Abstract
Background
Noug is an Ethiopian indigenous oilseed crop cultivated primarily for its oil and various economic importance. Evaluating the extent of genetic diversity within and among populations is one of the most important steps in breeding and conservation measures. Thus, this study aimed to uncover the extent of genetic diversity and population structure of noug accessions collected from different regions of Ethiopia using microsatellite markers.
Methods and results
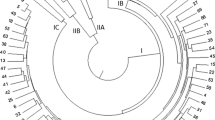
A total of 161 accessions from fourteen regions of Ethiopia, including some from Eritrea using 13 microsatellite markers were analyzed. All the 13 microsatellite markers were polymorphic and highly informative with a mean PIC value of 0.82. The analysis generated a total of 158 alleles with a mean of 12.15 per locus. The overall mean of Shannon information index and heterozygosity/gene diversity were 1.57 and 0.74, respectively suggesting the presence of higher genetic diversity across the collection regions. AMOVA revealed that 96.06% of the total genetic variation was attributed to within populations while only 3.94% was attributed to among populations. Likewise, the dendrogram clustering, PCoA, and the model-based population structure analysis didn’t exactly corresponded the grouping of the genotypes according to their regions of origin.
Conclusion
The microsatellites used in the present study are highly informative and could be targeted for developing markers for future marker-assisted breeding. Genotypes collected from Shewa, Wollo, Gojjam, Tigray, and B/G showed a higher genetic diversity and private alleles as compared to other populations. Hence, these areas can be considered as hotspots which could help for the identification of genotypes that can be used in breeding programs as well as for the implementation of further conservation programs.




Similar content being viewed by others
Data availability
This is part of M.Sc. thesis for the first author and can be found on the University’s repository available at “http://etd.aau.edu.et/handle/123456789/27424”.
References
Murthy HN, Hiremath SC, Salimath SS (1993) Origin, evolution and genome differentiation in Guizotia abyssinica and its wild species. Theoret Appl Genet 87:587–592. https://doi.org/10.1007/BF00221882
Dagne K (1994) Meiosis in interspecific hybrids and genomic interrelationships in Guizotia Cass. (Compositae). Hereditas 121:119–129. https://doi.org/10.1111/j.1601-5223.1994.00119.x
Hiremath SC, Murthy HN (1988) Domestication of niger (Guizotia abyssinica). Euphytica 37:225–228
Dempewolf H, Tesfaye M, Teshome A et al (2015) Patterns of domestication in the Ethiopian oil-seed crop noug (Guizotia abyssinica). Evol Appl 8:464–475. https://doi.org/10.1111/eva.12256
Getinet A, Sharma S (1996) Niger, Guizotia abyssinica (L. f.) Cass
CSA (Central Statistical Agency) (2019) Sample survey area and production of major crops (private peasant holdings, meher season). I:1–58
Alemayehu N, Alemaw G (1997) Highland Oilcrops: a two-decade research experience in Ethiopia
Geleta M, Asfaw Z, Bekele E, Teshome A (2002) Edible oil crops and their integration with the major cereals in North Shewa and South Welo, Central Highlands of Ethiopia: an ethnobotanical perspective. Hereditas 137:29–40. https://doi.org/10.1034/j.1601-5223.2002.1370105.x
Yadav S, Hussain Z, Suneja P et al (2012) Genetic divergence studies in niger (Guizotia abyssinica) germplasm. Biomass Bioenerg 44:64–69. https://doi.org/10.1016/j.biombioe.2012.04.011
MANR (2016) Plant Variety Release. Protection and Seed quality Control Directorate. Crop Variety Registered 330
Dagne K, Jonsson A (1997) Oil content and fatty acid composition of seeds of Guizotia cass (Compositae). J Sci Food Agric 73:274–278. https://doi.org/10.1002/(SICI)1097-0010(199703)73:3%3c274::AID-JSFA725%3e3.0.CO;2-F
Geleta M, Stymne S, Bryngelsson T (2011) Variation and inheritance of oil content and fatty acid composition in niger (Guizotia abyssinica). J Food Compos Anal 24:995–1003. https://doi.org/10.1016/j.jfca.2010.12.010
Kandel H, Porter P (2002) Niger (Guizotia abyssinica)(L. f.) Cass. Production in northwest Minnesota University of Minnesota extension service
Terefe M, Girma D (2022) Development of molecular resources for the genetic improvement of noug (Guizotia abyssinica (L. f) Cass): a mini review. CABI Agriculture and Bioscience https://doi.org/10.1186/s43170-022-00121-7
Mkwaila W, Terpstra KA, Ender M, Kelly JD (2011) Identification of QTL for agronomic traits and resistance to white mold in wild and landrace germplasm of common bean. Plant Breed 130:665–672. https://doi.org/10.1111/j.1439-0523.2011.01876.x
Glaszmann JC, Kilian B, Upadhyaya HD, Varshney RK (2010) Accessing genetic diversity for crop improvement. Curr Opin Plant Biol 13:167–173. https://doi.org/10.1016/j.pbi.2010.01.004
Singh Kesawat M, Das BK (2009) Molecular markers: it’s application in crop improvement types of molecular markers. J Crop Sci Biotech 12:169–181
Hasan N, Choudhary S, Naaz N et al (2021) Recent advancements in molecular marker-assisted selection and applications in plant breeding programmes. J Genet Eng Biotechnol 19:1–26. https://doi.org/10.1186/s43141-021-00231-1
Geleta M, Bryngelsson T, Bekele E, Dagne K (2007) Genetic diversity of Guizotia abyssinica (L. f.) Cass. (Asteraceae) from Ethiopia as revealed by random amplified polymorphic DNA (RAPD). Genet Resour Crop Evol 54:601–614. https://doi.org/10.1007/s10722-006-0018-0
Geleta M, Bryngelsson T, Bekele E, Dagne K (2008) Assessment of genetic diversity of Guizotia abyssinica (L.f.) Cass. (Asteraceae) from Ethiopia using amplified fragment length polymorphism. Plant Genet Resour 6:41–51. https://doi.org/10.1017/S1479262108913903
Petros Y, Merker A, Zeleke H (2007) Analysis of genetic diversity of Guizotia abyssinica from Ethiopia using inter simple sequence repeat markers. Hereditas 144:18–24. https://doi.org/10.1111/j.2007.0018-0661.01969.x
Hussain Z, Yadav S, Kumar S et al (2015) Molecular characterization of niger [Guizotia abyssinica (L.f.) Cass.] germplasms diverse for oil parameters. Indian J Biotechnol 14:344–350
Dempewolf H, Kane NC, Ostevik KL et al (2010) Establishing genomic tools and resources for Guizotia abyssinica (L.f.) Cass.-the development of a library of expressed sequence tags, microsatellite loci, and the sequencing of its chloroplast genome. Mol Ecol Resour 10:1048–1058. https://doi.org/10.1111/j.1755-0998.2010.02859.x
Abebaw M, Solomon A (2017) Genetic diversity assessment of Guzoita abyssinica using EST derived simple sequence repeats (SSRs) markers. Afr J Plant Sci 11:79–85. https://doi.org/10.5897/ajps2016.1512
Aboye B, Gebrselassie W, Disasa T (2020) Estimating-the-genetic-diversity-of-ethiopian-noug-guizotia-abyssinica-lf-cass-genotypes-using-ssr-markers
Tsehay S, Ortiz R, Johansson E et al (2020) New transcriptome-based SNP markers for Noug (Guizotia abyssinica) and their conversion to KASP markers for population genetics analyses. Genes 11:1373
Gebeyehu A, Hammenhag C, Tesfaye K, Vetukuri RR (2022) RNA-seq provides novel genomic resources for noug (Guizotia abyssinica ) and reveals microsatellite frequency and distribution in its transcriptome. Front Plant Sci 13:1–16. https://doi.org/10.3389/fpls.2022.882136
Koilkonda P, Sato S, Tabata S et al (2012) Large-scale development of expressed sequence tag-derived simple sequence repeat markers and diversity analysis in Arachis spp. Mol Breed 30:125–138. https://doi.org/10.1007/s11032-011-9604-8
DARTs (2000) Plant DNA Extraction Protocol for DArT. Doc 2–3
Pavel AB, Vasile CI (2012) PyElph—a software tool for gel images analysis and phylogenetics. BMC Bioinform. https://doi.org/10.1186/1471-2105-13-9
Liu K, Muse SV (2005) PowerMaker: an integrated analysis environment for genetic maker analysis. Bioinformatics 21:2128–2129. https://doi.org/10.1093/bioinformatics/bti282
Peakall R, Smouse PE (2012) GenALEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28:2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Kalinowski ST (2005) HP-RARE 1.0: a computer program for performing rarefaction on measures of allelic richness. Mol Ecol Notes 5:187–189. https://doi.org/10.1111/j.1471-8286.2004.00845.x
Excoffier L, Laval G, Schneider S (2005) Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evolut Bioinform 1:117693430500100. https://doi.org/10.1177/117693430500100003
Kumar S, Stecher G, Li M et al (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Takezaki N, Nei M, Tamura K (2010) POPTREE2: software for constructing population trees from allele frequency data and computing other population statistics with windows interface. Mol Biol Evol 27:747–752. https://doi.org/10.1093/molbev/msp312
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Kopelman NM, Mayzel J, Jakobsson M et al (2015) Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Resour 15:1179–1191
Li Y, Liu J (2018) StructureSelector: a web-based software to select and visualize the optimal number of clusters using multiple methods. Mol Ecol Resour 18:176–177
Botstein D, White RL, Skolnick M, Davis RW (1980) Botstein. Am J Hum Gen 32:314–331
Lacape J-M, Dessauw D, Rajab M et al (2007) Microsatellite diversity in tetraploid Gossypium germplasm: assembling a highly informative genotyping set of cotton SSRs. Mol Breed 19:45–58
Slatkin M (1987) Gene flow and the geographic structure of natural populations. Science 236:787–792
Iskakova AN, Romanova AA, Voronina EN et al (2014) Allele frequency and genotype distribution of 9 SNPs in the Kazakh population. J Pharmacogenomics Pharmacoproteomics 5:645–2153
Nei M, Roychoudhury AK (1974) Sampling variances of heterozygosity and genetic distance. Genetics 76:379–390. https://doi.org/10.1093/genetics/76.2.379
Filippi CV, Aguirre N, Rivas JG et al (2015) Population structure and genetic diversity characterization of a sunflower association mapping population using SSR and SNP markers. BMC Plant Biol 15:1–12. https://doi.org/10.1186/s12870-014-0360-x
Geleta M, Bekele E, Dagne K, Bryngelsson T (2010) Phylogenetics and taxonomic delimitation of the genus Guizotia (Asteraceae) based on sequences derived from various chloroplast DNA regions. Plant Syst Evol 289:77–89. https://doi.org/10.1007/s00606-010-0334-x
Zeinalzadeh-Tabrizi H, Haliloglu K, Ghaffari M, Hosseinpour A (2018) Assessment of genetic diversity among sunflower genotypes using microsatellite markers. Mol Biol Res Commun 7:143–152. https://doi.org/10.22099/mbrc.2018.30434.1340
Genet T, Belete K (2000) Phenotypic diversity in the Ethiopian noug germplasm. Afr Crop Sci J 8:137–143
Wright S (1949) The genetical structure of populations. Ann Eugen 15:323–354
Duminil J, Fineschi S, Hampe A et al (2015) From life-history traits ? 169:662–672
Hamrick JL, Godt MJW (1990) Allozyme diversity in plant species. Plant Popul Genet Breed Genet Resour 43–63
Ahmed W, Feyissa T, Tesfaye K, Farrakh S (2021) Genetic diversity and population structure of date palms (Phoenix dactylifera L.) in Ethiopia using microsatellite markers. J Genet Eng Biotechnol 19:1–14. https://doi.org/10.1186/s43141-021-00168-5
Ramachandran TK, Menon PM (1979) Pollination mechanism and inbreeding depression in niger (Guizotia abyssinica Cass.)[oil crops, India]. Madras Agric J
Bekele E, Geleta M, Dagne K et al (2007) Molecular phylogeny of genus Guizotia (Asteraceae) using DNA sequences derived from ITS. Genet Resour Crop Evol 54:1419–1427. https://doi.org/10.1007/s10722-006-9126-0
Acknowledgements
The authors are grateful to the Ethiopian Biodiversity Institute (EBI) for the provision of noug-seed samples. The authors are also thankful to the Ethiopian Institute of Agricultural Research (EIAR) and Addis Ababa University, Institute of Biotechnology, for the financial support.
Funding
NA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interest
The authors declare that they have no competing interest.
Ethical approval and consent to participate
NA.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
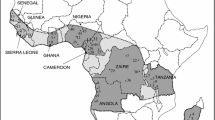
11033_2022_8005_MOESM2_ESM.jpg
Supplementary file2 Map showing noug sample collection regions (NB: all the sample areas, points, and boundaries are approximate and have nothing to do with political boundaries). The map was constructed using, ArcGIS_Desktop_1022_es_140418 (JPG 188 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Terefe, M., Birmeta, G., Girma, D. et al. Analysis of genetic diversity and population structure of oilseed crop noug (Guizotia abyssinica) accessions collected from Ethiopia. Mol Biol Rep 50, 43–55 (2023). https://doi.org/10.1007/s11033-022-08005-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-08005-9