranate (Punica granatum L.) caused by an insertion in the coding region of the leucoanthocyanidin dioxygenase (LDOX; ANS) gene. PLoS ONE 10:e0142777. https://doi.org/10.1371/journal.pone.0142777
Câmara JS, Locatelli M, Pereira JAM, Oliveira H, Arlorio M, Fernandes I, Perestrelo R, Freitas V, Bordiga M (2022) Behind the scenes of anthocyanins-from the health benefits to potential applications in food, pharmaceutical and cosmetic fiels. Nutrients 14:5133. https://doi.org/10.3390/nu14235133
Cappellini F, Marinelli A, Toccaceli M, Tonelli C, Petroni K (2021) Anthocyanins: from mechanisms of regualtion in plants to health benefits in foods. Front Plant Sci 12:748049. https://doi.org/10.3389/fpls.2021.748049
Chaves-Silva S, dos Santos AL, Chalfun-Júnior A, Zhao J, Peres LEP, Benedito VA (2018) Understanding the genetic regulation of anthocyanin biosynthesis in plants– tools for breeding purple varieties of fruits and vegetables. Phytochemistry 153:11–27. https://doi.org/10.1016/j.phytochem.2018.05.013
Chen L, Hu B, Qin Y, Hu G, Zhao J (2019) Advance of the negative regulation of anthocyanin biosynthesis by MYB transcription factors. Plant Physiol Biochem 136:178–187. https://doi.org/10.1016/j.plaphy.2019.01.024
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13:1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Deluc L, Bogs J, Walker AR, Ferrier T, Decendit A, Merillon JM, Robinson SP, Barrieu F (2008) The transcription factor VvMYB5b contributes to the regulation of anthocyanin and proanthocyanidin biosynthesis in developing grape berries. Plant Physiol 147:2041–2053. https://doi.org/10.1104/pp.108.118919
Dubos C, Stracke R, Grotewold E, Weisshaar B, Martin C, Lepiniec L (2010) MYB transcription factors in Arabidopsis. Trends Plant Sci 15:573–581. https://doi.org/10.1016/j.tplants.2010.06.005
Espley RV, Hellens RP, Putterill J, Stevenson DE, Kutty-Amma S, Allan AC (2007) Red colouration in apple fruit is due to the activity of the MYB transcription factor, MdMYB10. Plant J 49:414–427. https://doi.org/10.1111/j.1365-313X.2006.02964.x
Fahmy HA, Farag MA (2021) Ongoing and potential novel trends of pomegranate fruit peel; a comprehensive review of its health benefits and future perspectives as nutraceutical. J Food Biochem 46:e14024. https://doi.org/10.1111/jfbc.14024
Fan M, Li X, Zhang Y, Yang M, Wu S, Yin H, Liu W, Fan Z, Li J (2023) Novel insight into anthocyanin metabolism and molecular characterization of its key regulators in Camellia sasanqua. Plant Mol Biol 111:249–262. https://doi.org/10.1007/s11103-022-01324-2
Fellah B, Bannour M, Rocchetti G, Lucini L, Ferchichi A (2018) Phenolic profiling and antioxidant capacity in flowers, leaves and peels of Tunisian cultivars of Punica granatum L. J Food Sci Technol 55:3606–3615. https://doi.org/10.1007/s13197-018-3286-8
Feng S, Wang Y, Song Y, Xu Y, Chen X (2010) Anthocyanin biosynthesis in pears is regulated by a R2R3-MYB transcription factor PyMYB10. Planta. 232:245–255. https://doi.org/10.1007/s00425-010-1170-5
Harel-Beja R, Tian L, Freilich S, Habashi R, Borochov-Neori H, Lahav T, Trainin T, Doron-Faigenboim A, Ophir R, Bar-Ya’akov I, Amir R, Holland D (2019) Gene expression and metabolite profiling analyses of developing pomegranate fruit peel reveal interactions between anthocyanin and punicalagin production. Tree Genet Genomes 15:22. https://doi.org/10.1007/s11295-019-1329-6
He J, Liu Y, Pan Q, Cui X, Duan C (2010) Different anthocyanin profiles of the skin and the pulp of Yan73 (Muscat Hamburg × Alicante Bouschet) grape berries. Molecules 15:1141–1153. https://doi.org/10.3390/molecules15031141
Hernández F, Melgarejo P, Tomás-Barberán FA, Artés F (1999) Evolution of juice anthocyanins during ripening of new selected pomegranate (Punica granatum) clones. Eur Food Res Technol 210:39–42. https://doi.org/10.1007/s002170050529
Hichri I, Barrieu F, Bogs J, Kappel C, Delrot S, Lauvergeat V (2011) Recent advances in the transcriptional regulation of the flavonoid biosynthetic pathway. J Exp Bot 62:2465–2483. https://doi.org/10.1093/jxb/erq442
Huang Z, Wang B, Williams P, Pace RD (2009) Identification of anthocyanins in muscadine grapes with HPLC-ESI-MS. LWT-Food Sci Technol 42:819–824. https://doi.org/10.1016/j.lwt.2008.11.005
Jin J, Tian F, Yang D, Meng Y, Kong L, Luo J, Gao G (2017) PlantTFDB 4.0: toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res 45:D1040–D1045. https://doi.org/10.1093/nar/gkw982
Kaur R, Kapoor N, Aslam L, Mahajan R (2019) Molecular characterization of PgUFGT gene and R2R3-PgMYB transcription factor involved in flavonoid biosynthesis in four tissues of wild pomegranate (Punica granatum L). J Genet 98:94. https://doi.org/10.1007/s12041-019-1141-y
Khaksar G, Tabatabaei BES, Arzani A, Ghobadi C, Ebrahimie E (2015) Functional analysis of a pomegranate (Punica granatum L.) MYB transcription factor involved in the regulation of anthocyanin biosynthesis. Iran J Biotechnol 13:17–25. https://doi.org/10.15171/ijb.1045
Kim E, Hyun TK (2023) PlgMYBR1, an R2R3-MYB transcription factor, plays as a negative regulator of anthocyanin biosynthesis in Platycodon grandiflorus. 3 Biotech 13: 75. https://doi.org/10.1007/s13205-023-03490-6
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357–360. https://doi.org/10.1038/nmeth.3317
Lafferty DJ, Espley RV, Deng CH, Günther CS, Plunkett B, Turner JL, Jaakola L, Karppinen K, Allan AC, Albert NW (2022) Hierarchical regulation of MYBPA1 by anthocyanin- and proanthocyanidin-related MYB proteins is coserved in Vaccinium species. J Exp Bot 73:1344–1356. https://doi.org/10.1093/jxb/erab460
Li Y, Shan X, Zhou L, Gao R, Yang S, Wang S, Wang L, Gao X (2019) The R2R3-MYB factor FhMYB5 from Freesia Hybrida contributes to the regulation of anthocyanin and proanthocyanidin biosynthesis. Front Plant Sci 9:1935. https://doi.org/10.3389/fpls.2018.01935
Li S, Zhang Y, Shi L, Cao S, Chen W, Yang Z (2023) Involvement of a MYB transcription factor in anthocyanin biosynthesis during Chinese bayberry (Morella rubra) fruit ripening. Horticulturae 12:894. https://doi.org/10.3390/biology12070894
Liu W, Feng Y, Yu S, Fan Z, Li X, Li J, Yin H (2021) The flavonoid biosynthesis network in plants. Int J Mol Sci 22:12824. https://doi.org/10.3390/ijms222312824
Liu X, Zhai Y, Liu J, Xue J, Markovic T, Wang S, Zhang X (2023) Comparative transcriptome sequencing analysis to postulate the scheme of regulated leaf coloration in Perilla frutescens. Plant Mol Biol 112:119–142. https://doi.org/10.1007/s11103-023-01342-8
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2)−ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Lloyd A, Brockman A, Aguirre L, Campbell A, Bean A, Cantero A, Gonzalez A (2017) Advances in the MYB-bHLH-WD repeat (MBW) pigment regulatory model: addition of a WRKY factor and co-option of an anthocyanin MYB for betalain regulation. Plant Cell Physiol 58:1431–1441. https://doi.org/10.1093/pcp/pcx075
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17:10–12. https://doi.org/10.14806/ej.17.1.200
Miller JC, Chezem WR, Clay NK (2016) Ternary WD40 repeat-containing protein complexes: evolution, composition and roles in plant immunity. Front Plant Sci 6:1108. https://doi.org/10.3389/fpls.2015.01108
Ono NN, Britton MT, Fass JN, Nicolet CM, Lin D, Tian L (2011) Exploring the transcriptome landscape of pomegranate fruit peel for natural product biosynthetic gene and SSR marker discovery. J Integr Plant Biol 53:800–813. https://doi.org/10.1111/j.1744-7909.2011.01073.x
Peng Y, Lin-Wang K, Cooney JM, Wang T, Espley RV, Allan AC (2019) Differential regulation of the anthocyanin profile in purple kiwifruit (Actinidia species). Hort Res 6:3. https://doi.org/10.1038/s41438-018-0076-4
Peng Y, Thrimawithana AH, Cooney JM, Jensen DJ, Espley RV, Allan AC (2020) The proanthocyanin-related transcription factors MYBC1 and WRKY44 regulated branch points in the kiwifruit anthocyanin pathway. Sci Rep 10:14161. https://doi.org/10.1038/s41598-020-70977-0
Rivera-López J, Ordorica-Falomir C, Wesche-Ebeling P (1999) Changes in anthocyanin concentration in Lychee (Litchi chinensis Sonn.) pericarp during maturation. Food Chem 65:195–200. https://doi.org/10.1016/S0308-8146(98)00195-2
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140. https://doi.org/10.1093/bioinformatics/btp616
Rouholamin S, Zahedi B, Nazarian-Firouzabadi F, Saei A (2015) Expression analysis of anthocyanin biosynthesis key regulatory genes involved in pomegranate (Punica granatum L). Sci Hortic 186:84–88. https://doi.org/10.1016/j.scienta.2015.02.017
Russo M, Fanali C, Tripodo G, Dugo P, Muleo R, Dugo L, De Gara L, Mondello L (2018) Analysis of phenolic compounds in different parts of pomegranate (Punica granatum) fruit by HPLC-PDA-ESI/MS and evaluation of their antioxidant activity: application to different Italian varieties. Anal Bioanal Chem 410:3507–3520. https://doi.org/10.1007/s00216-018-0854-8
Salehi B, Sharifi-Rad J, Cappellini F, Reiner Z, Zorzan D, Imran M, Sener B, Kilic M, El-Shazly M, Fahmy NM, Al-Sayed E, Martorell M, Tonelli C, Petroni K, Docea A O, Calina D, Maroyi A (2020) The therapeutic potential of anthocyanins: current approaches based on their molecular mechanism of action. Front Pharmacol 11: 1300. https://doi.org/10.3389/fphar.2020.01300
Shen N, Wang T, Gan Q, Liu S, Wang L, Jin B (2022) Plant flavonoids: classification, distribution, biosynthesis, and antioxidant activity. Food Chem 383:132531. https://doi.org/10.1016/j.foodchem.2022.132531
Shu C, Wu S, Li H, Tian J (2023) Health benefits of anthocyanin-containing foods, beverages, and supplelments have unpredictable relation to gastrointestinal microbiota: a systematic review and meta-analysis of random clinical trials. Nutr Res 116:48–59. https://doi.org/10.1016/j.nutres.2023.04.002
Singh SP, Pal RK, Saini MK, Singh J, Gaikwad N, Parashuram S, Kaur C (2019) Targeted metabolite profiling to gain chemometric insight into Indian pomegranate cultivars and elite germplasm. J Sci Food Agric 99:5073–5082. https://doi.org/10.1002/jsfa.9751
Stover E, Mercure EW (2007) The pomegranate: a new look at the fruit of paradise. HortScience 42:1088–1092. https://doi.org/10.21273/hortsci.42.5.1088
Stracke R, Werber M, Weisshaar B (2001) The R2R3-MYB gene family in Arabidopsis thaliana. Curr Opin Plant Biol 4:447–456. https://doi.org/10.1016/s1369-5266(00)00199-0
Sun C, Wang C, Zhang W, Liu S, Wang W, Yu X, Song T, Yu M, Yu W, Qu S (2021) The R2R3-type MYB transcription factor MdMYB90-like is responsible for the enhanced skin color of an apple bud sport mutant. Hort Res 8:156. https://doi.org/10.1038/s41438-021-00590-3
Sunil L, Shetty NP (2022) Biosynthesis and regulation of anthocyanin pathway genes. Appl Microbiol Biotechnol 106:1783–1798. https://doi.org/10.1007/s00253-022-11835-z
Trainin T, Harel-Beja R, Bar-Ya’akov I, Ben-Simhon Z, Yahalomi R, Borochov-Neori H, Ophir R, Sherman A, Doron-Faigenboim A, Holland D (2021) Fine mapping of the “black” peel color in pomegranate (Punica granatum L.) strongly suggests that a mutation in the Anthocyanidin Reductase (ANR) gene is responsible for the trait. Front Plant Sci 12. https://doi.org/10.3389/fpls.2021.642019
Tuan PA, Bai S, Yaegaki H, Tamura T, Hihara S, Moriguchi T, Oda K (2015) The crucial role of PpMYB10.1 in anthocyanin accumulation in peach and relationships between its allelic type and skin color phenotype. BMC Plant Biol 15:280. https://doi.org/10.1186/s12870-015-0664-5
Walker AR, Lee E, Bogs J, McDavid DAJ, Thomas MR, Robinson SP (2007) White grapes arose through the mutation of two similar and adjacent regulatory genes. Plant J 49:772–785. https://doi.org/10.1111/j.1365-313X.2006.02997.x
Wang S, Zhang X, Li B, Zhao X, Shen Y, Yuan Z (2022) Genome-wide identification and characterization of bZIP gene family and cloning of candidate genes for anthocyanin biosynthesis in pomegranate (Punica granatum). BMC Plant Biol 22:170. https://doi.org/10.1186/s12870-022-03560-6
Xie S, Lei Y, Chen H, Li J, Chen H, Zhang Z (2020) R2R2-MYB transcription factors regulate anthocyanin biosynthesis in grapevine vegetative tissues. Front Plant Sci 11:527. https://doi.org/10.3389/fpls.2020.00527
Xu F, Ning Y, Zhang W, Liao Y, Li L, Cheng H, Cheng S (2014) An R2R3-MYB transcription factor as a negative regulator of the flavonid biosynthesis pathway in Ginkgo biloba. Funct Integr Genomics 14: 177–189. https://doi.org/10.1007/s10142-013-0352-1
Xu W, Dubos C, Lepiniec L (2015) Transcriptional control of flavonoid biosynthesis by MYB-bHLH-WDR complexes. Trends Plant Sci 20:176–185. https://doi.org/10.1016/j.tplants.2014.12.001
Yan H, Pei X, Zhang H, Li X, Zhang X, Zhao M, Chiang VL, Sederoff RR, Zhao X (2021) MYB-mediated regulation of anthocyanin biosynthesis. Int J Mol Sci 22:3103. https://doi.org/10.3390/ijms22063103
Yoshida K, Ma D, Constabel CP (2015) The MYB182 protein down-regulates proanthocyanidin and anthocyanin biosynthesis in poplar by repressing both structural and regulatory flavonoid genes. Plant Physiol 167:693–710. https://doi.org/10.1104/pp.114.253674
Yuan Y, Chiu LW, Li L (2009) Transcriptional regulation of anthocyanin biosynthesis in red cabbage. Planta 230:1141–1153. https://doi.org/10.1007/s00425-009-1013-4
Yuan Z, Fang Y, Zhang T, Fei Z, Han F, Liu C, Liu M, Xiao W, Zhang W, Wu S, Zhang M, Ju Y, Xu H, Dai H, Liu Y, Chen Y, Wang L, Zhou J, Guan D, Yan M, Xia Y, Huang X, Liu D, Wei H, Zheng H (2018) The pomegranate (Punica granatum L.) genome provides insights into fruit quality and ovule developmental biology. Plant Biotechnol J 16:1363–1374. https://doi.org/10.1111/pbi.12875
Zhang P, Zhu H (2023) Anthocyanins in plant food: current status, genetic modification, and future perspectives. Molecules 28:866. https://doi.org/10.3390/molecules28020866
Zhang X, Wang W, Li J, Ding Y, Tian JL, Wang Z, Xiong B, Xu T, Kou G, Zheng Y, Zhong Z, Wang Y, Deng Q, Liang D, Deng H, Liao L (2023) Analysis of anthocyanin accumulation and related gene expression during fig fruit development. Plant Mol Biol Rep 41:317–332. https://doi.org/10.1007/s11105-023-01370-0
Zhao X, Yuan Z (2021) Anthocyanins from pomegranate (Punica grantum L.) and their role in antioxidant capacities in vitro. Chem Biodivers 18:e2100399. https://doi.org/10.1002/cbdv.202100399
Zhao X, Yuan Z, Fang Y, Yin Y, Feng L (2013) Characterization and evaluation of major anthocyanins in pomegranate (Punica granatum L.) peel of different cultivars and their development phases. Eur Food Res Technol 236:109–117. https://doi.org/10.1007/s00217-012-1869-6
Zhao X, Yuan Z, Feng L, Fang Y (2015) Cloning and expression of anthocyanin biosynthetic genes in red and white pomegranate. J Plant Res 128:687–696. https://doi.org/10.1007/s10265-015-0717-8
Zhao X, Zhang Y, Long T, Wang S, Yang J (2022) Regulation mechanism of plant pigments biosynthesis: anthocyanins, carotenoids, and betalains. Metabolites 12:871. https://doi.org/10.3390/metabo12090871
Zheng Y, Jiao C, Sun H, Rosli HG, Pombo MA, Zhang P, Banf M, Dai X, Martin GB, Giovannoni JJ, Zhao PX, Rhee SY, Fei Z (2016) iTAK: a program for genome-wide prediction and classification of plant transcription factors, transcriptional regulators, and protein kinases. Mol Plant 9:1667–1670. https://doi.org/10.1016/j.molp.2016.09.014
Acknowledgements
The authors are grateful to Zhaoxiang Hao and Ying Chen of Zaozhuang Pomegranate Research Center for their help with the preparation of pomegranate samples.
Funding
This work was supported by National Natural Science Foundation of China (31901341), the Priority Academic Program Development of Jiangsu High Education Institutions (PAPD), and China Scholarship Council (202208320188).
Author information
Authors and Affiliations
Contributions
XZ and YZ conceived and designed the experimens; XZ and YF performed the experiments; DK conducted the data analysis; XZ wrote the manuscript; YT and YZ revised the manuscript. All authos have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have not disclosed any competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, X., Feng, Y., Ke, D. et al. Comparative transcriptomic and metabolomic profiles reveal fruit peel color variation in two red pomegranate cultivars. Plant Mol Biol 114, 51 (2024). https://doi.org/10.1007/s11103-024-01446-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11103-024-01446-9