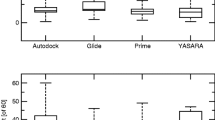
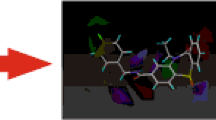
Molecular docking of the extracellular domain of the GABAB1a subunit (whose spatial structure was identified by molecular simulation of a full-size GABAB receptor) with the GABA antagonists phenylethylamine and CGP54626 was examined. The spatial structure of the molecules of the above antagonists was obtained using the PubChem database of chemical compounds and mixtures. Detection and analysis of the interaction sites of phenylethylamine with the extracellular domain of subunit GABAB1а of the GABAB receptor was performed using the SwissDock web-service. Docking results were estimated using the FullFitness parameter and free energy analysis of binding of the antagonists to the receptor. The results of molecular docking of phenylethylamine and CGP54626 to the extracellular domain of subunit GABAB1a of the GABAB receptor, as well as a comparative analysis of these data with the results of molecular docking of baclofen to the mentioned receptor, have demonstrated that all binding sites for both antagonists and agonist, located either under or in the “claws” of the subunit molecule, are completely or partially structured by specific groups of amino acid residues. The sites differ from each other in their molecular volumes and density of filling with the above residues. According to the mentioned estimated parameters, the binding sites for baclofen, phenylethylamine, and CGP54626 located in the claws have higher motility and binding energy than similar sites positioned under the claws. In the latter case, the respective sites have higher “rigidity” and lower motility. It has been revealed that the main principle of forming the binding sites for antagonists is the formation of the main matrix elements of baclofen-binding sites located under the claws or in the latter, respectively, with subsequent creation of new groups of amino acids and/or supplying the existing ones with additional amino acid residues. As for antagonist/agonist binding sites, this process is likely to occur via the hydrogen bonds with the constituents of the former, namely amino acid residues threonine and serine, which are the most reaction-capable ones due to their hydroxylic groups. It has been assumed that one of the possible mechanisms of conformation-dependent impact of baclofen-, phenylethylamine-, and CGP54626-binding sites under the claws on the formation of sites to these compounds in the claws is based on the fact that agonist/antagonist-activated transfers of “rigidly”-packed sites, located under the claws and causing conformational transformations of polypeptide chains, are realized, in general, for greater distances. Stabilization of interaction between such sites is likely to occur via the formation of bonds between cations (positively charged radicals of polar charged amino acids in the sites under the claws) and π-electrons of aromatic rings of nonpolar and polar uncharged amino acids in the sites located in the claws.
Similar content being viewed by others
References
M. J. Jembrek and J. Vlainic, “GABA receptors: pharmacological potential and pitfalls,” Curr. Pharm. Des., 21, No. 34, 4943-4959 (2015).
B. Bettler, K. Kaupmann, J. Mosbacher, and M. Gassmann, “Molecular structure and physiological functions of GABA(B) receptors,” Physiol. Rev., 84, No. 3, 835-867 (2004).
N. G. Bowery and G. D. Pratt, “GABAB receptors as targets for drug action,” Arzneimittelforschung, 42, No. 2A, 215-223 (1992).
A. Bouron, “Modulation of spontaneous quantal release of neurotransmitters in the hippocampus,” Prog. Neurobiol., 63, No. 6, 613-635 (2001).
J. H. White, A. Wise, M. J. Main, et al., “Heterodimerization is required for the formation of a functional GABAB receptor,” Nature, 396, No. 6712, 679-682 (1998).
N.G. Bowery, B. Bettler, W. Froestl, et al., “International Union of Pharmacology. XXXIII. Mammalian gammaaminobutyric acid(B) receptors: structure and function,” Pharmacol. Rev., 54, No. 2, 247-264 (2002).
Y. Geng, M. Bush, L. Mosyak, et al., “Structural mechanism of ligand activation in human GABA(B) receptor,” Nature, 504, No. 7479, 254-259 (2013).
J.P. Pin, J. Kniazeff, V. Binet, et al., “Activation mechanism of the heterodimeric GABA(B) receptor,” Biochem. Pharmacol., 68, No. 8, 1565-1572 (2004).
Y. Geng, D. Xiong, L. Mosyak, et al., “Structure and functional interaction of the extracellular domain of human GABA(B) receptor GBR2,” Nat. Neurosci., 15, No. 7, 970-978 (2012).
J. Liu, D. Maurel, S. Etzol, et al., “Molecular determinants involved in the allosteric control of agonist affinity in the GABAB receptor by the GABAB2 subunit,” J. Biol. Chem., 279, No. 16, 15824-15830 (2004).
P. Rondard, S. Huang, C. Monnier, et al., “Functioning of the dimeric GABA(B) receptor extracellular domain revealed by glycan wedge scanning,” EMBO J., 27, No. 9, 1321-1332 (2008).
B. A. Clark and S. G. Cull-Candy, “Activity-dependent recruitment of extrasynaptic NMDA receptor activation at an AMPA receptor-only synapse,” J. Neurosci., 22, No. 11, 4428-4436 (2002).
J. P. Pin, J. Kniazeff, C. Goudet, et al., “The activation mechanism of class-C G-protein coupled receptors,” Biol. Cell., 96, No. 5, 335-342 (2004).
R. Cossart, M. Esclapez, J. C. Hirsch, et al., “GluR5 kainate receptor activation in interneurons increases tonic inhibition of pyramidal cells,” Nat. Neurosci., 1, No. 6, 470-478 (1998).
R. Cossart, J. Epsztein, R. Tyzio, et al., “Quantal release of glutamate generates pure kainate and mixed AMPA/kainate EPSCs in hippocampal neurons,” Neuron, 35, No. 1, 147-159 (2002).
B. Duthey, S. Caudron, J. Perroy, et al., “A single subunit (GB2) is required for G-protein activation by the heterodimeric GABA(B) receptor,” J. Biol. Chem., 277, No. 5, 3236-3241 (2002).
M. Havlickova, L. Prezeau, B. Duthey, et al., “The intracellular loops of the GB2 subunit are crucial for G-protein coupling of the heteromeric gammaaminobutyrate B receptor,” Mol. Pharmacol., 62, No. 2, 343-350 (2002).
K. A. Jones, B. Borowsky, J. A. Tamm, et al., “GABAB receptors function as a heteromeric assembly of the subunits GABABR1 and GABABR2,” Nature, 396, No. 6712, 674-679 (1998).
H.-C. Kornau, “GABAB receptors and synaptic modulation,” Cell Tissue Res., 326, No. 2, 517-533 (2006).
A. Yu. Nyporko, A. M. Naumenko, A. Golius, et al., “3D reconstruction of a full-size GABAB receptor,” Neurophysiology, 47, No. 5, 364-375 (2015).
A. M. Naumenko, L. M. Shapoval, A. Yu. Nyporko, et al., “Computer simulation of molecular interaction between baclofen and the GABAB receptor,” Neurophysiology, 49, No. 1, 2-7 (2017).
M. D. Berry, “The effects of pargyline and 2-phenylethylamine on D1-like dopamine receptor binding,” J. Neural. Transm., 118, No. 7, 1115-1118 (2011).
R. L. O’Reilly and B. A. Davis, “Phenylethylamine and schizophrenia,” Prog. Neuropsychopharmacol. Biol. Psychiat., 18, No. 1, 63-75 (1994).
F. Brugger, U. Wicki, H. R. Olpe, et al., “The action of new potent GABAB receptor antagonists in the hemisected spinal cord preparation of the rat,” Eur. J. Pharmacol., 235, No. 1, 153-155 (1993).
M. J. Asay and S. K. Boyd, “Characterization of the binding of [3H]CGP54626 to GABAB receptors in the male bullfrog (Rana catesbeiana),” Brain Res., 1094, No. 1, 76-85 (2006).
S. Kim, P. A. Thiessen, E. E. Bolton, et al., “PubChem substance and compound databases,” Nucleic. Acids Res., 44, No. D1, 1202-1213 (2016).
Y. Wang, T. Suzek, J. Zhang, et al., “PubChem BioAssay: 2014 update,” Nucleic. Acids Res., 42(Database issue), D1075–D1082 (2014).
A. Grosdidier, V. Zoete, and O. Michielin, “SwissDock, a protein-small molecule docking web service based on EADock DSS,” Nucleic. Acids Res., 39 (Web Server issue), W270-W277 (2011).
A. Grosdidier, V. Zoete, and O. Michielin, “Fast docking using the CHARMM force field with EADock DSS,” J. Comput. Chem., 32, No. 10, 2149-2159 (2011).
V. Zoete, M. A. Cuendet, A. Grosdidier, and O. Michielin, “SwissParam, a fast force field generation tool for small organic molecules,” J. Comput. Chem., 32, No. 11, 2359-2368 (2011).
R. S. Spolar, J. R. Livingstone, and M. T. Record, Jr., “Use of liquid hydrocarbon and amide transfer data to estimate contributions to thermodynamic functions of protein folding from the removal of nonpolar and polar surface from water,” Biochemistry, 31, No. 16, 3947-3955 (1992).
K. T. О’Neil and W. F. DeGrado, “A thermodynamic scale for the helix-forming tendencies of the commonly occurring amino acids,” Science, 250, No. 4981, 646-651 (1990).
K. A. Dill, “Dominant forces in protein folding,” Biochemistry, 29, No. 31, 7133-7155 (1990).
C. R. Cantor and P. R. Schimmel, Biophysical Chemistry, Part 1. The Conformation of Biological Macromolecules (Their Biophysical Chemistry; pt. 1), W. H. Freeman (Publ.) (1984).
M. Tomita and V. T. Marchesi, “Amino-acid sequence and oligosaccharide attachment sites of human erythrocyte glycophorin,” Proc. Natl. Acad. Sci. USA, 72, No. 8, 2964-2968 (1976).
R. Aurora, T. P. Creamer, R. Srinivasan, and G. D. Rose, “Local interaction in protein folding: lessons from the α-helix,” J. Biol. Chem., 272, No. 3, 1413-1416 (1997).
F. M. Richards, “The interpretation of protein structures: total volume, group volume distributions and packing density,” J. Mol. Biol., 82, No. 1, 1-14 (1974).
W. Kauzmann, K. Moore, and D. Schultz, “Protein densities from X-ray crystallographic coordinates,” Nature, 248, No. 447, 447-449 (1974).
J. P. Gallivan and D. A. Dougherty, “A computational study of cation–π interactions vs. salt bridges in aqueous media: Implications for protein engineering,” J. Am. Chem. Soc., 122, No. 5, 870-874 (2000).
I. N. Mezhevoy and V. G. Badelin, “Dissolution and solvation enthalpies of L-serine in aqueous-alcoholic solutions at 298.15 K,” J. Phys. Chem., 82, No. 4, 789-791 (2008) [in Russian].
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Naumenko, A.M., Shapoval, L.M., Davydovska, T.L. et al. Molecular Docking of Phenylethylamine and CGP54626 to an Extracellular Domain of the GABAB-Receptor. Neurophysiology 50, 230–242 (2018). https://doi.org/10.1007/s11062-018-9743-1
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11062-018-9743-1