Abstract
Virtual screening a collection of ~ 25,000 ChemBridge molecule collection identified two nitrogenous heterocyclic molecules, 12 and 15, with potential dual inhibitory properties against trypanosomal cruzain and rhodesain cysteine proteases. Similarity search in DrugBank found the two virtual hits with novel chemical structures with unreported anti-trypanosomal activities. Investigations into the binding mechanism by molecular dynamics simulations for 100 ns revealed the molecules were able to occupy the binding sites and stabilise the protease complexes. Binding affinities calculated using the MM/PBSA method for the last 20 ns showed that the virtual hits have comparable binding affinities to other known inhibitors from literature suggesting both molecules as promising scaffolds with dual cruzain and rhodesain inhibition properties, i.e. 12 has predicted ΔGbind values of − 38.1 and − 38.2 kcal/mol to cruzain and rhodesain, respectively, and 15 has predicted ΔGbind values of − 34.4 and − 25.8 kcal/mol to rhodesain. Per residue binding free energy decomposition studies and visual inspection at 100 ns snapshots revealed hydrogen bonding and non-polar attractions with important amino acid residues that contributed to the ΔGbind values. The interactions are similar to those previously reported in the literature. The overall ADMET predictions for the two molecules were favourable for drug development with acceptable pharmacokinetic profiles and adequate oral bioavailability.
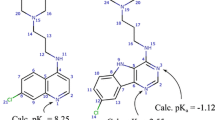
Graphical Abstract













Similar content being viewed by others
Data availability
All data generated from this work is included in this published article and supplementary information. Any materials related to the work may be requested from the corresponding author.
Code availability
Not applicable.
References
Baker CH, Welburn SC (2018) The long wait for a new drug for human African Trypanosomiasis. Trends Parasitol 34:818–27. https://doi.org/10.1016/j.pt.2018.08.006
World Health Organization (2021) Trypanosomiasis, human African (sleeping sickness). https://www.who.int/news-room/fact-sheets/detail/trypanosomiasis-human-african-(sleeping-sickness). Accessed 15 December 2021
De Koning PH (2020) the drugs of sleeping sickness: their mechanisms of action and resistance, and a brief history. Trop Med Infect Dis 5(1):14. https://doi.org/10.3390/tropicalmed5010014
Sajid M, McKerrow JH (2002) Cysteine proteases of parasitic organisms. Mol Biochem Parasitol 120:1–21. https://doi.org/10.1016/S0166-6851(01)00438-8
Ferreira LG, Andricopulo AD (2017) Targeting cysteine proteases in trypanosomatid disease drug discovery. Pharmacol Ther 180:49–61. https://doi.org/10.1016/j.pharmthera.2017.06.004
Di Chio C, Previti S et al (2020) Development of novel benzodiazepine-based peptidomimetics as inhibitors of rhodesain from trypanosoma brucei rhodesiense. ChemMedChem 15:995–1001. https://doi.org/10.1002/cmdc.202000158
Engel JC, Doyle PS, Hsieh I, McKerrow JH (1998) Cysteine protease inhibitors cure an experimental Trypanosoma cruzi infection. J Exp Med 188:725–734. https://doi.org/10.1084/jem.188.4.725
Fujii N, Mallari JP, Hansell EJ et al (2005) Discovery of potent thiosemicarbazone inhibitors of rhodesain and cruzain. Bioorg Med Chem Lett 15:121–123. https://doi.org/10.1016/j.bmcl.2004.10.023
Ehmke V, Winkler E et al (2013) Optimization of triazine nitriles as rhodesain inhibitors: structure-activity relationships, bioisosteric imidazopyridine nitriles, and X-ray crystal structure analysis with human cathepsin L. ChemMedChem 8:967–975. https://doi.org/10.1002/cmdc.201300112
Neitz RJ, Bryant C et al (2015) Tetrafluorophenoxymethyl ketone cruzain inhibitors with improved pharmacokinetic properties as therapeutic leads for Chagas’ disease. Bioorg Med Chem Lett 25:4834–4837. https://doi.org/10.1016/j.bmcl.2015.06.066
Makhoba XH, Viegas C Jr, Mosa RA, Viegas FPD, Pooe OJ (2020) potential impact of the multi-target drug approach in the treatment of some complex diseases. Drug Des Devel Ther 14:3235–3249. https://doi.org/10.2147/DDDT.S257494
Khare S, Nagle AS et al (2016) Proteasome inhibition for treatment of leishmaniasis, Chagas disease and sleeping sickness. Nature 537:229–233. https://doi.org/10.1038/nature19339
Lionta E, Spyrou G, Vassilatis DK, Cournia Z (2014) Structure-based virtual screening for drug discovery: principles, applications and recent advances. Curr Top Med Chem 14:1923–1938. https://doi.org/10.2174/1568026614666140929124445
Homeyer N, Gohlke H (2012) Free energy calculations by the molecular mechanics poisson−boltzmann surface area method. Mol Inform 31:114–122. https://doi.org/10.1002/minf.201100135
Allinger NL (1977) Conformational analysis. 130. MM2. A hydrocarbon force field utilizing V1 and V2 torsional terms. J Am Chem Soc 99:8127–8134. https://doi.org/10.1021/ja00467a001
McGrath ME, Eakin AE, Engel JC, McKerrow JH, Craik CS, Fletterick RJ (1995) The crystal structure of cruzain: a therapeutic target for Chagas’ disease. J Mol Biol 247:251–259. https://doi.org/10.1006/jmbi.1994.0137
Berman HM, Westbrook J et al (2000) The protein data bank. Nucleic Acids Res 28:235–242. https://doi.org/10.1093/nar/28.1.235
Mooij WT, Verdonk ML (2005) General and targeted statistical potentials for protein-ligand interactions. Proteins 61:272–287. https://doi.org/10.1002/prot.20588
Jones G, Willett P, Glen RC, Leach AR, Taylor R (1997) Development and validation of a genetic algorithm for flexible docking. J Mol Biol 267:727–748. https://doi.org/10.1006/jmbi.1996.0897
Abraham MJ, van der Spoel D, Lindahl E, Hess B, Gromacs development team team (2016) GROMACS User Manual version 5.1.4
Berendsen HJC, van der Spoel D, van Drunen R (1995) GROMACS: a message-passing parallel molecular dynamics implementation. Comput Phys Commun 91:43–56. https://doi.org/10.1016/0010-4655(95)00042-E
Malde AK, Zuo L et al (2011) An automated force field topology builder (ATB) and repository: version 1.0. J Chem Theory Comput 7:4026–4037. https://doi.org/10.1021/ct200196m
Stroet M, Caron B, Visscher KM, Geerke DP, Malde AK, Mark AE (2018) Automated topology builder version 3.0: prediction of solvation free enthalpies in water and hexane. J Chem Theory Comput 14:5834–5845. https://doi.org/10.1021/acs.jctc.8b00768
Schmid N, Eichenberger AP et al (2011) Definition and testing of the GROMOS force-field versions 54A7 and 54B7. Eur Biophys J 40:843–856. https://doi.org/10.1007/s00249-011-0700-9
Berweger CD, van Gunsteren WF, Müller-Plathe F (1995) Force field parametrization by weak coupling. Re-engineering SPC water Chem Phys Lett 232:429–436. https://doi.org/10.1016/0009-2614(94)01391-8
Berendsen HJC, Postma JPM, van Gunsteren WF, DiNola A, Haak JR (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81:3684–3690. https://doi.org/10.1063/1.448118
Parrinello M, Rahman A (1981) Polymorphic transitions in single crystals: a new molecular dynamics method. J Appl Phys 52:7182–7190. https://doi.org/10.1063/1.328693
Hess B, Bekker H, Berendsen HJC, Fraaije JGEM (1997) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18:1463–1472. https://doi.org/10.1002/(SICI)1096-987X(199709)18:12%3c1463::AID-JCC4%3e3.0.CO;2-H
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an Nṡlog(N) method for Ewald sums in large systems. J Chem Phys 98:10089. https://doi.org/10.1063/1.464397
Kumari R, Kumar R, Lynn A (2014) g_mmpbsa—A GROMACS tool for high-throughput MM-PBSA calculations. J Chem Info Model 54:1951–1962. https://doi.org/10.1021/ci500020m
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7:42717. https://doi.org/10.1038/srep42717
Giraldo C, Gómez S et al (2016) Insight into the mechanism of the Michael reaction. ChemPhysChem 17:2022–2034. https://doi.org/10.1002/cphc.201600166
Wishart DS, Knox C et al (2008) DrugBank: a knowledgebase for drugs, drug actions and drug targets. Nucleic Acids Res 36:D901–D906. https://doi.org/10.1093/nar/gkm958
Lang DK, Kaur R, Arora R, Saini B, Arora S (2020) Nitrogen-containing heterocycles as anticancer agents: an overview. Anticancer Agents Med Chem 20:2150–2168. https://doi.org/10.2174/1871520620666200705214917
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (1997) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 23:3–25. https://doi.org/10.1016/S0169-409X(96)00423-1
Veber DF, Johnson SR, Cheng HY, Smith BR, Ward KW, Kopple KD (2002) Molecular properties that influence the oral bioavailability of drug candidates. J Med Chem 45:2615–2623. https://doi.org/10.1021/jm020017n
Martin YC (2005) A bioavailability score. J Med Chem 48:3164–3170. https://doi.org/10.1021/jm0492002
Ditzinger F, Price DJ et al (2019) Lipophilicity and hydrophobicity considerations in bioenabling oral formulations approaches—a PEARRL review. J Pharm Pharmacol 71:464–482. https://doi.org/10.1111/jphp.12984
Ertl P, Schuffenhauer A (2009) Estimation of synthetic accessibility score of drug-like molecules based on molecular complexity and fragment contributions. J Cheminformatics 1:8. https://doi.org/10.1186/1758-2946-1-8
Ferreira RS, Simeonov A et al (2010) Complementarity between a docking and a high- throughput screen in discovering new cruzain inhibitors. J Med Chem 53:4891–4905. https://doi.org/10.1021/jm100488w
Silva LR, Guimarães AS et al (2021) Computer-aided design of 1,4-naphthoquinone-based inhibitors targeting cruzain and rhodesain cysteine proteases. Bioorg Med Chem 41:116213. https://doi.org/10.1016/j.bmc.2021.116213
Ogungbe IV, Setzer WN (2009) Comparative molecular docking of antitrypanosomal natural products into multiple Trypanosoma brucei drug targets. Molecules 14(4):1513–1536. https://doi.org/10.3390/molecules14041513
de Souza ML, de Oliveira Rezende C et al (2020) Discovery of potent, reversible, and competitive cruzain inhibitors with trypanocidal activity: a structure-based drug design approach. J Chem Inf Model 60:1028–1041. https://doi.org/10.1021/acs.jcim.9b00802
Wiggers HJ, Rocha JR et al (2013) Non-peptidic cruzain inhibitors with trypanocidal activity discovered by virtual screening and in vitro assay. PLoS Negl Trop Dis 7:e2370. https://doi.org/10.1371/journal.pntd.0002370
Ferreira RAA, Pauli I et al (2019) Structure-based and molecular modeling studies for the discovery of cyclic imides as reversible cruzain inhibitors with potent anti-trypanosoma cruzi activity. Front Chem 7:798. https://doi.org/10.3389/fchem.2019.00798
Rogers KE, Keränen H et al (2012) Novel cruzain inhibitors for the treatment of Chagas’ disease. Chem Biol Drug Des 80:398–405. https://doi.org/10.1111/j.1747-0285.2012.01416.x
Funding
This research work is partially supported by the Chulabhorn Royal Academy research grant project code 631-CS01. This research work is supported in part by the grant from the Center of Excellence on Environmental Health and Toxicology (EHT), OPS, Ministry of Higher Education, Science Research and Innovation. The authors would like to thank Center of Excellence on Environmental Health and Toxicology (EHT), OPS, Ministry of Higher Education, Science, Research and Innovation for their excellent technical support.
Author information
Authors and Affiliations
Contributions
The experimental procedures, planning and analysis were performed by CE, CZ, TS and ASB. The manuscript draft was prepared and reviewed by CE, CB, RS, WN, TS, SR and ASB. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare no competing interests.
Ethical approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Eurtivong, C., Zimmer, C., Schirmeister, T. et al. A structure-based virtual high-throughput screening, molecular docking, molecular dynamics and MM/PBSA study identified novel putative drug-like dual inhibitors of trypanosomal cruzain and rhodesain cysteine proteases. Mol Divers (2023). https://doi.org/10.1007/s11030-023-10600-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11030-023-10600-2