ABSTRACT
Purpose
To investigate the protein–protein interactions of a highly concentrated antibody solution that could cause oligomerization or aggregation and to develop a better understanding of the optimization of drug formulations.
Methods
In this study, we used Raman spectroscopy to investigate the structure and interactions of a highly concentrated antibody solution over a wide range of concentrations (10–200 mg/mL) with the aid of a multivariate analysis.
Results
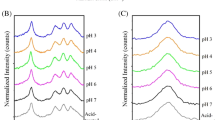
Our analysis of the amide I band, I 856 /I 830 of Tyr, and the relative intensity at 1004 cm−1 of the Phe and OH stretching region at around 3000 cm−1 showed that across this wide range of concentrations, the secondary structure of the IgG molecules did not change; however, short-range attractive interactions around the Tyr and Phe residues occurred as the distance between the IgG molecules decreased with increasing concentration. Analysis of the OH stretching region at around 3000 cm−1 showed that these short-range attractive interactions correlated with the amount of hydrated water around the IgG molecules.
Conclusions
Our data show that Raman spectroscopy can provide valuable information of the protein–protein interactions based on conformational approaches to support conventional colloidal approaches, especially for analyses of highly concentrated solutions.












Similar content being viewed by others
Abbreviations
- MCR-ALS:
-
Multivariate curve resolution-alternating least squares
- PCA:
-
Principal components analysis
- REV:
-
Reduced-eigenvalue
- RMSECV:
-
Root-mean-square prediction error of cross-validation
REFERENCES
Uchiyama S. Liquid formulation for antibody drugs. Biochim Biophys Acta. 1844;2014:2041–52.
Yadav S, Liu J, Shire SJ, Kalonia DS. Specific interactions in high concentration antibody solutions resulting in high viscosity. J Pharm Sci. 2010;99(3):1152–68.
Stradner A, Sedgwick H, Cardinaux F, Poon WCK, Egelhaaf SU, Schurtenberger P. Equilibrium cluster formation in concentrated protein solutions and colloids. Nature. 2004;432:492–5.
Wen Z. Raman spectroscopy of protein pharmaceuticals. J Pharm Sci. 2007;96(10):2861–71.
Oladepo SA, Xiong K, Hong Z, Asher SA, Handen J, Lednev IK. UV resonance raman investigations of peptide and protein structure and dynamics. Chem Rev. 2012;112(5):2604–28.
Ota C, Noguchi S, Tsumoto K. The molecular interaction of a protein in highly concentrated solution investigated by Raman spectroscopy. Biopolymers. 2015;103(4):237–46.
Malinowski ER. Factor analysis in chemistry. 2nd ed. New York: Wiley-Interscience; 1991.
Lobo AP, Valles BS, Tascón NF, Madrera RR. Calibration models for routine analysis of cider by mid-infrared spectroscopy. LWT Food Sci Technol. 2006;39(9):1026–32.
Tauler R. Application of non-linear optimization methods to the estimation of multivariate curve resolution solutions and of their feasible band boundaries in the investigation of two chemical and environmental simulated data sets. Anal Chim Acta. 2007;595(1–2):289–98.
Maiti NC, Apetri MM, Zagorski MG, Carey PR, Anderson VE. Raman spectroscopic characterization of secondary structure in natively unfolded proteins : α-Synuclein. J Am Chem Soc. 2004;126(8):2399–408.
Apetri MM, Maiti NC, Zagorski MG, Carey PR, Anderson VE. Secondary structure of a-synuclein oligomers: characterization by raman and atomic force microscopy. J Mol Biol. 2006;355(1):63–71.
Vinaykin M, Benderskii AV. Vibrational sum-frequency spectrum of the water bend at the air/water interface. J Phys Chem Lett. 2012;3(22):3348–52.
Chen H, Lin H, Chen H, Mai F, Liu Y, Lin C, et al. Innovative strategy with potential to increase hemodialysis efficiency and safety. Sci Rep. 2014;4(4425):1–5.
Shen YR, Ostroverkhov V. Sum-frequency vibrational spectroscopy on water interfaces: polar orientation of water molecules at interfaces. Chem Rev. 2006;106:1140–54.
Davis JG, Gierszal KP, Wang P, Ben-Amotz D. Water structural transformation at molecular hydrophobic interfaces. Nature. 2012;491:582–5.
Englert L, Biela A, Zayed M, Heine A, Hangauer D, Klebe G. Displacement of disordered water molecules from hydrophobic pocket creates enthalpic signature: binding of phosphonamidate to the S1′-pocket of thermolysin. Biochim Biophys Acta. 1800;2010:1192–202.
Phillip WS, Matthew RL, Demetri TM, George MW. Is it the shape of the cavity, or the shape of the water in the cavity? Eur Phys J Special Topics. 2014;223:853–91.
Lord RC, Yu N. Laser excited Raman spectroscopy of biomolecules: I. Native lysozyme and its constituent amino acids. J Mol Biol. 1970;50(2):509–24.
Xu M, Ermolenkov VV, He W, Uversky VN, Fredriksen L, Lednev IK. Lysozyme fibrillation: deep UV Raman spectroscopic characterization of protein structural transformation. Biopolymers. 2005;79:58–61.
Xu M, Shashilov VA, Ermolenkov VV, Fredriksen L, Zagorevski D, Lednev IK. Hen egg white lysozyme fibrillation: a deep UV resonance Raman spectroscopic study. J Biophotonics. 2008;1(3):215–29.
Gelden JD, Gussem KD, Vandenabeele P, Moens L. Reference database of Raman spectra of biological molecules. J Raman Spectrosc. 2007;38:1133–47.
Sarbon NM, Badii F, Howell NK. Preparation and characterization of chicken skin gelatin as an alternative to mammalian gelatin. Food Hydrocoll. 2013;30(1):143–51.
McCammon JA, Wolynes PG, Karplus M. Picosecond dynamics of tyrosine side chains in proteins. Biochemistry. 1979;18:927–42.
Brandl M, Weiss MS, Jabs A, Suhnel J, Hilgenfeld R. C-H π-interactions in proteins. J Mol Biol. 2001;307:357–77.
Siamwiza MN, Lord RC, Chen MC, Takamatsu T, Harada I, Matsuura H, et al. Interpretation of the doublet at 850 and 830 cm−1 in the Raman spectra of Tyrosyl residues in proteins and certain model compounds. Biochemistry. 1975;14(22):4870–6.
Chari R, Jerath K, Badkar AV, Kalonia DS. Long and short range electrostatic interactions affect the rheology of highly concentrated antibody solutions. Pharm Res. 2009;26:2607–18.
Takeuchi H, Harada I. Normal coordinate analysis of the indole ring. Spectrochim Acta. 1986;142A:1069–78.
Miura T, Takeuchi H, Harada I. Characterization of individual tryptophan side chains in proteins using Raman spectroscopy and hydrogen deuterium exchange kinetics. Biochemistry. 1988;27:88–94.
Scherer TM, Liu J, Shire SJ, Minton AP. Intermolecular interactions of IgG1 monoclonal antibodies at high concentrations characterized by light scattering. J Phys Chem B. 2010;114:12948–57.
Saito S, Hasegawa J, Kobayashi N, Tomitsuka T, Uchiyama S, Fukui K. Effects of ionic strength and sugars on the aggregation propensity of monoclonal antibodies: influence of colloidal and conformational stabilities. Pharm Res. 2013;30:1263–80.
Thomas MS. Role of cosolute–protein interactions in the dissociation of monoclonal antibody clusters. J Phys Chem B. 2015;119:13027–38.
ACKNOWLEDGMENTS AND DISCLOSURES
We thank our colleague Dr. Daisuke Irikura (Horiba, Ltd.) for helpful comments on the results and discussion.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
The results of PCA (REV, RMSECV) are supplied as Supporting Information 1,3,5 (Fig. S1,3,5). Concentration dependence of Raman spectra in CH and OH stretching region (Fig. S2), the abstract spectra of MCR-ALS analysis (Fig. S4) and distance between the molecules of the IgG solution as a function of concentration (Fig. S6) are also supplied as Supporting Information 2,4,6. (DOCX 940 kb)
Rights and permissions
About this article
Cite this article
Ota, C., Noguchi, S., Nagatoishi, S. et al. Assessment of the Protein–Protein Interactions in a Highly Concentrated Antibody Solution by Using Raman Spectroscopy. Pharm Res 33, 956–969 (2016). https://doi.org/10.1007/s11095-015-1842-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11095-015-1842-8