Abstract
Objective
Brain metastases (BM) are associated with poor prognosis and increased mortality rates, making them a significant clinical challenge. Studying BMs can aid in improving early detection and monitoring. Systematic comparisons of anatomical distributions of BM from different primary cancers, however, remain largely unavailable.
Methods
To test the hypothesis that anatomical BM distributions differ based on primary cancer type, we analyze the spatial coordinates of BMs for five different primary cancer types along principal component (PC) axes. The dataset includes 3949 intracranial metastases, labeled by primary cancer types and with six features. We employ PC coordinates to highlight the distinctions between various cancer types. We utilized different Machine Learning (ML) algorithms (RF, SVM, TabNet DL) models to establish the relationship between primary cancer diagnosis, spatial coordinates of BMs, age, and target volume.
Results
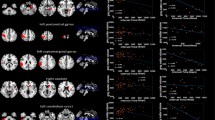
Our findings revealed that PC1 aligns most with the Y axis, followed by the Z axis, and has minimal correlation with the X axis. Based on PC1 versus PC2 plots, we identified notable differences in anatomical spreading patterns between Breast and Lung cancer, as well as Breast and Renal cancer. In contrast, Renal and Lung cancer, as well as Lung and Melanoma, showed similar patterns. Our ML and DL results demonstrated high accuracy in distinguishing BM distribution for different primary cancers, with the SVM algorithm achieving 97% accuracy using a polynomial kernel and TabNet achieving 96%. The RF algorithm ranked PC1 as the most important discriminating feature.
Conclusions
In summary, our results support accurate multiclass ML classification regarding brain metastases distribution.




Similar content being viewed by others
Data Availability
Data that are used in this study are part of the International Radiosurgery Registry Foundation (Study #: IRRF 02-15-2022), & Topography of Brain Metastases by Primary Cancer Genetic Subtypes & Data collection was approved by institutional review boards at each of the institutions affiliated with the IRRF. Due to retrospective design, informed consents were not obtained.
References
Cardinal T, Pangal D, Strickland BA, Newton P, Mahmoodifar S, Mason J, Craig D, Simon T, Tew BY, Yu M et al (2022) Anatomical and topographical variations in the distribution of brain metastases based on primary cancer origin and molecular subtypes: a systematic review. Neuro-Oncol Adv 4(1):170. https://doi.org/10.1093/noajnl/vdab170
In GK, Mason J, Lin S, Newton PK, Kuhn P, Nieva J (2017) Development of metastatic brain disease involves progression through lung metastases in egfr mutated non-small cell lung cancer. Converg Sci Phys Oncol 3(3):035002. https://doi.org/10.1088/2057-1739/aa7a8d
Newton PK, Mason J, Venkatappa N, Jochelson MS, Hurt B, Nieva J, Comen E, Norton L, Kuhn P (2015) Spatiotemporal progression of metastatic breast cancer: a markov chain model highlighting the role of early metastatic sites. NPJ Breast Cancer 1(1):1–9. https://doi.org/10.1038/npjbcancer.2015.18
Newton PK, Mason J, Hurt B, Bethel K, Bazhenova L, Nieva J, Kuhn P (2014) Entropy, complexity and markov diagrams for random walk cancer models. Sci Rep 4(1):7558. https://doi.org/10.1038/srep07558
Newton PK, Mason J, Bethel K, Bazhenova L, Nieva J, Norton L, Kuhn P (2013) Spreaders and sponges define metastasis in lung cancer: a markov chain monte carlo mathematical model. Can Res 73(9):2760–2769. https://doi.org/10.1158/0008-5472.CAN-12-4488
Newton PK, Mason J, Bethel K, Bazhenova LA, Nieva J, Kuhn P (2012) A stochastic markov chain model to describe lung cancer growth and metastasis. PLoS ONE 7(4):34637. https://doi.org/10.1371/journal.pone.0034637
Schroeder T, Bittrich P, Kuhne J, Noebel C, Leischner H, Fiehler J, Schroeder J, Schoen G, Gellisen S (2020) Mapping distribution of brain metastases: does the primary tumor matter? J Neuro-Oncol 147:229–235. https://doi.org/10.1007/s11060-020-03419-6
Neman J, Franklin M, Madaj Z, Deshpande K, Triche TJ, Sadlik G, Carmichael JD, Chang E, Yu C, Strickland BA et al (2021) Use of predictive spatial modeling to reveal that primary cancers have distinct central nervous system topography patterns of brain metastasis. J Neurosurg 136(1):88–96. https://doi.org/10.3171/2021.1.JNS203536
Mahmoodifar S, Pangal DJ, Cardinal T, Craig D, Simon T, Tew BY, Yang W, Chang E, Yu M, Neman J et al (2022) A quantitative characterization of the spatial distribution of brain metastases from breast cancer and respective molecular subtypes. J Neuro-Oncol 160(1):241–251. https://doi.org/10.1007/s11060-022-04147-9
Fidler IJ, Yano S, Zhang R-d, Fujimaki T, Bucana CD (2002) The seed and soil hypothesis: vascularisation and brain metastases. Lancet Oncol 3(1):53–57. https://doi.org/10.1016/s1470-2045(01)00622-2
Fidler IJ (2011) The role of the organ microenvironment in brain metastasis. In: Seminars in cancer biology, vol 21. Elsevier, pp 107–112. https://doi.org/10.1016/j.semcancer.2010.12.009
Kirby M (2000) Geometric data analysis: an empirical approach to dimensionality reduction and the study of patterns. John Wiley & Sons Inc, New York
Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O, Blondel M, Prettenhofer P, Weiss R, Dubourg V et al (2011) Scikit-learn: machine learning in python. J Mach Learn Res 12:2825–2830
Chawla NV, Bowyer KW, Hall LO, Kegelmeyer WP (2002) Smote: synthetic minority over-sampling technique. J Artif Intell Res 16:321–357. https://doi.org/10.1613/jair.953
Lemaître G, Nogueira F, Aridas CK (2017) Imbalanced-learn: a python toolbox to tackle the curse of imbalanced datasets in machine learning. J Mach Learn Res 18(1):559–563
Breiman L (2001) Random forests. Mach Learn 45:5–32
Gupta P, Garg S (2020) Breast cancer prediction using varying parameters of machine learning models. Procedia Comput Sci 171:593–601. https://doi.org/10.1016/j.procs.2020.04.064
Ramaswamy S, Tamayo P, Rifkin R, Mukherjee S, Yeang C-H, Angelo M, Ladd C, Reich M, Latulippe E, Mesirov JP et al (2001) Multiclass cancer diagnosis using tumor gene expression signatures. Proc Natl Acad Sci 98(26):15149–15154. https://doi.org/10.1073/pnas.211566398
He K, Zhang X, Ren S, Sun J (2016) Deep residual learning for image recognition. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 770–778. https://doi.org/10.1109/CVPR.2016.90
Arik SÖ, Pfister T (2021) Tabnet: attentive interpretable tabular learning. In: Proceedings of the AAAI conference on artificial intelligence, vol 35, pp 6679–6687. https://doi.org/10.1609/aaai.v35i8.16826
Quattrocchi CC, Errante Y, Gaudino C, Mallio CA, Giona A, Santini D, Tonini G, Zobel BB (2012) Spatial brain distribution of intra-axial metastatic lesions in breast and lung cancer patients. J Neuro-Oncol 110:79–87. https://doi.org/10.1007/s11060-012-0937-x
Funding
Partial funding through the USC Norris Comprehensive Cancer Center’s Multi-Level Cancer Risk Prediction Models pilot Project Award, ‘Molecular, Clinical and Neuro-imaging Determinants of Spatiotemporal Pathogenesis of Cancer-Specific Brain Metastases: Data Analysis and Longitudinal Modeling’ (12/01/2020-11/30/2021) is gratefully acknowledged.
Author information
Authors and Affiliations
Contributions
S.M., D.P., J.M., and P.N. wrote the main manuscript text. D.P., B.S., J.N., and G.Z. provided and interpreted the data. S.M. and P.N. prepared figures and plots. T.K.E, S.P., Y.S., A.H., D.M., M.T., J.S., S.P., and G.M. collected the data. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mahmoodifar, S., Pangal, D.J., Neman, J. et al. Comparative analysis of the spatial distribution of brain metastases across several primary cancers using machine learning and deep learning models. J Neurooncol (2024). https://doi.org/10.1007/s11060-024-04630-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11060-024-04630-5