Abstract
Background
Coronavirus-19 is still considered a pandemic that influences the world. Other molecular alterations should be clearer besides the increasing cytokine storm and pro-inflammatory molecules. Hypoxic conditions that induce HIF-1α lead to stimulate gene expression of STC-2 that targets PAPP-A expression. This study aimed to determine gene expression levels of PAPP-A, STC-2, and HIF-1α in COVID-19 infection. We also aimed to reveal the relationship of these genes with laboratory and clinical data of COVID-19 patients.
Materials and Results
We extracted RNA from peripheral blood samples of COVID-19(+) and COVID-19(−) individuals. The real-time PCR method was used to measure mRNA expression of PAPP-A, STC-2, and HIF-1α. Gene expression analysis was evaluated by the 2−ΔΔCt method. PAPP-A, STC-2, and HIF-1α mRNA expressions of severe patients were higher than healthy individuals (p = 0.0451, p = 0.4466, p < 0.0001, respectively). Correlation analysis of gene expression patterns of severe patients demonstrated a positive correlation between PAPP-A and STC-2 (p < 0.0001, r = 0.8638).
Conclusion
This is the first study that investigates the relation of PAPP-A, STC-2, and HIF-1α gene expression in patients with COVID-19 infection. Besides the routine laboratory findings, PAPP-A, STC-2, and HIF-1α mRNA expressions may be considered to patients’ prognosis as a sign of increased cytokines and pro-inflammatory molecules.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) that causes Coronavirus disease 2019 (COVID-19) infection still is a health problem in the world. Though there are several vaccines to diminish infection effects, all people can be infected with SARS-CoV-2, but the elderly are more vulnerable to severe forms of COVID-19. Chronic diseases make treatment more complicated. COVID-19 may be considered a systemic disease, as inflammatory cells destroy many organs at the same time [1].
Pregnancy‐associated plasma protein‐A (PAPP-A) is a metalloproteinase that is a member of the pappalysin family of metzincin metalloproteinase [2]. PAPP-A is expressed in many different cells including human osteoblasts [3], vascular smooth muscle cells [4], and ovarian granulosa cells [5]. Proinflammatory cytokine expressions regulate PAPP-A mRNA expression in human fibroblasts, and osteoblasts [4, 6]. Macrophage‐derived pro‐inflammatory cytokines lead to PAPP-A secretion from vascular smooth muscle and endothelial cells [7].
Stanniocalcin-2 (STC-2) is a secreted glycoprotein expressed in many types of tissues [8] and is a member of the stanniocalcin family [9]. STC-2 has a potential role in the inhibition of PAPP-A [10]. STC-2 promoter has hypoxia response elements (HRE) so it is a target of hypoxia-inducible factor 1α (HIF-1α) [11]. Oxidative stress and hypoxia increase the STC-2 level [12].
Inflammation related to hypoxia is so important clinically [13]. In the acute phase of sepsis strong pro-inflammatory response is seen, however in the late phase of the sepsis immune-suppressed state allows for tissue repair [14]. HIFs are heterodimeric proteins that have α and β subunits. HIF-1α has an active role in acute hypoxia condition [15]. HIF-1α is a transcription factor that regulates cell adaptive response to hypoxia [15]. HIF-1α binds to target gene promoters including HRE and regulates their expression [16]. It regulates many genes which have roles in survival, immune reaction, cytokine production, vascularization, and cellular homeostasis [15, 17, 18]. HIF-1β localized in the nucleus but HIF-1α located in the cytoplasm and translocate to the nucleus under hypoxic conditions [19]. Cytokine production and metabolism clinical symptomology influence HIF-1α [15]. Hypoxic conditions stimulate HIF-1α expression. However, HIF-1α gene expression attenuates in process of time [20]. HIF-1α was found to be decreased in septic patients and this decline was correlated with disease severity [21].
Although the absence of several vaccines that do not prevent spreading the disease, there is no specific medication for the treatment of the infection. The molecular changes of the disease need to be clearer. HIF-1α is a transcription factor that is upregulated by hypoxic conditions [19]. Hypoxic conditions stimulate gene expression of STC-2 [12] and STC-2 target gene PAPP-A. So investigating the relation of these three genes in COVID-19 infection may contribute to a better understanding of molecular changes in the disease. In line with this information, we investigated the gene expression level of PAPP-A, STC-2 in COVID-19(+) patients.
Materials and methods
Ethical consideration
This study was realized in compliance with the Declaration of Helsinki principles and required permission was obtained from the Ministry of Health, Turkey. Ethical approval was gained Ethics Committee of the Ataturk University Clinical Researches of Ethical Committee (Decision no;68). The participants were older than eighteen and informed consent was obtained from all individual participants or their legal representatives included in the study.
Patients and study design
To this study, 40 healthy volunteers and 70 COVID-19(+) patients (non-severe and sever patients) were enrolled. All participants’ situations were confirmed via COVID-19 polymerase chain reaction (PCR) test. EDTA whole blood from participants were picked up at Erzurum Regional Education and Research Hospital and storage at – 20 °C for a few days. 40 COVID-19 (+) patients had severe symptoms including decreased oxygen saturation, suspected respiratory infection symptom, shortness of breath, however 30 COVID-19(+) patients had non-severe symptoms including fever, fatigue, cough, muscle pain, sore throat, headache. Pregnant women were excluded from the study.
RNA extraction, cDNA synthesis, real-time PCR
Blood from volunteers (100 µL) was used for extraction of total RNA by EcoPURE Total RNA Kit (EcoTech Biotechnology, Erzurum, Turkey) according to the manufacturer’s instructions. Total RNA was quantified via Epoch Spectrophotometer System and Take3 Plate (BioTek, USA) and stored at – 20 °C until cDNA synthesis. iScript cDNA Synthesis Kit was used to convert RNA to cDNA. Gene expressions of PAPP-A, STC-2, and HIF-1α were realized with ABI StepOne Plus Real-Time PCR System (ABI, USA). Β-actin was an internal control gene. 20 μL PCR reaction included 10 μL SsoAdvanced Universal SYBR Green Supermix (BioRad, Richmond, CA), 1 μL of Forward primer and 1 μL of Reverse primer, 4 μL of cDNA, and 4 μL PCR-grade water. RT-PCR conditions were 30 s at 95 °C, 10 s at 95 °C, and 1 min at 60 °C. 2−ΔΔCt method was used to analyze gene expression of target genes [22].
Primer sequences used in this study are as follows;
PAPP-A
Forward Primer:5′-ACAAAGACCCACGCTACTTTTT-3′
Reverse Primer:5′-CATGAACTGCCCATCATAGGTG-3′
STC-2
Forward Primer:5′-GGGTGTGGCGTGTTTGAATG-3′
Reverse Primer:5′-TTTCCAGCGTTGTGCAGAAAA-3′
HIF-1α
Forward Primer:5′-CATAAAGTCTGCAACATGGAAGGT-3′
Reverse Primer:5′-ATTTGATGGGTGAGGAATGGGTT-3′
β-actin
Forward Primer:5′-CATGTACGTTGCTATCCAGGC-3′
Reverse Primer:5′-CTCCTTAATGTCACGCACGAT-3′
Data collection of patients
Demographic, clinical information and laboratory findings of patients was gained from hospital database. To evaluate laboratory findings, white blood cell (WBC) count, Neutrophil (Neu), Lymphocyte (LYM), platelet, C-reactive protein (CRP), procalcitonin, fibrinogen, D-dimer, alanine aminotransferase (ALT), aspartate aminotransferase (AST), albumin, creatinine, hemoglobin, hematocrit (HCT), lactate dehydrogenase (LDH), bilirubin (total), blood urea nitrogen (BUN) were taken into account.
Statistically analysis
All experimental results were assessed with GraphPad Prism (version 5; San Diego, CA). To evaluate data distribution Kolmogorov–Smirnov test was used. Statistical differences between two groups were realized by two-tailed Student's t test, for normally distributed data, or the Mann–Whitney U as a nonparametric equivalent of the Student's t test. To comparison, multiple subgroup analysis Kruskal–Wallis test was performed. Correlation of gene expression levels was carried out with Spearman correlation analysis. p values < 0.05 were considered statistically significant.
Results and discussion
Results
Characteristics and laboratory findings of the COVID-19 cohorts
The mean ± SD age of the groups including healthy individuals, non-severe patients, severe patients were 48 ± 15, 46 ± 16, and 68 ± 10, respectively. The age range of patients (non-severe and severe patients) was 20–90 years. The percentage of male patients was 53 and of female patients was 47. The lungs of 77% of the patients were affected by the COVID-19 infection. Some patients had more than one comorbidity. Severe patients mostly consisted of elderly individuals so they had more comorbidities including 60% hypertension, 25% heart diseases, 33% diabetes mellitus, 23% chronic lung diseases. The healthy individuals and non-severe patients had fewer comorbidities than the severe patients. In the healthy individuals, just 4 individuals were with comorbidities that are 5% hypertension and 5% chronic lung diseases. The non-severe patients involve 9 patients with comorbidities including 13% to hypertension, 10% to diabetes mellitus, and 3% to chronic lung diseases. Comorbidities may have contributed to the severity of infection for severe patients. The percentage of patients who died in the hospital is 11.4%. Most of them were intensive care patients.
Laboratory findings of non-severe and severe patients are demonstrated in Table 1. Severe patients had elevated WBC counts, Neu, ALT, AST, bilirubin(total), LDH, procalcitonin, CRP, D-dimer, creatinine, fibrinogen, BUN compared with non-severe patients (p = 0.0304; p = 0.0461; p = 0.4149; p = 0.0066; p = 0.2174; p < 0.0001; p = 0.0039; p < 0.0001; p < 0.0001; p < 0.0001; p = 0.0073; p < 0.0001, respectively).
Decreased LYM, PLT, hemoglobin, HCT, and albumin values were observed in the severe patients when compared with the non-severe patients (p < 0.0001; p = 0.0272; p = 0.0043; p = 0.0458; p < 0.0001, respectively).
Altered gene expression levels of PAPP-A, STC-2, and HIF-1α
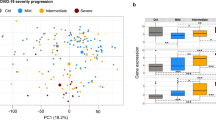
We compared PAPP-A, STC-2, and HIF-1α gene expression levels in healthy individuals, non-severe and severe patients. We observed that PAPP-A expressions among cohorts showed significant differences (p = 0.0451). There was an increased gene expression in severe patients but decreased in non-severe patients (Fig. 1A). STC-2 had the same pattern but it is not significant statically (p = 0.3485) (Fig. 1B). STC-2 expression was higher in severe patients however lower in non-severe patients than healthy individuals. Furthermore, HIF-1α showed different expression pattern among cohorts (Fig. 1C) (p < 0.0001). Expression of HIF-1α was higher in severe patients than healthy individuals but lower than non-severe patients. This data demonstrates that gene expression of PAPP-A, STC-2 HIF-1α showed different patterns among cohorts however all genes showed increased gene expression levels in severe patients than controls.
Correlation analysis of gene expression
We used the gene expression levels of severe patients to reveal the correlation of the genes. When we examined PAPP-A and STC-2 we observed a strong positive correlation (r = 0.8638, p < 0.0001) (Fig. 2). PAPP-A and HIF-1α gene expressions have negative correlation, not significantly (r = − 0.07298 p = 0.6964). Correlation analysis also revealed a negative but not significant correlation for STC-2 and HIF-1α (r = − 0.2573 p = 0.1863) in severe patients.
Variable gene expression levels according to laboratory findings of COVID-19 patients
We investigated whether there is any relationship between patients' demographic information including age and gender and gene expressions. When we evaluated according to demographic information, we did not find any significant difference in gene expressions between patients (p > 0.05). Besides that, the results showed that patients whose lungs were affected by COVID-19 infection had lower levels of HIF-1α gene expression than patients whose lungs were not affected by COVID-19 infection (p = 0.0019).
When the PAPP-A, STC-2 and HIF-1α gene expressions of patients with low, high and normal laboratory findings were compared, no significant difference was found between the patients (p > 0.05). However, patients with higher creatinine levels showed decreased HIF-1α expression compared with patients with normal creatinine levels (p = 0.0437). Patients with low hemoglobin levels showed decreased PAPP-A and STC-2 gene expressions (p = 0.0433 p = 0.0224, respectively).
Discussion
SARS-CoV-2 infection still keeps the mystery since unexplained underlying molecular mechanisms and reasons causing cytokine storms. Routine laboratory findings have been used to estimate the prognosis of patients with COVID-19. Therefore, it is imperative to reveal the new biomarkers that underlie COVID-19 infection that affect treatment. So, we investigated PAPP-A, STC-2, and HIF-1α gene expression levels in COVID-19(+) patients.
PAPP-A is a protein that is under the regulation of pro-inflammatory cytokines including IL-1β and TNF-α [23]. A recent retrospective study of COVID-19 demonstrated elevated serum levels of PAPP-A protein in early infected patients and suggested PAPP-A as a biomarker to the early stage of COVID-19 [24]. SARS-CoV-2 study in pregnant patients and control group demonstrated no differences of PAPP-A gene expression between cohorts [25]. Trilla et al. elevated PAPP-A levels of pregnant patients who is infected with SARS-CoV-2 and showed higher PAPP-A level in symptomatic women than asymptomatic and negative women [26]. Serrano et al. was found no differences for PAPP-A level between pregnant women and general population [27]. In our study PAPP-A mRNA levels were increased in severe patients but decreased in non-severe patients. The discrepancy of the studies is based on differences of cohorts and/or different stages of the disease. We suggest that PAPP-A may be used as a sign for the advanced disease stage.
At the search of the literature, there was no study about STC-2 in COVID-19 patients. Haijin Lv et al. demonstrated anti-oxidant and anti-inflammatory properties of STC-2 in mesenchymal stromal cells [28]. In vivo and in vitro suppression effect of STC-2 on ROS was shown [28]. We demonstrated increased mRNA level of STC-2 in severe COVID-19 patients however decreased mRNA level in non-severe patients but statically not significant. Increased oxidative stress and hypoxia at the different stages of COVID-19 infection may regulate STC-2 expression, disparately. So, we propose that STC-2 may represent different levels of hypoxia and oxidative stress in severe COVID-19 infection.
The role of hypoxia in COVID-19 infection has been shown by many studies. HIF-1α is a hypoxia-induced molecule. Zhang et al. suggested HIF-1α signaling pathway as common key pathway for SARS-CoV-2 infection [29]. Critically ill patients with COVID-19 infection were showed increased HIF-1α expression [30]. Codo et al. showed higher HIF-1α expression in blood monocytes from severe COVID-19 patients than healthy donors. When they treated blood monocyte with HIF-1α inhibitor they saw decreased HIF-1α expression and abated SARS-CoV-2 replication, besides that treated with HIF-1α stabilizator aggravated SARS-CoV-2 infection and HIF-1α expression [31]. Zhu et al. claimed that HIF-1α expression is associated with macrophage inflammation in COVID-19 patients [32]. Tian et al. suggested SARS-CoV-2 ORF-3A induce mitochondrial damage that cause to promotion of HIF-1α expression [33]. Rolfo et al. investigated placental HIF-1α level in third-trimester pregnancies with asymptomatic and symptomatic COVID-19 patients. They found that overexpression of HIF-1α in the placenta of pregnant patients with asymptomatic and symptomatic COVID-19 [34]. In our study, HIF-1α showed an increased level of mRNA expression in all patients than healthy individuals. When we compared cohorts, non-severe patients’ HIF-1α expression was higher than severe patients. This may be explained by the advanced stage of disease and attenuating HIF-1α expression by the time of progress. Conflicting diseases may be the results of various patients and/or types of different infectious agents caused to diseases.
PAPP-A and STC-2 relation was studied before at several diseases but not to COVID-19 infection. STC-2 is a glycoprotein that binds PAPP-A and limited its proteolytic activity [10]. According to the previous studies in the literature the proteolytic activity of STC-2 on PAPP-A may be changed by some factors including disease type, tissues type, patients. Panagiotou et al. showed increased PAPP-A and STC-2 by exercise in healthy individuals [35]. Ortega et al. demonstrated increased PAPP-A and decreased STC-2 gene expression in patients with chronic venous disease [36]. Hjortebjerg et al. revealed unchanged PAPP-A and but decreased STC-2 expression after Roux-en-Y gastric bypass operation in obese subjects [37]. In pulmonary disease patients, plasma and tissue fluid were compared and results showed a negative correlation between PAPP-A and STC-2 in tissue fluid but did not show in serum [38]. In our study, we found a strong positive correlation between STC-2 and PAPP-A mRNA expression. The effect of increased oxidative stress and pro-inflammatory molecules that influence PAPP-A [23] and STC-2 [28] may increase the level of both gene expressions. Relation of HIF-1α and PAPP-A was studied in the endometrial tumor before and results demonstrated positive associations [39]. In our study, we did not find a significant correlation between HIF-1α and PAPP-A gene expressions. Hypoxia induces STC-2 expression and STC-2 is also a target gene of HIF-1α [16]. In our study, we found a negative but not significant correlation between HIF-1α and STC-2 gene expression levels in severe COVD-19 patients. The extended patient population may show a statically significant correlation.
This study has some limitations. First, we enrolled less patient population than we target. Second, we could not access all patients’ information. Gene expressions can vary depending on many factors such as genetic and epigenetic changes, intercellular and intracellular signaling. It is often very difficult to attribute changes in gene expressions to a single cause. However, it is necessary to consider some parameters that may contribute to alteration in HIF-1α, PAPP-A and STC-2 gene expression. Third, another limitation of this study is the inability to analyze the presence of specific immune cell markers [40] that may cause differences in gene expression in this study. Oxidative stress and/or hypoxia are the major part of the disease. In vitro analysis that could demonstrate whether hypoxia, oxidative stress, or both contribute to changes in gene expression were not performed in this study.
Conclusion
In conclusion, this is the first study to investigate together the expression of PAPP-A, STC-2, and HIF-1α mRNA in COVID-19 infection. This study firstly showed a positive correlation between PAPP-A and STC-2 in COVID-19(+) patients. Increasing oxidative stress and hypoxia and elevated levels of pro-inflammatory molecules may lead to the alteration of gene expression of PAPP-A, STC-2, and HIF-1α mRNA. PAPP-A, STC-2, and HIF-1α expressions may use a sign of disease severity. Correlation of PAPP-A and STC-2 may be considered in addition to routine laboratory tests that show disease prognosis since their regulation effected by hypoxia and pro-inflammatory molecules. More in vivo and in vitro studies are needed to more clearly reveal the relationship between PAPP-A, STC-2 and HIF-1α in COVID-19 infection.
References
Huang C, Wang Y, Li X et al (2020) Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. The Lancet 395:497–506. https://doi.org/10.1016/S0140-6736(20)30183-5
Boldt HB, Overgaard MT, Laursen LS et al (2001) Mutational analysis of the proteolytic domain of pregnancy-associated plasma protein-A (PAPP-A): classification as a metzincin. Biochem J 358:359. https://doi.org/10.1042/0264-6021:3580359
Conover CA, Chen B-K, Resch ZT (2004) Regulation of pregnancy-associated plasma protein-A expression in cultured human osteoblasts. Bone 34:297–302. https://doi.org/10.1016/j.bone.2003.10.011
Conover CA, Bale LK, Harrington SC et al (2006) Cytokine stimulation of pregnancy-associated plasma protein A expression in human coronary artery smooth muscle cells: inhibition by resveratrol. Am J Physiol Physiol 290:C183–C188. https://doi.org/10.1152/ajpcell.00199.2005
Conover CA, Faessen GF, Ilg KE et al (2001) Pregnancy-associated plasma protein-A is the insulin-like growth factor binding protein-4 protease secreted by human ovarian granulosa cells and is a marker of Dominant follicle selection and the corpus luteum. Endocrinology 142:2155–2155. https://doi.org/10.1210/endo.142.5.8286
Resch ZT, Chen B-K, Bale LK et al (2004) Pregnancy-associated plasma protein A gene expression as a target of inflammatory cytokines. Endocrinology 145:1124–1129. https://doi.org/10.1210/en.2003-1313
Conover CA, Harrington SC, Bale LK, Oxvig C (2007) Surface association of pregnancy-associated plasma protein-A accounts for its colocalization with activated macrophages. Am J Physiol Circ Physiol 292:H994–H1000. https://doi.org/10.1152/ajpheart.00798.2006
Chang AC-M, Reddel RR (1998) Identification of a second stanniocalcin cDNA in mouse and human: stanniocalcin 2. Mol Cell Endocrinol 141:95–99. https://doi.org/10.1016/S0303-7207(98)00097-5
Ishibashi K, Miyamoto K, Taketani Y et al (1998) Molecular cloning of a second human stanniocalcin homologue (STC2). Biochem Biophys Res Commun 250:252–258. https://doi.org/10.1006/bbrc.1998.9300
Jepsen MR, Kløverpris S, Mikkelsen JH et al (2015) Stanniocalcin-2 inhibits mammalian growth by proteolytic inhibition of the insulin-like growth factor axis. J Biol Chem 290:3430–3439. https://doi.org/10.1074/jbc.M114.611665
Law AYS, Lai KP, Ip CKM et al (2008) Epigenetic and HIF-1 regulation of stanniocalcin-2 expression in human cancer cells. Exp Cell Res 314:1823–1830. https://doi.org/10.1016/j.yexcr.2008.03.001
Ito D, Walker JR, Thompson CS et al (2004) Characterization of stanniocalcin 2, a novel target of the mammalian unfolded protein response with cytoprotective properties. Mol Cell Biol 24:9456–9469. https://doi.org/10.1128/MCB.24.21.9456-9469.2004
Eltzschig HK, Carmeliet P (2011) Hypoxia and inflammation. N Engl J Med 364:656–665. https://doi.org/10.1056/NEJMra0910283
Van Wyngene L, Vandewalle J, Libert C (2018) Reprogramming of basic metabolic pathways in microbial sepsis: therapeutic targets at last? EMBO Mol Med 10:e8712. https://doi.org/10.15252/emmm.201708712
Fitzpatrick SF (2019) Immunometabolism and sepsis: a role for HIF? Front Mol Biosci 6:e00085. https://doi.org/10.3389/fmolb.2019.00085
Semenza GL, Jiang B-H, Leung SW et al (1996) Hypoxia response elements in the aldolase A, enolase 1, and lactate dehydrogenase A gene promoters contain essential binding sites for hypoxia-inducible factor 1. J Biol Chem 271:32529–32537. https://doi.org/10.1074/jbc.271.51.32529
Adams J, Difazio L, Rolandelli R et al (2009) HIF-1: a key mediator in hypoxia (Review). Acta Physiol Hung 96:19–28. https://doi.org/10.1556/APhysiol.96.2009.1.2
Yoon D, Pastore YD, Divoky V et al (2006) Hypoxia-inducible Factor-1 deficiency results in dysregulated erythropoiesis signaling and iron homeostasis in mouse development. J Biol Chem 281:25703–25711. https://doi.org/10.1074/jbc.M602329200
Kallio PJ (1998) Signal transduction in hypoxic cells: inducible nuclear translocation and recruitment of the CBP/p300 coactivator by the hypoxia-inducible factor-1alpha. EMBO J 17:6573–6586. https://doi.org/10.1093/emboj/17.22.6573
Prabhakar NR (2001) Invited review: oxygen sensing during intermittent hypoxia: cellular and molecular mechanisms. J Appl Physiol 90:1986–1994. https://doi.org/10.1152/jappl.2001.90.5.1986
Schäfer ST, Frede S, Winning S et al (2013) Hypoxia-inducible factor and target gene expression are decreased in patients with sepsis. Anesthesiology 118:1426–1436. https://doi.org/10.1097/ALN.0b013e31828baa67
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Conover CA, Oxvig C (2017) PAPP-A: a promising therapeutic target for healthy longevity. Aging Cell 16:205–209. https://doi.org/10.1111/acel.12564
Sanchez BG, Gasalla JM, Sánchez-Chapado M et al (2021) Increase in ischemia-modified albumin and pregnancy-associated plasma protein-A in COVID-19 patients. J Clin Med 10:5474. https://doi.org/10.3390/jcm10235474
Cosma S, Carosso AR, Borella F et al (2021) Prenatal biochemical and ultrasound markers in COVID-19 pregnant patients: a prospective case-control study. Diagnostics 11:398. https://doi.org/10.3390/diagnostics11030398
Trilla C, Mora J, Crovetto F et al (2022) First-trimester SARS-CoV-2 infection: clinical presentation, inflammatory markers and obstetric outcomes. Fetal Diagn Ther. https://doi.org/10.1159/000523974
Serrano B, Mendoza M, Garcia-Aguilar P et al (2022) Shared risk factors for COVID-19 and preeclampsia in the first trimester: an observational study. Acta Obstet Gynecol Scand. https://doi.org/10.1111/aogs.14371
Lv H, Liu Q, Sun Y et al (2020) Mesenchymal stromal cells ameliorate acute lung injury induced by LPS mainly through stanniocalcin-2 mediating macrophage polarization. Ann Transl Med 8:334–334. https://doi.org/10.21037/atm.2020.02.105
Zhang L, Li M, Wang Z et al (2022) Cardiovascular risk after SARS-CoV-2 infection is mediated by IL18/IL18R1/HIF-1 signaling pathway axis. Front Immunol 12:e780804. https://doi.org/10.3389/fimmu.2021.780804
Taniguchi-Ponciano K, Vadillo E, Mayani H et al (2021) Increased expression of hypoxia-induced factor 1α mRNA and its related genes in myeloid blood cells from critically ill COVID-19 patients. Ann Med 53:197–207. https://doi.org/10.1080/07853890.2020.1858234
Codo AC, Davanzo GG, de Monteiro LB et al (2020) Elevated glucose levels favor SARS-CoV-2 infection and monocyte response through a HIF-1α/glycolysis-dependent axis. Cell Metab 32:437-446.e5. https://doi.org/10.1016/j.cmet.2020.07.007
Zhu B, Wu Y, Huang S et al (2021) Uncoupling of macrophage inflammation from self-renewal modulates host recovery from respiratory viral infection. Immunity 54:1200-1218.e9. https://doi.org/10.1016/j.immuni.2021.04.001
Tian M, Liu W, Li X et al (2021) HIF-1α promotes SARS-CoV-2 infection and aggravates inflammatory responses to COVID-19. Signal Transduct Target Ther 6:308. https://doi.org/10.1038/s41392-021-00726-w
Rolfo A, Cosma S, Nuzzo AM et al (2022) Increased placental anti-oxidant response in asymptomatic and symptomatic COVID-19 third-trimester pregnancies. Biomedicines 10:634. https://doi.org/10.3390/biomedicines10030634
Panagiotou G, Anastasilakis AD, Kynigopoulos G et al (2017) Physiological parameters regulating circulating levels of the IGFBP-4/Stanniocalcin-2/PAPP-A axis. Metabolism 75:16–24. https://doi.org/10.1016/j.metabol.2017.07.003
Ortega MA, Fraile-Martínez O, Asúnsolo Á et al (2020) Chronic venous disease patients showed altered expression of IGF-1/PAPP-A/STC-2 axis in the vein wall. Biomed Res Int 2020:1–8. https://doi.org/10.1155/2020/6782659
Hjortebjerg R, Bojsen-Møller KN, Søeby M et al (2021) Metabolic improvement after gastric bypass correlates with changes in IGF-regulatory proteins stanniocalcin-2 and IGFBP-4. Metabolism 124:154886. https://doi.org/10.1016/j.metabol.2021.154886
Espelund US, Bjerre M, Hjortebjerg R et al (2017) Insulin-like growth factor bioactivity, stanniocalcin-2, pregnancy-associated plasma protein-A, and IGF-binding protein-4 in pleural fluid and serum from patients with pulmonary disease. J Clin Endocrinol Metab 102:3526–3534. https://doi.org/10.1210/jc.2017-00033
Iunusova N V, Spirina L V, Kondakova LA, et al [Relationship between the expression levels of PAPP-A metalloproteinase and growth and transcriptional factors in endometrial cancer]. Izv Akad Nauk Seriia Biol 284–91
Jin M, Shi N, Wang M et al (2020) CD45: a critical regulator in immune cells to predict severe and non-severe COVID-19 patients. Aging 12:19867–19879. https://doi.org/10.18632/aging.103941
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Karabulut Uzunçakmak, S., Naldan, M.E., Dirican, E. et al. Preliminary investigation of gene expression levels of PAPP-A, STC-2, and HIF-1α in SARS-Cov-2 infected patients. Mol Biol Rep 49, 8693–8699 (2022). https://doi.org/10.1007/s11033-022-07710-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07710-9