Abstract
Key message
The induction of miR399 and miR398 and the inhibition of miR156, miR159, miR160, miR171, miR2111, and miR2643 were observed under Pi deficiency in alfalfa. The miRNA-mediated genes involved in basic metabolic process, root and shoot development, stress response and Pi uptake.
Abstract
Inorganic phosphate (Pi) deficiency is known to be a limiting factor in plant development and growth. However, the underlying miRNAs associated with the Pi deficiency-responsive mechanism in alfalfa are unclear. To elucidate the molecular mechanism at the miRNA level, we constructed four small RNA (sRNA) libraries from the roots and shoots of alfalfa grown under normal or Pi-deficient conditions. In the present study, alfalfa plants showed reductions in biomass, photosynthesis, and Pi content and increases in their root-to-shoot ratio and citric, malic, and succinic acid contents under Pi limitation. Sequencing results identified 47 and 44 differentially expressed miRNAs in the roots and shoots, respectively. Furthermore, 909 potential target genes were predicted, and some targets were validated by RLM-RACE assays. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses showed prominent enrichment in signal transducer activity, binding and basic metabolic pathways for carbohydrates, fatty acids and amino acids; cellular response to hormone stimulus and response to auxin pathways were also enriched. qPCR results verified that the differentially expressed miRNA profile was consistent with sequencing results, and putative target genes exhibited opposite expression patterns. In this study, the miRNAs associated with the response to Pi limitation in alfalfa were identified. In addition, there was an enrichment of miRNA-targeted genes involved in biological regulatory processes such as basic metabolic pathways, root and shoot development, stress response, Pi transportation and citric acid secretion.









Similar content being viewed by others
References
Allen E, Xie Z, Gustafson AM, Carrington JC (2005) microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell 121:207–221. https://doi.org/10.1016/j.cell.2005.04.004
Ameline-Torregrosa C, Wang BB, O’Bleness MS, Deshpande S, Zhu H, Roe B, Young ND, Cannon SB (2007) Identification and characterization of nucleotide-binding site-leucine-rich repeat genes in the model plant Medicago truncatula. Plant Physiol 146:5–21. https://doi.org/10.1104/pp.107.104588
Ames BN (1966) Assay of inorganic phosphate, total phosphate and phosphatases. Method Enzymol 8:115–118
Araujo WL, Ishizaki K, Nunes-Nesi A, Larson TR, Tohge T, Krahnert I, Witt S, Obata T, Schauer N, Graham IA, Leaver CJ, Fernie AR (2010) Identification of the 2-hydroxyglutarate and isovaleryl-CoA dehydrogenases as alternative electron donors linking lysine catabolism to the electron transport chain of Arabidopsis mitochondria. Plant Cell 22:1549–1563. https://doi.org/10.1105/tpc.110.075630
Aravind J, Rinku S, Pooja B, Shikha M, Kaliyugam S, Mallikarjuna MG, Kumar A, Rao AR, Nepolean T (2017) Identification, characterization, and functional validation of drought-responsive microRNAs in subtropical maize inbreds. Front Plant Sci 8:941. https://doi.org/10.3389/fpls.2017.00941
Aung B, Gruber MY, Amyot L, Omari K, Bertrand A, Hannoufa A (2015) Ectopic expression of LjmiR156 delays flowering, enhances shoot branching, and improves forage quality in alfalfa. Plant Biotechnol R 9:379–393. https://doi.org/10.1007/s11816-015-0375-2
Bari R, Datt Pant B, Stitt M, Scheible WR (2006) PHO2, microRNA399, and PHR1 define a phosphate-signaling pathway in plants. Plant Physiol 141:988–999. https://doi.org/10.1104/pp.106.079707
Bazin J, Khan GA, Combier JP, Bustos-Sanmamed P, Debernardi JM, Rodriguez R, Sorin C, Palatnik J, Hartmann C, Crespi M, Lelandais-Briere C (2013) miR396 affects mycorrhization and root meristem activity in the legume Medicago truncatula. Plant J 74:920–934. https://doi.org/10.1111/tpj.12178
Bielecka M, Watanabe M, Morcuende R, Scheible WR, Hawkesford MJ, Hesse H, Hoefgen R (2014) Transcriptome and metabolome analysis of plant sulfate starvation and resupply provides novel information on transcriptional regulation of metabolism associated with sulfur, nitrogen and phosphorus nutritional responses in Arabidopsis. Front Plant Sci 5:805. https://doi.org/10.3389/fpls.2014.00805
Bonnet E, He Y, Billiau K, Van de Peer Y (2010) TAPIR, a web server for the prediction of plant microRNA targets, including target mimics. Bioinformatics 26:1566–1568. https://doi.org/10.1093/bioinformatics/btq233
Boualem A, Laporte P, Jovanovic M, Laffont C, Plet J, Combier JP, Niebel A, Crespi M, Frugier F (2008) MicroRNA166 controls root and nodule development in Medicago truncatula. Plant J 54:876–887. https://doi.org/10.1111/j.1365-313X.2008.03448.x
Branscheid A, Sieh D, Pant BD, May P, Devers EA, Elkrog A, Schauser L, Scheible WR, Krajinski F (2010) Expression pattern suggests a role of miR399 in the regulation of the cellular response to local Pi increase during arbuscular mycorrhizal symbiosis. Mol Plant Microbe Interact 23:915–926. https://doi.org/10.1094/Mpmi-23-7-0915
Brinch-Pedersen H, Sorensen LD, Holm PB (2002) Engineering crop plants: getting a handle on phosphate. Trends Plant Sci 7:118–125. https://doi.org/10.1016/S1360-1385(01)02222-1
Cabeza RA, Liese R, Lingner A, von Stieglitz I, Neumann J, Salinas-Riester G, Pommerenke C, Dittert K, Schulze J (2014) RNA-seq transcriptome profiling reveals that Medicago truncatula nodules acclimate N(2) fixation before emerging P deficiency reaches the nodules. J Exp Bot 65:6035–6048. https://doi.org/10.1093/jxb/eru341
Cakir O, Candar-Cakir B, Zhang B (2016) Small RNA and degradome sequencing reveals important microRNA function in Astragalus chrysochlorus response to selenium stimuli. Plant Biotechnol J 14:543–556. https://doi.org/10.1111/pbi.12397
Chen X (2009) Small RNAs and their roles in plant development. Annu Rev Cell Dev Biol 25:21–44. https://doi.org/10.1146/annurev.cellbio.042308.113417
Chiou TJ, Lin SI (2011) Signaling network in sensing phosphate availability in plants. Annu Rev Plant Biol 62:185–206. https://doi.org/10.1146/annurev-arplant-042110-103849
Chiou TJ, Aung K, Lin SI, Wu CC, Chiang SF, Su CL (2006) Regulation of phosphate homeostasis by microRNA in Arabidopsis. Plant Cell 18:412–421. https://doi.org/10.1105/tpc.105.038943
Cohu CM, Abdel-Ghany SE, Gogolin Reynolds KA, Onofrio AM, Bodecker JR, Kimbrel JA, Niyogi KK, Pilon M (2009) Copper delivery by the copper chaperone for chloroplast and cytosolic copper/zinc-superoxide dismutases: regulation and unexpected phenotypes in an Arabidopsis mutant. Mol Plant 2:1336–1350. https://doi.org/10.1093/mp/ssp084
Dai X, Zhao PX (2011) psRNATarget: a plant small RNA target analysis server. Nucleic Acids Res 39:W155–W159. https://doi.org/10.1093/nar/gkr319
Denisov IG, Makris TM, Sligar SG, Schlichting I (2005) Structure and chemistry of cytochrome P450. Chem Rev 105:2253–2277. https://doi.org/10.1021/cr0307143
Devaiah BN, Nagarajan VK, Raghothama KG (2007) Phosphate homeostasis and root development in Arabidopsis are synchronized by the zinc finger transcription factor ZAT6. Plant Physiol 145:147–159. https://doi.org/10.1104/pp.107.101691
Ellis CM, Nagpal P, Young JC, Hagen G, Guilfoyle TJ, Reed JW (2005) AUXIN RESPONSE FACTOR1 and AUXIN RESPONSE FACTOR2 regulate senescence and floral organ abscission in Arabidopsis thaliana. Development 132:4563–4574. https://doi.org/10.1242/dev.02012
Fan W, Zhang S, Du H, Sun X, Shi Y, Wang C (2014) Genome-wide identification of different dormant Medicago sativa L. MicroRNAs in response to fall dormancy. PLoS One 9:e114612. https://doi.org/10.1371/journal.pone.0114612
Fredeen AL, Raab TK, Rao IM, Terry N (1990) Effects of phosphorus nutrition on photosynthesis in Glycine max (L.) Merr. Planta 181:399–405. https://doi.org/10.1007/BF00195894
Friedlander MR, Mackowiak SD, Li N, Chen W, Rajewsky N (2012) miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucl Acids Res 40:37–52. https://doi.org/10.1093/nar/gkr688
Fujii H, Chiou TJ, Lin SI, Aung K, Zhu JK (2005) A miRNA involved in phosphate-starvation response in Arabidopsis. Curr Biol 15:2038–2043. https://doi.org/10.1016/j.cub.2005.10.016
Fukaki H, Tasaka M (2009) Hormone interactions during lateral root formation. Plant Mol Biol 69:437–449. https://doi.org/10.1007/s11103-008-9417-2
Gao F, Wang N, Li H, Liu J, Fu C, Xiao Z, Wei C, Lu X, Feng J, Zhou Y (2016a) Identification of drought-responsive microRNAs and their targets in Ammopiptanthus mongolicus by using high-throughput sequencing. Sci Rep 6:34601. https://doi.org/10.1038/srep34601
Gao R, Austin RS, Amyot L, Hannoufa A (2016b) Comparative transcriptome investigation of global gene expression changes caused by miR156 overexpression in Medicago sativa. BMC Genom 17:658. https://doi.org/10.1186/s12864-016-3014-6
Gupta S, Kumari M, Kumar H, Varadwaj PK (2017a) Genome-wide analysis of miRNAs and Tasi-RNAs in Zea mays in response to phosphate deficiency. Funct Integr Genom 17:335–351. https://doi.org/10.1007/s10142-016-0538-4
Gupta S, Yadav BS, Raj U, Freilich S, Varadwaj PK (2017b) Transcriptomic analysis of soil grown T. aestivum cv. root to reveal the changes in expression of genes in response to multiple nutrients deficiency. Front Plant Sci 8:1025. https://doi.org/10.3389/fpls.2017.01025
Ha M, Kim VN (2014) Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol 15:509–524. https://doi.org/10.1038/nrm3838
Hernandez G, Ramirez M, Valdes-Lopez O, Tesfaye M, Graham MA, Czechowski T, Schlereth A, Wandrey M, Erban A, Cheung F, Wu HC, Lara M, Town CD, Kopka J, Udvardi MK, Vance CP (2007) Phosphorus stress in common bean: root transcript and metabolic responses. Plant Physiol 144:752–767. https://doi.org/10.1104/pp.107.096958
Hsieh LC, Lin SI, Shih AC, Chen JW, Lin WY, Tseng CY, Li WH, Chiou TJ (2009) Uncovering small RNA-mediated responses to phosphate deficiency in Arabidopsis by deep sequencing. Plant Physiol 151:2120–2132. https://doi.org/10.1104/pp.109.147280
Hu Y, Cash D (2009) Alfalfa management guide for ningxia. In: Cash D (ed) Chap. 1: global status and development trends of alfalfa. United Nations Food and Agriculture Organization, Beijing, pp 1–14
Jagadeeswaran G, Zheng Y, Li YF, Shukla LI, Matts J, Hoyt P, Macmil SL, Wiley GB, Roe BA, Zhang W, Sunkar R (2009) Cloning and characterization of small RNAs from Medicago truncatula reveals four novel legume-specific microRNA families. New Phytol 184:85–98. https://doi.org/10.1111/j.1469-8137.2009.02915.x
Jiang C, Gao X, Liao L, Harberd NP, Fu X (2007) Phosphate starvation root architecture and anthocyanin accumulation responses are modulated by the gibberellin-DELLA signaling pathway in Arabidopsis. Plant Physiol 145:1460–1470. https://doi.org/10.1104/pp.107.103788
Jones-Rhoades MW, Bartel DP, Bartel B (2006) MicroRNAs and their regulatory roles in plants. Annu Rev Plant Biol 57:19–53. https://doi.org/10.1146/annurev.arplant.57.032905.105218
Kuo HF, Chiou TJ (2011) The role of microRNAs in phosphorus deficiency signaling. Plant Physiol 156:1016–1024. https://doi.org/10.1104/pp.111.175265
Lambers H, Finnegan PM, Laliberte E, Pearse SJ, Ryan MH, Shane MW, Veneklaas EJ (2011) Phosphorus nutrition of proteaceae in severely phosphorus-impoverished soils: are there lessons to be learned for future crops? Plant Physiol 156:1058–1066. https://doi.org/10.1104/pp.111.174318
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–359. https://doi.org/10.1038/nmeth.1923
Li X, Zeng R, Liao H (2016) Improving crop nutrient efficiency through root architecture modifications. J Integr Plant Biol 58:193–202. https://doi.org/10.1111/jipb.12434
Li Y, Wan L, Bi S, Wan X, Li Z, Cao J, Tong Z, Xu H, He F, Li X (2017) Identification of drought-desponsive microRNAs from roots and leaves of alfalfa by high-throughput sequencing. Genes. https://doi.org/10.3390/genes8040119
Liang G, He H, Yu DQ (2012) Identification of nitrogen starvation-responsive microRNAs in Arabidopsis thaliana. PLoS One. https://doi.org/10.1371/journal.pone.0048951
Liu J, Versaw WK, Pumplin N, Gomez SK, Blaylock LA, Harrison MJ (2008) Closely related members of the Medicago truncatula PHT1 phosphate transporter gene family encode phosphate transporters with distinct biochemical activities. J Biol Chem 283:24673–24681. https://doi.org/10.1074/jbc.M802695200
Liu MY, Chen WW, Xu JM, Fan W, Yang JL, Zheng SJ (2013a) The role of VuMATE1 expression in aluminium-inducible citrate secretion in rice bean (Vigna umbellata) roots. J Exp Bot 64:1795–1804. https://doi.org/10.1093/jxb/ert039
Liu N, Yang J, Guo S, Xu Y, Zhang M (2013b) Genome-wide identification and comparative analysis of conserved and novel microRNAs in grafted watermelon by high-throughput sequencing. PLoS ONE 8:e57359. https://doi.org/10.1371/journal.pone.0057359
Long RC, Li MN, Kang JM, Zhang TJ, Sun Y, Yang QC (2015) Small RNA deep sequencing identifies novel and salt-stress-regulated microRNAs from roots of Medicago sativa and Medicago truncatula. Physiol Plant 154:13–27. https://doi.org/10.1111/ppl.12266
Lopez-Arredondo DL, Leyva-Gonzalez MA, Gonzalez-Morales SI, Lopez-Bucio J, Herrera-Estrella L (2014) Phosphate nutrition: improving low-phosphate tolerance in crops. Annu Rev Plant Biol 65:95–123. https://doi.org/10.1146/annurev-arplant-050213-035949
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. https://doi.org/10.1186/S13059-014-0550-8
Lynch JP (2011) Root phenes for enhanced soil exploration and phosphorus acquisition: tools for future crops. Plant Physiol 156:1041–1049. https://doi.org/10.1104/pp.111.175414
Ma XF, Wright E, Ge Y, Bell J, Xi Y, Bouton JH, Wang ZY (2009) Improving phosphorus acquisition of white clover (Trifolium repens L.) by transgenic expression of plant-derived phytase and acid phosphatase genes. Plant Sci 176:479–488. https://doi.org/10.1016/j.plantsci.2009.01.001
Magalhaes JV, Liu J, Guimaraes CT, Lana UGP, Alves VMC, Wang YH, Schaffert RE, Hoekenga OA, Pineros MA, Shaff JE, Klein PE, Carneiro NP, Coelho CM, Trick HN, Kochian LV (2007) A gene in the multidrug and toxic compound extrusion (MATE) family confers aluminum tolerance in sorghum. Nat Genet 39:1156–1161. https://doi.org/10.1038/ng2074
Meyers BC, Axtell MJ, Bartel B, Bartel DP, Baulcombe D, Bowman JL, Cao X, Carrington JC, Chen XM, Green PJ, Griffiths-Jones S, Jacobsen SE, Mallory AC, Martienssen RA, Poethig RS, Qi YJ, Vaucheret H, Voinnet O, Watanabe Y, Weigel D, Zhui JK (2008) Criteria for annotation of plant microRNAs. Plant Cell 20:3186–3190. https://doi.org/10.1105/tpc.108.064311
Nguyen GN, Rothstein SJ, Spangenberg G, Kant S (2015) Role of microRNAs involved in plant response to nitrogen and phosphorous limiting conditions. Front Plant Sci 6:629. https://doi.org/10.3389/fpls.2015.00629
Pant BD, Buhtz A, Kehr J, Scheible WR (2008) MicroRNA399 is a long-distance signal for the regulation of plant phosphate homeostasis. Plant J 53:731–738. https://doi.org/10.1111/j.1365-313X.2007.03363.x
Pant BD, Pant P, Erban A, Huhman D, Kopka J, Scheible WR (2015) Identification of primary and secondary metabolites with phosphorus status-dependent abundance in Arabidopsis, and of the transcription factor PHR1 as a major regulator of metabolic changes during phosphorus limitation. Plant Cell Environ 38:172–187. https://doi.org/10.1111/pce.12378
Rausch C, Bucher M (2002) Molecular mechanisms of phosphate transport in plants. Planta 216:23–37. https://doi.org/10.1007/s00425-002-0921-3
Ryan PR, Delhaize E, Jones DL (2001) Function and mechanism of organic anion exudation from plant roots. Annu Rev Plant Biol 52:527–560. https://doi.org/10.1146/annurev.arplant.52.1.527
Ryan PR, Raman H, Gupta S, Horst WJ, Delhaize E (2009) A second mechanism for aluminum resistance in wheat relies on the constitutive efflux of citrate from roots. Plant Physiol 149:340–351. https://doi.org/10.1104/pp.108.129155
Schruff MC, Spielman M, Tiwari S, Adams S, Fenby N, Scott RJ (2006) The AUXIN RESPONSE FACTOR 2 gene of Arabidopsis links auxin signalling, cell division, and the size of seeds and other organs. Development 133:251–261. https://doi.org/10.1242/dev.02194
Secco D, Jabnoune M, Walker H, Shou H, Wu P, Poirier Y, Whelan J (2013) Spatio-temporal transcript profiling of rice roots and shoots in response to phosphate starvation and recovery. Plant Cell 25:4285–4304. https://doi.org/10.1105/tpc.113.117325
Sha A, Chen Y, Ba H, Shan Z, Zhang X, Wu X, Qiu D, Chen S, Zhou X (2012) Identification of Glycine Max MicroRNAs in response to phosphorus deficiency. J Plant Biol 55:268–280. https://doi.org/10.1007/s12374-011-0255-4
Sheng P, Tan J, Jin M, Wu F, Zhou K, Ma W, Heng Y, Wang J, Guo X, Zhang X, Cheng Z, Liu L, Wang C, Liu X, Wan J (2014) Albino midrib 1, encoding a putative potassium efflux antiporter, affects chloroplast development and drought tolerance in rice. Plant Cell Rep 33:1581–1594. https://doi.org/10.1007/s00299-014-1639-y
Shu YJ, Liu Y, Li W, Song LL, Zhang J, Guo CH (2016) Genome-wide investigation of microRNAs and their targets in response to freezing stress in Medicago sativa L. based on high-throughput sequencing. G3-Genes Genom Genet 6:755–765. https://doi.org/10.1534/g3.115.025981
Sorin C, Declerck M, Christ A, Blein T, Ma L, Lelandais-Briere C, Njo MF, Beeckman T, Crespi M, Hartmann C (2014) A miR169 isoform regulates specific NF-YA targets and root architecture in Arabidopsis. New Phytol 202:1197–1211. https://doi.org/10.1111/nph.12735
Srivastava PK, Moturu TR, Pandey P, Baldwin IT, Pandey SP (2014) A comparison of performance of plant miRNA target prediction tools and the characterization of features for genome-wide target prediction. BMC Genom 15:348. https://doi.org/10.1186/1471-2164-15-348
Trull MC, Guiltinan MJ, Lynch JP, Deikman J (1997) The responses of wild-type and ABA mutant Arabidopsis thaliana plants to phosphorus starvation. Plant Cell Environ 20:85–92. https://doi.org/10.1046/j.1365-3040.1997.d01-4.x
Valdés-López O, Yang SS, Aparicio-Fabre R, Graham PH, Reyes JL, Vance CP, Hernández G (2010) MicroRNA expression profile in common bean (Phaseolus vulgaris) under nutrient deficiency stresses and manganese toxicity. New Phytol 187:805–818. https://doi.org/10.1111/j.1469-8137.2010.03320.x
Voinnet O (2009) Origin, biogenesis, and activity of plant microRNAs. Cell 136:669–687. https://doi.org/10.1016/j.cell.2009.01.046
Wang Y, Zhang C, Hao Q, Sha A, Zhou R, Zhou X, Yuan L (2013) Elucidation of miRNAs-mediated responses to low nitrogen stress by deep sequencing of two soybean genotypes. PLoS ONE 8:e67423. https://doi.org/10.1371/journal.pone.0067423
Wen M, Shen Y, Shi SH, Tang T (2012) miREvo: an integrative microRNA evolutionary analysis platform for next-generation sequencing experiments. BMC Bioinform 13:140. https://doi.org/10.1186/1471-2105-13-140
Yan YS, Wang HC, Hamera S, Chen XY, Fang RX (2014) miR444a has multiple functions in the rice nitrate-signaling pathway. Plant J 78:44–55. https://doi.org/10.1111/tpj.12446
Yu N, Niu Q-W, Ng K-H, Chua N-H (2015) The role of miR156/SPLs modules in Arabidopsis lateral root development. Plant J 83:673–685. https://doi.org/10.1111/tpj.12919
Zeng HQ, Zhu YY, Huang SQ, Yang ZM (2010) Analysis of phosphorus-deficient responsive miRNAs and cis-elements from soybean (Glycine max L.). J Plant Physiol 167:1289–1297. https://doi.org/10.1016/j.jplph.2010.04.017
Zhao Y, Chan Z, Gao J, Xing L, Cao M, Yu C, Hu Y, You J, Shi H, Zhu Y, Gong Y, Mu Z, Wang H, Deng X, Wang P, Bressan RA, Zhu JK (2016) ABA receptor PYL9 promotes drought resistance and leaf senescence. Proc Natl Acad Sci USA 113:1949–1954. https://doi.org/10.1073/pnas.1522840113
Zhou ZS, Zeng HQ, Liu ZP, Yang ZM (2012) Genome-wide identification of Medicago truncatula microRNAs and their targets reveals their differential regulation by heavy metal. Plant Cell Environ 35:86–99. https://doi.org/10.1111/j.1365-3040.2011.02418.x
Zhu YY, Zeng HQ, Dong CX, Yin XM, Shen QR, Yang ZM (2010) microRNA expression profiles associated with phosphorus deficiency in white lupin (Lupinus albus L.). Plant Sci 178:23–29. https://doi.org/10.1016/j.plantsci.2009.09.011
Acknowledgements
This work was supported by China Forage and Grass Research System (CARS-34) , and the Agricultural Science and Technology Innovation Program (ASTIP-IAS14).
Author information
Authors and Affiliations
Contributions
ZL, XL and ZT conceived and designed the experiments. ZL, HX, YL, JC, XW, ZL and ZM performed the experiments. ZL analyzed the experimental data and drafted the manuscript. FH, YW and LW revised the draft manuscript. XL and ZT supervised all analyses and revised the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
Flow diagram for the analysis of small RNAs. Supplementary material 1 (TIF 42397 KB)
Supplementary Fig. 2
The distribution of different small RNAs in different chromosome. Supplementary material 2 (TIF 29101 KB)
Supplementary Fig. 3
Percent nucleotide bias of the 20th -24th bases of novel miRNAs from 12 libraries. The brown box indicates guanine, the green box indicates cytosine, the blue box indicates uracil, and the red box indicates adenine. Supplementary material 3 (TIF 57371 KB)
Supplementary Fig. 4
Secondary structures of predicted novel miRNAs. Supplementary material 4 (TIF 55753 KB)
Supplementary Fig. 5
Pearson correlation of miRNA expression between different libraries. Blue indicates that the correlation coefficient is 1. Supplementary material 5 (TIF 42593 KB)
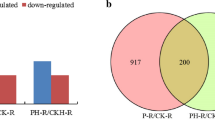
Supplementary Fig. 6
Common and specific differentially expressed miRNAs in roots and shoots under Pi-deficient conditions. (a) Conserved miRNA response to Pi deficiency; (b) Novel miRNA response to Pi deficiency. Supplementary material 6 (TIF 13509 KB)
Rights and permissions
About this article
Cite this article
Li, Z., Xu, H., Li, Y. et al. Analysis of physiological and miRNA responses to Pi deficiency in alfalfa (Medicago sativa L.). Plant Mol Biol 96, 473–492 (2018). https://doi.org/10.1007/s11103-018-0711-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-018-0711-3