Abstract
Plants are subjected to biotic and abiotic stresses regularly, which irreparably harm agricultural production. Eco-friendly and sustainable technology to deal with this challenge is to breed abiotic stress tolerant cultivars. To generate crop plants conferring resistance against stresses, conventional breeding was used in the past, but because of the complex heredity of abiotic stress tolerance traits, such techniques remain insufficient in making greater enhancement. Genome-engineering based on CRISPR-Cas9 (clustered regularly interspaced short palindromic repeats-CRISPR associated protein9) has shown enormous potential in developing climate-resilient cultivars. Likewise, the development of chickpea transgenic lines by knockout of 4CL and REV7 genes exhibits drought tolerance which establishes a foundation for future studies in chickpea. In addition, the CRISPR-Cas9 system can boost yield potential under abiotic stress situations by producing non-transgenic plants having the required characteristics. This review article discusses the validation of gene function based on the CRISPR-Cas9 for the development of abiotic stress-tolerant crop plants, emphasizing the chickpea to open the new ventures of generating abiotic stress-tolerant chickpea varieties.

Similar content being viewed by others
References
Akbar H, Gheewala SH (2020) Effect of climate change on cash crops yield in Pakistan. Arab J Geosci 13:1–15
Calleja-Cabrera J, Boter M, Oñate-Sánchez L, Pernas M (2020) Root growth adaptation to climate change in crops. Front Plant Sci 11:544
Razzaq MK, Rauf S, Khurshid M et al (2019) Pollen Viability an Index of Abiotic Stresses Tolerance and Methods for the Improved Pollen Viability. Pakistan Journal of Agricultural Research32
Boscaiu M, Fita A (2020) Physiological and molecular characterization of crop resistance to abiotic stresses
Fita A, Rodríguez-Burruezo A, Boscaiu M, Prohens J, Vicente O (2015) Breeding and domesticating crops adapted to drought and salinity: a new paradigm for increasing food production. Front Plant Sci 6:978
Driedonks N, Rieu I, Vriezen WH (2016) Breeding for plant heat tolerance at vegetative and reproductive stages. Plant Reprod 29:67–79
Hussain F, Rafiq M, Ghias M et al (2017) Genetic diversity for seed yield and its components using principal component and cluster analysis in. Life Sci. J 14
Li M, Chen L, Zeng J et al (2020) Identification of additive–epistatic QTLs conferring seed traits in soybean using recombinant inbred lines. Frontiers in plant science 11
Ren J, Lee J, Na D (2020) Recent advances in genetic engineering tools based on synthetic biology. J Microbiol 58:1–10
Zhang Y, Massel K, Godwin ID, Gao C (2018) Applications and potential of genome editing in crop improvement. Genome Biol 19:1–11
Ashkani S, Rafii MY, Shabanimofrad M et al (2015) Molecular breeding strategy and challenges towards improvement of blast disease resistance in rice crop. Front Plant Sci 6:886
Fuchs M (2017) Pyramiding resistance-conferring gene sequences in crops. Curr Opin Virol 26:36–42
Liu H-J, Jian L, Xu J et al (2020) High-throughput CRISPR/Cas9 mutagenesis streamlines trait gene identification in maize. Plant Cell 32:1397–1413
Klap C, Yeshayahou E, Bolger AM et al (2017) Tomato facultative parthenocarpy results from Sl AGAMOUS-LIKE 6 loss of function. Plant Biotechnol J 15:634–647
Zhang K, Raboanatahiry N, Zhu B, Li M (2017) Progress in genome editing technology and its application in plants. Front Plant Sci 8:177
Razzaq MK, Aleem M, Mansoor S et al (2021) Omics and CRISPR-Cas9 Approaches for Molecular Insight, Functional Gene Analysis, and Stress Tolerance Development in Crops. Int J Mol Sci 22:1292
Das A, Datta S, Thakur S et al (2017) Expression of a chimeric gene encoding insecticidal crystal protein Cry1Aabc of Bacillus thuringiensis in chickpea (Cicer arietinum L.) confers resistance to gram pod borer (Helicoverpa armigera Hubner.). Front Plant Sci 8:1423
Rezaei MK, Deokar A, Tar’an B (2016) Identification and expression analysis of candidate genes involved in carotenoid biosynthesis in chickpea seeds. Front Plant Sci 7:1867
Lohani N, Jain D, Singh MB, Bhalla PL (2020) Engineering multiple abiotic stress tolerance in canola, Brassica napus. Front Plant Sci 11:3
Drinkwater KF, Beaugrand G, Kaeriyama M et al (2010) On the processes linking climate to ecosystem changes. J Mar Syst 79:374–388
Dai A (2011) Characteristics and trends in various forms of the Palmer Drought Severity Index during 1900–2008. Journal of Geophysical Research: Atmospheres116
He M, He C-Q, Ding N-Z (2018) Abiotic stresses: general defenses of land plants and chances for engineering multistress tolerance. Front Plant Sci 9:1771
Waqas MA, Kaya C, Riaz A et al (2019) Potential mechanisms of abiotic stress tolerance in crop plants induced by thiourea. Frontiers in plant science,1336
Zhang H, Zhu J, Gong Z, Zhu J-K (2021) Abiotic stress responses in plants. Nature Reviews Genetics,1–16
Yuan F, Yang H, Xue Y et al (2014) OSCA1 mediates osmotic-stress-evoked Ca2 + increases vital for osmosensing in Arabidopsis. Nature 514:367–371
Sahoo MR, Devi TR, Dasgupta M, Nongdam P, Prakash N (2020) Reactive oxygen species scavenging mechanisms associated with polyethylene glycol mediated osmotic stress tolerance in Chinese potato. Sci Rep 10:1–9
Bouzroud S, Gasparini K, Hu G et al (2020) Down regulation and loss of auxin response factor 4 function using CRISPR/Cas9 alters plant growth, stomatal function and improves tomato tolerance to salinity and osmotic stress. Genes 11:272
Zargar SM, Mahajan R, Bhat JA, Nazir M, Deshmukh R (2019) Role of silicon in plant stress tolerance: opportunities to achieve a sustainable cropping system. 3 Biotech 9:73
Sharma P, Jha AB, Dubey RS, Pessarakli M (2012) Reactive oxygen species, oxidative damage, and antioxidative defense mechanism in plants under stressful conditions. Journal of botany 2012
Fahad S, Bajwa AA, Nazir U et al (2017) Crop production under drought and heat stress: plant responses and management options. Front Plant Sci 8:1147
Qamar R, Ghias M, Hussain F et al (2018) Effect of drought on morpho-physiological traits of sunflower (Helianthus annuus L) hybrids and their parental inbred lines. Pakistan Journal of Agricultural Research31
Kulik A, Wawer I, Krzywińska E, Bucholc M, Dobrowolska G (2011) SnRK2 protein kinases—key regulators of plant response to abiotic stresses. OMICS 15:859–872
Li P, Li YJ, Zhang FJ et al (2017) The Arabidopsis UDP-glycosyltransferases UGT79B2 and UGT79B3, contribute to cold, salt and drought stress tolerance via modulating anthocyanin accumulation. Plant J 89:85–103
Li R, Liu C, Zhao R et al (2019) CRISPR/Cas9-Mediated SlNPR1 mutagenesis reduces tomato plant drought tolerance. BMC Plant Biol 19:1–13
Ke Q, Kim HS, Wang Z et al (2017) Down-regulation of GIGANTEA‐like genes increases plant growth and salt stress tolerance in poplar. Plant Biotechnol J 15:331–343
Cai R, Dai W, Zhang C et al (2017) The maize WRKY transcription factor ZmWRKY17 negatively regulates salt stress tolerance in transgenic Arabidopsis plants. Planta 246:1215–1231
Jiang B, Shi Y, Zhang X et al (2017) PIF3 is a negative regulator of the CBF pathway and freezing tolerance in Arabidopsis. Proceedings of the National Academy of Sciences 114, E6695-E6702
Tiwari M, Sharma D, Singh M, Tripathi RD, Trivedi PK (2014) Expression of OsMATE1 and OsMATE2 alters development, stress responses and pathogen susceptibility in Arabidopsis. Sci Rep 4:1–12
Tang N, Ma S, Zong W et al (2016) MODD mediates deactivation and degradation of OsbZIP46 to negatively regulate ABA signaling and drought resistance in rice. Plant Cell 28:2161–2177
Gupta D, Bhattacharjee O, Mandal D et al (2019) CRISPR-Cas9 system: A new-fangled dawn in gene editing. Life Sci 232:116636
Urnov FD, Rebar EJ, Holmes MC, Zhang HS, Gregory PD (2010) Genome editing with engineered zinc finger nucleases. Nat Rev Genet 11:636–646
Kim Y-G, Shi Y, Berg JM, Chandrasegaran S (1997) Site-specific cleavage of DNA–RNA hybrids by zinc finger/FokI cleavage domain fusions. Gene 203:43–49
Joung JK, Sander JD (2013) TALENs: a widely applicable technology for targeted genome editing. Nat Rev Mol Cell Biol 14:49–55
Mali P, Aach J, Stranges PB et al (2013) CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat Biotechnol 31:833–838
Ma X, Zhang Q, Zhu Q et al (2015) A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8:1274–1284
Doudna JA, Charpentier E (2014) The new frontier of genome engineering with CRISPR-Cas9. Science 346:1258096
Zhang J-H, Adikaram P, Pandey M, Genis A, Simonds WF (2016) Optimization of genome editing through CRISPR-Cas9 engineering. Bioengineered 7:166–174
Gaillochet C, Develtere W, Jacobs TB (2021) CRISPR screens in plants: approaches, guidelines, and future prospects. Plant Cell 33:794–813
Kuppu S, Ron M, Marimuthu MP et al (2020) A variety of changes, including CRISPR/Cas9-mediated deletions, in CENH3 lead to haploid induction on outcrossing. Plant Biotechnol J 18:2068–2080
Feng Z, Mao Y, Xu N et al (2014) Multigeneration analysis reveals the inheritance, specificity, and patterns of CRISPR/Cas-induced gene modifications in Arabidopsis. Proceedings of the National Academy of Sciences 111, 4632–4637
Mubarik MS, Wang X, Khan SH et al (2021) Engineering broad-spectrum resistance to cotton leaf curl disease by CRISPR-Cas9 based multiplex editing in plants. GM Crops & Food, pp 1–12
Chilcoat D, Liu Z-B, Sander J (2017) Use of CRISPR/Cas9 for crop improvement in maize and soybean. Prog Mol Biol Transl Sci 149:27–46
Johansen IE, Liu Y, Jørgensen B et al (2019) High efficacy full allelic CRISPR/Cas9 gene editing in tetraploid potato. Sci Rep 9:1–7
Zaidi SS-e-A, Tashkandi M, Mansoor S, Mahfouz MM (2016) Engineering plant immunity: using CRISPR/Cas9 to generate virus resistance. Front Plant Sci 7:1673
Zhang Q (2007) Strategies for developing green super rice. Proceedings of the national Academy of Sciences 104, 16402–16409
Zhang D, Zhang Z, Unver T, Zhang B (2020) CRISPR/Cas: A powerful tool for gene function study and crop improvement. Journal of Advanced Research
Sharma R, Upadhyay S, Bhattacharya S, Singh A (2021) Abiotic Stress-Responsive miRNA and Transcription Factor-Mediated Gene Regulatory Network in Oryza sativa: Construction and Structural Measure Study. Front Genet 12:130
Mickelbart MV, Hasegawa PM, Bailey-Serres J (2015) Genetic mechanisms of abiotic stress tolerance that translate to crop yield stability. Nat Rev Genet 16:237–251
Jain M (2015) Function genomics of abiotic stress tolerance in plants: a CRISPR approach. Front Plant Sci 6:375
Chen S, Zhang N, Zhang Q et al (2019) Genome editing to integrate seed size and abiotic stress tolerance traits in Arabidopsis reveals a role for DPA4 and SOD7 in the regulation of inflorescence architecture. Int J Mol Sci 20:2695
Cui Y, Jiang N, Xu Z, Xu Q (2020) Heterotrimeric G protein are involved in the regulation of multiple agronomic traits and stress tolerance in rice. BMC Plant Biol 20:1–13
Li S, Lin Y-CJ, Wang P et al (2019) The AREB1 transcription factor influences histone acetylation to regulate drought responses and tolerance in Populus trichocarpa. Plant Cell 31:663–686
Wang B, Xie G, Liu Z et al (2019) Mutagenesis reveals that the OsPPa6 gene is required for enhancing the alkaline tolerance in rice. Front Plant Sci 10:759
He J, Meng S, Zhao T et al (2017) An innovative procedure of genome-wide association analysis fits studies on germplasm population and plant breeding. Theor Appl Genet 130:2327–2343
Chen H-J, Su C-T, Lin C-H, Huang G-J, Lin Y-H (2010) Expression of sweet potato cysteine protease SPCP2 altered developmental characteristics and stress responses in transgenic Arabidopsis plants. J Plant Physiol 167:838–847
Zhou H, He M, Li J et al (2016) Development of commercial thermo-sensitive genic male sterile rice accelerates hybrid rice breeding using the CRISPR/Cas9-mediated TMS5 editing system. Sci Rep 6:1–12
Osakabe K, Osakabe Y, Toki S (2010) Site-directed mutagenesis in Arabidopsis using custom-designed zinc finger nucleases. Proceedings of the National Academy of Sciences 107, 12034–12039
Antoniou C, Chatzimichail G, Xenofontos R et al (2017) Melatonin systemically ameliorates drought stress-induced damage in M edicago sativa plants by modulating nitro‐oxidative homeostasis and proline metabolism.Journal of Pineal Research62, e12401
Xiao Y, Karikari B, Wang L, Chang F, Zhao T (2021) Structure characterization and potential role of soybean phospholipases A multigene family in response to multiple abiotic stress uncovered by CRISPR/Cas9 technology. Environ Exp Bot 188:104521
Ahmad A, Ashraf S, Munawar N, Jamil A, Ghaffar A, Shahbaz M (2021) CRISPR/Cas-Mediated Abiotic Stress Tolerance in Crops; in CRISPR Crops. Springer, pp 177–211
Hrbáčková M, Dvořák P, Takáč T et al (2020) Biotechnological perspectives of omics and genetic engineering methods in alfalfa. Front Plant Sci 11:592
Jaganathan D, Ramasamy K, Sellamuthu G, Jayabalan S, Venkataraman G (2018) CRISPR for crop improvement: an update review. Front Plant Sci 9:985
Varshney RK, Song C, Saxena RK et al (2013) Draft genome sequence of chickpea (Cicer arietinum) provides a resource for trait improvement. Nat Biotechnol 31:240–246
Millan T, Clarke HJ, Siddique KH et al (2006) Chickpea molecular breeding: new tools and concepts. Euphytica 147:81–103
Mantri NL, Ford R, Coram TE, Pang EC (2010) Evidence of unique and shared responses to major biotic and abiotic stresses in chickpea. Environ Exp Bot 69:286–292
Jha UC, Chaturvedi SK, Bohra A, Basu PS, Khan MS, Barh D (2014) Abiotic stresses, constraints and improvement strategies in chickpea. Plant Breeding 133:163–178
Garg R, Bhattacharjee A, Jain M (2015) Genome-scale transcriptomic insights into molecular aspects of abiotic stress responses in chickpea. Plant Mol Biology Report 33:388–400
Sabaghpour SH, Mahmodi AA, Saeed A, Kamel M, Malhotra R (2006) Study on chickpea drought tolerance lines under dryland condition of Iran. Indian J Crop Sci 1:70–73
Mafakheri A, Siosemardeh A, Bahramnejad B, Struik P, Sohrabi Y (2010) Effect of drought stress on yield, proline and chlorophyll contents in three chickpea cultivars. Aust J Crop Sci 4:580–585
Rahbarian R, Khavari-Nejad R, Ganjeali A, Bagheri A, Najafi F (2011) Drought stress effects on photosynthesis, chlorophyll fluorescence and water relations in tolerant and susceptible chickpea (Cicer arietinum L.) genotypes. Acta Biologica Cracoviensia s, Botanica
Barnabás B, Jäger K, Fehér A (2008) The effect of drought and heat stress on reproductive processes in cereals. Plant Cell Environ 31:11–38
Singh A, Nath O, Singh S, Kumar S, Singh IK (2018) Genome-wide identification of the MAPK gene family in chickpea and expression analysis during development and stress response. Plant Gene 13:25–35
Azeem F, Ahmad B, Atif RM et al (2018) Genome-wide analysis of potassium transport-related genes in chickpea (Cicer arietinum L.) and their role in abiotic stress responses. Plant Mol Biology Report 36:451–468
Mugabe D, Coyne CJ, Piaskowski J et al (2019) Quantitative trait loci for cold tolerance in chickpea. Crop Sci 59:573–582
Jaudal M, Zhang L, Che C et al (2016) Mt VRN 2 is a Polycomb VRN 2-like gene which represses the transition to flowering in the model legume Medicago truncatula. Plant J 86:145–160
Samineni S, Kamatam S, Thudi M, Varshney RK, Gaur PM (2016) Vernalization response in chickpea is controlled by a major QTL. Euphytica 207:453–461
Saradadevi R, Palta JA, Siddique KH (2017) ABA-mediated stomatal response in regulating water use during the development of terminal drought in wheat. Front Plant Sci 8:1251
Paul PJ, Samineni S, Thudi M et al (2018) Molecular mapping of QTLs for heat tolerance in chickpea. Int J Mol Sci 19:2166
Hayes B, Bowman P, Chamberlain A, God dard ME (2009) Genomic selection in dairy cattle: Progress and challenges. J Dairy Sci 92:433–443
Fleury D, Jefferies S, Kuchel H, Langridge P (2010) Genetic and genomic tools to improve drought tolerance in wheat. J Exp Bot 61:3211–3222
Sharma KD, Nayyar H (2014) Cold stress alters transcription in meiotic anthers of cold tolerant chickpea (Cicer arietinum L.). BMC Res Notes 7:1–13
Agarwal G, Garg V, Kudapa H et al (2016) Genome-wide dissection of AP2/ERF and HSP90 gene families in five legumes and expression profiles in chickpea and pigeonpea. Plant Biotechnol J 14:1563–1577
Garg R, Shankar R, Thakkar B et al (2016) Transcriptome analyses reveal genotype-and developmental stage-specific molecular responses to drought and salinity stresses in chickpea. Sci Rep 6:1–15
Kudapa H, Garg V, Chitikineni A, Varshney RK (2018) The RNA-Seq‐based high resolution gene expression atlas of chickpea (Cicer arietinum L.) reveals dynamic spatio‐temporal changes associated with growth and development. Plant Cell Environ 41:2209–2225
Kumar M, Chauhan AS, Yusuf MA, Sanyal I, Chauhan PS (2019) Transcriptome sequencing of chickpea (Cicer arietinum L.) genotypes for identification of drought-responsive genes under drought stress condition. Plant Mol Biology Report 37:186–203
Jain M, Misra G, Patel RK et al (2013) A draft genome sequence of the pulse crop chickpea (C icer arietinum L.). Plant J 74:715–729
Deokar AA, Kondawar V, Kohli D et al (2015) The CarERF genes in chickpea (Cicer arietinum L.) and the identification of CarERF116 as abiotic stress responsive transcription factor. Funct Integr Genom 15:27–46
Sen S, Chakraborty J, Ghosh P, Basu D, Das S (2017) Chickpea WRKY70 regulates the expression of a homeodomain-leucine zipper (HD-zip) I transcription factor CaHDZ12, which confers abiotic stress tolerance in transgenic tobacco and chickpea. Plant Cell Physiol 58:1934–1952
Acharjee S, Sarmah BK (2013) Biotechnologically generating ‘super chickpea’for food and nutritional security. Plant Sci 207:108–116
Bhatnagar-Mathur P, Vadez V, Devi MJ, Lavanya M, Vani G, Sharma KK (2009) Genetic engineering of chickpea (Cicer arietinum L.) with the P5CSF129A gene for osmoregulation with implications on drought tolerance. Mol Breeding 23:591–606
Rani A, Devi P, Jha UC, Sharma KD, Siddique KH, Nayyar H (2020) Developing climate-resilient chickpea involving physiological and molecular approaches with a focus on temperature and drought stresses. Front Plant Sci 10:1759
Das Bhowmik SS, Cheng AY, Long H et al (2019) Robust genetic transformation system to obtain non-chimeric transgenic chickpea. Front Plant Sci 10:524
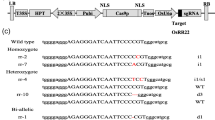
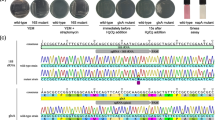
Badhan S, Ball AS, Mantri N (2021) First report of CRISPR/Cas9 mediated DNA-free editing of 4CL and RVE7 genes in chickpea protoplasts. Int J Mol Sci 22:396
Montague TG, Cruz JM, Gagnon JA, Church GM, Valen E (2014) CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res 42:W401–W407
Lei Y, Lu L, Liu H-Y, Li S, Xing F, Chen L-L (2014) CRISPR-P: a web tool for synthetic single-guide RNA design of CRISPR-system in plants. Mol Plant 7:1494–1496
Li H, Yang Y, Hong W, Huang M, Wu M, Zhao X (2020) Applications of genome editing technology in the targeted therapy of human diseases: mechanisms, advances and prospects. Signal Transduct Target therapy 5:1–23
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have declared that no competing or conflicts of interest exist.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Razzaq, M.K., Akhter, M., Ahmad, R.M. et al. CRISPR-Cas9 based stress tolerance: New hope for abiotic stress tolerance in chickpea (Cicer arietinum). Mol Biol Rep 49, 8977–8985 (2022). https://doi.org/10.1007/s11033-022-07391-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07391-4