Abstract
Mitochondria and chloroplast are very important organelles for organism, participating in basic life activity. Their genomes contain many repeats which can lead to a variation of genome structure. Oryza is an important genus for human beings’ nutrition. Several mitochondrial and chloroplast genomes of Oryza have been sequenced, which help us to insight the distribution and evolution of the repeats in Oryza species. In this paper, we compared six mitochondrial and 13 chloroplast genomes of Oryza and found that the structures of mitochondrial genomes were more diverse than chloroplast genomes. Since repeats can change the structure of the genome, resulting in the structural diversity of the genome, we analyzed all repeats and found 31 repeats in mitochondrial and 13 repeats in chloroplast genomes. Further, we developed 21 pairs of MRS molecular markers and 12 pairs of CRS molecular markers based on mitochondrial repeats and chloroplast repeats, respectively. These molecular markers can be used to detect the repeat-mediated recombination in Oryza mitochondrial and chloroplast genomes by PCR or fluorescence quantification.





Similar content being viewed by others
References
Altschul SFGW, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Alverson AJ, Wei X, Rice DW, Stern DB, Barry K, Palmer JD (2010) Insights into the evolution of mitochondrial genome size from complete sequences of Citrullus lanatus and Cucurbita pepo (Cucurbitaceae). Mol Biol Evol 27:1436–1448
Alverson AJ, Rice DW, Dickinson S, Barry K, Palmer JD (2011) Origins and recombination of the bacterial-sized multichromosomal mitochondrial genome of cucumber. Plant Cell 23:2499–2513
Arrieta-Montiel MP, Mackenzie SA (2011) Plant mitochondrial genomes and recombination. In: Kempken F (ed) Plant mitochondria. Springer, New York, pp 65–82
Arrieta-Montiel MP, Shedge V, Davila J, Christensen AC, Mackenzie SA (2009) Diversity of the Arabidopsis mitochondrial genome occurs via nuclear-controlled recombination activity. Genetics 183:1261–1268
Beaudet D, Terrat Y, Halary S, de la Providencia IE, Hijri M (2013) Mitochondrial genome rearrangements in glomus species triggered by homologous recombination between distinct mtDNA haplotypes. Genome Biol Evol 5:1628–1643
Brazda V, Lysek J, Bartas M, Fojta M (2018) Complex analyses of short inverted repeat sequences in all sequenced chloroplast DNAs. Biomed Res Int 2018:1097018
Caparroz R, Rocha AV, Cabanne GS, Tubaro P, Aleixo A, Lemmon EM, Lemmon AR (2018) Mitogenomes of two neotropical bird species and the multiple independent origin of mitochondrial gene orders in Passeriformes. Mol Biol Rep 45:279–285
Cechova J, Lysek J, Bartas M, Brazda V (2018) Complex analyses of inverted repeat sequences in mitochondrial genomes revealed their importance and variability. Bioinformatics 34:1081–1085
Choi KS, Chung MG, Park S (2016) The complete chloroplast genome sequences of three Veroniceae Species (Plantaginaceae): comparative analysis and highly divergent regions. Front Plant Sci 7:355
Cole LW, Guo W, Mower JP, Palmer JD (2018) High and variable rates of repeat-mediated mitochondrial genome rearrangement in a genus of plants. Mol Biol Evol 35:2773–2785
Daniell H, Lin CS, Yu M, Chang WJ (2016) Chloroplast genomes: diversity, evolution, and applications in genetic engineering. Genome Biol 17:134
Darling AE, Mau B, Perna NT (2010) progressiveMauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS One 5:e11147
Duminil J (2014) Mitochondrial genome and plant taxonomy. Methods Mol Biol 1115:121–140
Gu C, Tembrock LR, Zheng S, Wu Z (2018) The complete chloroplast genome of Catha edulis: a comparative analysis of genome features with related species. Int J Mol Sci 19:525
Guo W, Zhu A, Fan W, Mower JP (2016) Complete mitochondrial genomes from the ferns Ophioglossum californicum and Psilotum nudum are highly repetitive with the largest organellar introns. New Phytol 213:391–403
Jansen RK, Cai Z, Raubeson LA, Daniell H, Depamphilis CW, Leebens-Mack J, Muller KF, Guisinger-Bellian M, Haberle RC, Hansen AK, Chumley TW, Lee SB, Peery R, McNeal JR, Kuehl JV, Boore JL (2007) Analysis of 81 genes from 64 plastid genomes resolves relationships in angiosperms and identifies genome-scale evolutionary patterns. Proc Natl Acad Sci U S A 104:19369–19374
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30:3059–3066
Lohse M, Drechsel O, Kahlau S, Bock R (2013) OrganellarGenomeDRAW--a suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucleic Acids Res 41:W575–W581
Miller-Messmer M, Kuhn K, Bichara M, Le Ret M, Imbault P, Gualberto JM (2012) RecA-dependent DNA repair results in increased heteroplasmy of the Arabidopsis mitochondrial genome. Plant Physiol 159:211–226
Park S, An B, Park S (2018) Reconfiguration of the plastid genome in Lamprocapnos spectabilis: IR boundary shifting, inversion, and intraspecific variation. Sci Rep 8:13568
Rogers SO, Bendich AJ (1985) Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues. Plant Mol Biol 5:69–76
Shearman JR, Sonthirod C, Naktang C, Pootakham W, Yoocha T, Sangsrakru D, Jomchai N, Tragoonrung S, Tangphatsornruang S (2016) The two chromosomes of the mitochondrial genome of a sugarcane cultivar: assembly and recombination analysis using long PacBio reads. Sci Rep 6:31533
Ge S, Sang T, Lu BR, Hong DY (1999) Phylogeny of rice genomes with emphasis on origins of allotetraploid species. Proc Natl Acad Sci U S A 96:14400–14405
Schattner P, Brooks AN, Lowe TM (2005) The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res 33:W686–W689
Sun Y, Moore MJ, Lin N, Adelalu KF, Meng A, Jian S, Yang L, Li J, Wang H (2017) Complete plastome sequencing of both living species of Circaeasteraceae (Ranunculales) reveals unusual rearrangements and the loss of the ndh gene family. BMC Genomics 18:592
Untergasser A, Cutcutache I, Koressaar T, Ye J, Faircloth BC, Remm M, Rozen SG (2012) Primer3--new capabilities and interfaces. Nucleic Acids Res 40:e115
Wallet C, Le Ret M, Bergdoll M, Bichara M, Dietrich A, Gualberto JM (2015) The RECG1 DNA translocase is a key factor in recombination surveillance, repair, and segregation of the mitochondrial DNA in Arabidopsis. Plant Cell 27:2907–2925
Wang X, Zhou T, Bai G, Zhao Y (2018) Complete chloroplast genome sequence of Fagopyrum dibotrys: genome features, comparative analysis and phylogenetic relationships. Sci Rep 8:12379
Weng ML, Blazier JC, Govindu M, Jansen RK (2014) Reconstruction of the ancestral plastid genome in Geraniaceae reveals a correlation between genome rearrangements, repeat sequences, and nucleotide substitution rates. Genome Biol Evol 31:645–659
Wicke S, Schneeweiss GM, dePamphilis CW, Muller KF, Quandt D (2011) The evolution of the plastid chromosome in land plants: gene content, gene order, gene function. Plant Mol Biol 76:273–297
Wyman SK, Jansen RK, Boore JL (2004) Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20:3252–3255
Xu YZ, Arrieta-Montiel MP, Virdi KS, de Paula WB, Widhalm JR, Basset GJ, Davila JI, Elthon TE, Elowsky CG, Sato SJ, Clemente TE, Mackenzie SA (2011) MutS HOMOLOG1 is a nucleoid protein that alters mitochondrial and plastid properties and plant response to high light. Plant Cell 23:3428–3441
Yang WL, Zou JN, Wang JJ, Li NW, Luo XY, Jiang XF, Li SQ (2020) Wide crossing diversify mitogenomes of rice. BMC Plant Biol 20:159
Yang Y, Zhou T, Duan D, Yang J, Feng L, Zhao G (2016) Comparative analysis of the complete chloroplast genomes of five Quercus species. Front Plant Sci 7:959
Yue F, Cui L, de Pamphilis CW, Moret BM, Tang J (2008) Gene rearrangement analysis and ancestral order inference from chloroplast genomes with inverted repeat. BMC Genomics 9(Suppl 1):S25
Zhang YJ, Ma PF, Li DZ (2011) High-throughput sequencing of six bamboo chloroplast genomes: phylogenetic implications for temperate woody bamboos (Poaceae: Bambusoideae). PLoS One 6:e20596
Zhao Q, Feng Q, Lu H, Li Y, Wang A, Tian Q, Zhan Q, Lu Y, Zhang L, Huang T, Wang Y, Fan D, Zhao Y, Wang Z, Zhou C, Chen J, Zhu C, Li W, Weng Q, Xu Q, Wang ZX, Wei X, Han B, Huang X (2018) Pan-genome analysis highlights the extent of genomic variation in cultivated and wild rice. Nat Genet 50:278–284
Acknowledgments
We thank Dr. Hongwei Xie and Chunyu Zhang for providing the wild rice accessions and Brassica varieties, respectively.
Funding
This work was partly granted by the National Key Research and Development Program (2016YFD0100903), the National Science Foundation (31571630) of China, and Huanghe Patent Program of Wuhan City.
Author information
Authors and Affiliations
Contributions
S.L. and W.Y. conceived and designed the research. W.Y., J.Z., Y.Y., and W.L. performed the research. W.Y., J.Z., and Y.Y. analyzed the data. W.Y. wrote the manuscript. S.L., J.Z., Y.Y., and W.L. revised the manuscript. All authors have read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Fig. S1
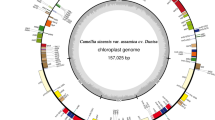
Mitochondrial and Chloroplast genomes of Oryza species. a. Mitochondrial genomes of six Oryza species. b. Chloroplast genome of thirteen Oryza species. The genes drawn outside of the circle are transcribed counterclockwise, while those inside are clockwise. Small single copy (SSC), large single copy (LSC), and inverted repeats (IRa, IRb) are indicated. The dark gray and light gray shading within the inner circle correspond to the percentage of G + C and A + T content, respectively. Gene function or identifiers are displayed using colors indicated by the inner legend (JPG 4462 kb)
Fig. S2
Detection of the stability of molecular markers in different individuals, tissues and developmental stages (JPG 2086 kb)
Fig. S3
Detection of the usability of molecular markers in Brassica (JPG 5701 kb)
Table S1
(DOCX 15 kb)
Table S2
(DOCX 16 kb)
Table S3
(DOCX 16 kb)
Table S4
(DOCX 15 kb)
ESM 1
(XLSX 73 kb)
ESM 2
(TXT 5 kb)
ESM 3
(TXT 509 bytes)
Rights and permissions
About this article
Cite this article
Yang, W., Zou, J., Yu, Y. et al. Repeats in mitochondrial and chloroplast genomes characterize the ecotypes of the Oryza. Mol Breeding 41, 7 (2021). https://doi.org/10.1007/s11032-020-01198-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-020-01198-6