Abstract
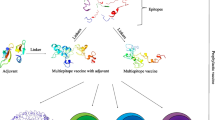
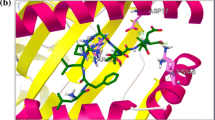
Infections with HCV, HBV and poliovirus are still considered to be substantial global health burdens. Vaccination is one of the most important preventive strategies against these infections. Multi-epitope vaccines are presented as novel strategies to circumvent the limitations associated with conventional vaccines. Given these circumstances, a multi-epitope protein was designed using the predicted high score epitopes of the antigens from HCV, HBV and Poliovirus. To this end, the sequences of HCV core protein, HBV small surface antigen and VPs of Poliovirus were collected and the consensus sequence of these antigens were obtained using BLAST and MSA analyses. Then, the physicochemical properties of these antigens along with their high score B and T-cell epitopes were predicted using various softwares. The obtained epitopes were connected with proper linkers to build the final 500 amino acids HHP protein. The secondary and tertiary structure of the HHP as well as its physicochemical properties and immunological properties were predicted using different tools. Assessment of various properties of the designed protein indicated that the HHP poly-epitope is an immunogenic and non-allergen antigen, which can derive humoral and cellular immune responses against HCV, HBV and Poliovirus infections.









Similar content being viewed by others
References
Adu F (2005) The virology of the polio virus. Ann Ibadan Postgrad Med 3(1):13–19
Ali M, Pandey RK, Khatoon N, Narula A, Mishra A, Prajapati VK (2017) Exploring dengue genome to construct a multi-epitope based subunit vaccine by utilizing immunoinformatics approach to battle against dengue infection. Sci Rep 7(1):9232
Arashkia A, Roohvand F, Memarnejadian A, Aghasadeghi MR, Rafati S (2010) Construction of HCV-polytope vaccine candidates harbouring immune-enhancer sequences and primary evaluation of their immunogenicity in BALB/c mice. Virus Genes 40(1):44
Bai Y, Shen W-C (2006) Improving the oral efficacy of recombinant granulocyte colony-stimulating factor and transferrin fusion protein by spacer optimization. Pharma Res 23(9):2116–2121
Benkert P, Biasini M, Schwede T (2010) Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics 27(3):343–350
Bhasin M, Lata S, Raghava GP (2007) TAPPred prediction of TAP-binding peptides in antigens. Methods Mol Biol 409:381–386
Chauhan V, Rungta T, Goyal K, Singh MP (2019) Designing a multi-epitope based vaccine to combat Kaposi Sarcoma utilizing immunoinformatics approach. Sci Rep 9(1):2517
Chen VB, Arendall WB, Headd JJ, Keedy DA, Immormino RM, Kapral GJ et al (2010) MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr Sect D 66(1):12–21
DeLano WL (2002) Pymol: an open-source molecular graphics tool. CCP4 Newsletter on protein crystallography. 40:82–92
Delpeyroux F, Chenciner N, Lim A, Malpiece Y, Blondel B, Crainic R et al (1986) A poliovirus neutralization epitope expressed on hybrid hepatitis B surface antigen particles. Science 233(4762):472–475
Delpeyroux F, Peillon N, Blondel B, Crainic R, Streeck R (1988) Presentation and immunogenicity of the hepatitis B surface antigen and a poliovirus neutralization antigen on mixed empty envelope particles. J Virol 62(5):1836–1839
Dimitrov I, Bangov I, Flower DR, Doytchinova I (2014) AllerTOP v. 2—a server for in silico prediction of allergens. J Mol Model 20(6):2278
Doytchinova IA, Flower DR (2007) VaxiJen: a server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinformatics 8(1):4
Drozdetskiy A, Cole C, Procter J, Barton GJ (2015) JPred4: a protein secondary structure prediction server. Nucleic Acids Res 43(W1):W389–W394
Emini EA, Jameson BA, Wimmer E (1984) Identification of a new neutralization antigenic site on poliovirus coat protein VP2. J Virol 52(2):719–721
Ende AR, Kim NH, Yeh MM, Harper J, Landis CS (2015) Fulminant hepatitis B reactivation leading to liver transplantation in a patient with chronic hepatitis C treated with simeprevir and sofosbuvir: a case report. J Med Case Rep 9(1):164
Fong TL, Di Bisceglie AM, Waggoner JG, Banks SM, Hoofnagle JH (1991) The significance of antibody to hepatitis C virus in patients with chronic hepatitis B. Hepatology 14(1):64–67
Ganji M, Khalili S, Mard-Soltani M, Khalesi B, Karkhah A, Amani J. A precisely designed immunotoxin against VCAM1 consisting of a humanized antibody variable domain fused to granzyme: an in silico approach. Int J Pept Res Ther. 2019:1–9
Gasteiger E, Hoogland C, Gattiker A, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the ExPASy server. The proteomics protocols handbook. Springer, Berlin, pp. 571–607
Geourjon C, Deleage G (1995) SOPMA: significant improvements in protein secondary structure prediction by consensus prediction from multiple alignments. Bioinformatics 11(6):681–684
Guan P, Hattotuwagama CK, Doytchinova IA, Flower DR (2006) MHCPred 2.0. Appl Bioinform 5(1):55–61
Guillen G, Aguilar J, Duenas S, Hermida L, Guzmán M, Penton E et al (2010) Virus-Like Particles as vaccine antigens and adjuvants: application to chronic disease, cancer immunotherapy and infectious disease preventive strategies. Procedia Vaccinol 2(2):128–133
He L, Cheng Y, Kong L, Azadnia P, Giang E, Kim J et al (2015) Approaching rational epitope vaccine design for hepatitis C virus with meta-server and multivalent scaffolding. Sci Rep 5:12501
Heo L, Park H, Seok C (2013) GalaxyRefine: protein structure refinement driven by side-chain repacking. Nucleic Acids Res 41(W1):W384–W388
Huang X-j, Lü X, Lei Y-f, Yang J, Yao M, Lan H-y et al (2013) Cellular immunogenicity of a multi-epitope peptide vaccine candidate based on hepatitis C virus NS5A, NS4B and core proteins in HHD-2 mice. J Virol Methods 189(1):47–52
Jespersen MC, Peters B, Nielsen M, Marcatili P (2017) BepiPred-2.0: improving sequence-based B-cell epitope prediction using conformational epitopes. Nucleic Acids Res 45(W1):W24–W29
Jorba J, Diop OM, Iber J, Henderson E, Sutter RW, Wassilak SG et al (2017) Update on vaccine-derived polioviruses—worldwide, January 2016–June 2017. MMWR Morbid Mortal Wkl Rep 66(43):1185
Kao J-H, Chen D-S (2002) Global control of hepatitis B virus infection. Lancet Infect Dis 2(7):395–403
Karpenko LI, Bazhan SI, Antonets DV, Belyakov IM (2014) Novel approaches in polyepitope T-cell vaccine development against HIV-1. Exp Rev Vaccines 13(1):155–173
Kazemi F, Arab SS, Mohajel N, Keramati M, Niknam N, Aslani MM et al (2018) Computational simulations assessment of mutations impact on streptokinase (SK) from a group G streptococci with enhanced activity–insights into the functional roles of structural dynamics flexibility of SK and stabilization of SK–µplasmin catalytic complex. J Biomol Struct Dyn. 28:1–12
Keşmir C, Nussbaum AK, Schild H, Detours V, Brunak S (2002) Prediction of proteasome cleavage motifs by neural networks. Protein Eng 15(4):287–296
Kew O, Pallansch M (2018) Breaking the last chains of poliovirus transmission: progress and challenges in global polio eradication. Annu Rev Virol 5(1):427–451
Khalili S, Rasaee MJ, Mousavi SL, Amani J, Jahangiri A, Borna H (2017) In silico prediction and in vitro verification of a novel multi-epitope antigen for HBV detection. Mol Genet Microbiol Virol 32(4):230–240
Khatoon N, Pandey RK, Prajapati VK (2017) Exploring Leishmania secretory proteins to design B and T cell multi-epitope subunit vaccine using immunoinformatics approach. Sci Rep 7(1):8285
Konstantinou D, Deutsch M (2015) The spectrum of HBV/HCV coinfection: epidemiology, clinical characteristics, viralinteractions and management. Ann Gastroenterol 28(2):221
Krieger E, Joo K, Lee J, Lee J, Raman S, Thompson J et al (2009) Improving physical realism, stereochemistry, and side-chain accuracy in homology modeling: four approaches that performed well in CASP8. Proteins 77(S9):114–122
Krogh A, Larsson B, Von Heijne G, Sonnhammer EL (2001) Predicting transmembrane protein topology with a hidden markov model: application to complete genomes1. J Mol Biol 305(3):567–580
Lüthy R, Bowie JU, Eisenberg D (1992) Assessment of protein models with three-dimensional profiles. Nature 356(6364):83
Magnan CN, Zeller M, Kayala MA, Vigil A, Randall A, Felgner PL et al (2010) High-throughput prediction of protein antigenicity using protein microarray data. Bioinformatics 26(23):2936–2943
Mard-Soltani M, Rasaee MJ, Khalili S, Sheikhi A, Hedayati M, Ghaderi-Zefrehi H et al (2018) The effect of differentially designed fusion proteins to elicit efficient anti-human thyroid stimulating hormone immune responses. Iran J Allergy Asthma Immunol 17(2):158–170
McGuffin LJ, Bryson K, Jones DT (2000) The PSIPRED protein structure prediction server. Bioinformatics 16(4):404–405
McLauchlan J (2000) Properties of the hepatitis C virus core protein: a structural protein that modulates cellular processes. J Viral Hepat 7(1):2–14
McMahon BJ (2014) Chronic hepatitis B virus infection. Med Clin North Am 98(1):39–54
Melief CJ, Van Der Burg SH (2008) Immunotherapy of established (pre) malignant disease by synthetic long peptide vaccines. Nat Rev Cancer 8(5):351
Memarnejadian A, Roohvand F (2010) Fusion of HBsAg and prime/boosting augment Th1 and CTL responses to HCV polytope DNA vaccine. Cell Immunol 261(2):93–98
Mishra S, Losikoff PT, Self AA, Terry F, Ardito MT, Tassone R et al (2014) Peptide-pulsed dendritic cells induce the hepatitis C viral epitope-specific responses of naïve human T cells. Vaccine 32(26):3285–3292
Mohamed SA, Mesa G (1997) Dual infection with hepatitis C and B viruses: clinical and histological study in Saudi patients. Hepato-gastroenterology 44(17):1404–1406
Mohammadzadeh S, Roohvand F, Memarnejadian A, Jafari A, Ajdary S, Salmanian A-H et al (2016) Co-expression of hepatitis C virus polytope–HBsAg and p19-silencing suppressor protein in tobacco leaves. Pharm Biol 54(3):465–473
Narula A, Pandey RK, Khatoon N, Mishra A, Prajapati VK (2018) Excavating chikungunya genome to design B and T cell multi-epitope subunit vaccine using comprehensive immunoinformatics approach to control chikungunya infection. Infect Genet Evol 61:4–15
Nezafat N, Ghasemi Y, Javadi G, Khoshnoud MJ, Omidinia E (2014) A novel multi-epitope peptide vaccine against cancer: an in silico approach. J Theor Biol 349:121–134
Offord V, Coffey T, Werling D (2010) LRRfinder: a web application for the identification of leucine-rich repeats and an integrative Toll-like receptor database. Dev Comp Immunol 34(10):1035–1041
Pan W, Chen DS, Lu YJ, Sun FF, Xu HW, Zhang YW et al (2017) Bioinformatic prediction of the epitopes of Echinococcus granulosus antigen 5. Biomed Rep 6(2):181–187
Papadopoulos N, Papavdi M, Pavlidou A, Konstantinou D, Kranidioti H, Kontos G et al (2018) Hepatitis B and C coinfection in a real-life setting: viral interactions and treatment issues. Ann Gastroenterol 31(3):365
Patient R, Hourioux C, Vaudin P, Pagès J-C, Roingeard P (2009) Chimeric hepatitis B and C viruses envelope proteins can form subviral particles: implications for the design of new vaccine strategies. New Biotechnol 25(4):226–234
Pelte C, Cherepnev G, Wang Y, Schoenemann C, Volk H-D, Kern F (2004) Random screening of proteins for HLA-A* 0201-binding nine-amino acid peptides is not sufficient for identifying CD8 T cell epitopes recognized in the context of HLA-A* 0201. J Immunol 172(11):6783–6789
PIWord L. JM 2003 structure validation by Calpha geometry: phi, psi and Cbeta deviation. Proteins.50:437450
Ponomarenko J, Bui H-H, Li W, Fusseder N, Bourne PE, Sette A et al (2008) ElliPro: a new structure-based tool for the prediction of antibody epitopes. BMC Bioinformatics 9(1):514
Qin Y, Liao P (2018) Hepatitis B virus vaccine breakthrough infection: surveillance of S gene mutants of HBV. Acta Virol 62(2):115–121
Ranjbar MM, Ghorban K, Alavian SM, Keyvani H, Dadmanesh M, Ardakany AR et al (2013) GB virus C/hepatitis G virus envelope glycoprotein E2: computational molecular features and immunoinformatics study. Hepat Mon 13(12):e15342
Reche PA, Glutting JP, Zhang H, Reinherz EL (2004) Enhancement to the RANKPEP resource for the prediction of peptide binding to MHC molecules using profiles. Immunogenetics 56(6):405–419
Reddy Chichili VP, Kumar V, Sivaraman J (2013) Linkers in the structural biology of protein–protein interactions. Protein Sci 22(2):153–167
Rizzetto M, Ciancio A (2008) Chronic HBV-related liver disease. Mol Aspects Med 29(1):72–84
Rui LX, Park YM, Choi JY, Kim BS, Jung G (1998) Detection of antibodies against DNA polymerase of hepatitis B virus in HBsAg-positive sera using ELISA. Korean J Intern Med 13(2):95–98
Saha S, Raghava G (ed) (2004) BcePr: prediction of continuous B-cell epitopes in antigenic sequences using physico-chemical properties. International conference on artificial immune systems, Springer
Saha S, Raghava G (2006a) AlgPred: prediction of allergenic proteins and mapping of IgE epitopes. Nucleic Acids Res 34(suppl_2):W202–W209
Saha S, Raghava G (2006b) Prediction of continuous B-cell epitopes in an antigen using recurrent neural network. Proteins 65(1):40–48
Scheel TK, Rice CM (2013) Understanding the hepatitis C virus life cycle paves the way for highly effective therapies. Nat Med 19(7):837
Schuler MM, Nastke MD, Stevanovikc S (2007) SYFPEITHI: database for searching and T-cell epitope prediction. Methods Mol Biol 409:75–93
Shahsavandi S, Ebrahimi MM, Sadeghi K, Mahravani H (2015) Design of a heterosubtypic epitope-based peptide vaccine fused with hemokinin-1 against influenza viruses. Virol Sin 30(3):200–207
Singh H, Raghava G (2001) ProPred: prediction of HLA-DR binding sites. Bioinformatics 17(12):1236–1237
Tan S-L. Hepatitis C (2006) Viruses: genomes and molecular biology. Horizon Scientific Press, Norwich
Taylor R (2000) Bioinformatics and molecular analysis section (BIMAS) HLA peptide binding predictions.
Vita R, Overton JA, Greenbaum JA, Ponomarenko J, Clark JD, Cantrell JR et al (2014) The immune epitope database (IEDB) 3.0. Nucleic acids research 43(D1):D405–D412
Wass MN, Kelley LA, Sternberg MJ (2010) 3DLigandSite: predicting ligand-binding sites using similar structures. Nucleic Acids Res 38(suppl_2):W469–W473
Wetz K, Habermehl K-O (1982) Specific cross-linking of capsid proteins to virus RNA by ultraviolet irradiation of poliovirus. J Gen Virol 59(2):397–401
Wiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35(suppl_2):W407–W410
Wriggers W, Chakravarty S, Jennings PA (2005) Control of protein functional dynamics by peptide linkers. Peptide Sci 80(6):736–746
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y (2015) The I-TASSER Suite: protein structure and function prediction. Nat Methods 12(1):7
Yazdanian M, Memarnejadian A, Mahdavi M, Motevalli F, Sadat SM, Vahabpour R et al. Evaluation of cellular responses for a chimeric HBsAg-HCV core DNA vaccine in BALB/c mice. Adv Biomed Res. 2015;4
Yim T-J, Tang S, Andino R (1996) Poliovirus recombinants expressing hepatitis B virus antigens elicited a humoral immune response in susceptible mice. Virology 218(1):61–70
Acknowledgements
This study was supported by Deputy of Shahid Beheshti University of Medical Sciences [Grant Number: 10221] and it was conducted at the Cellular and Molecular Biology Research Center, Shahid Beheshti University of Medical Sciences in partial fulfillment of the Ph.D thesis of Mrs. Armina Alagheband Bahrami.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The Authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Bahrami, A.A., Bandehpour, M., Khalesi, B. et al. Computational Design and Analysis of a Poly-Epitope Fusion Protein: A New Vaccine Candidate for Hepatitis and Poliovirus. Int J Pept Res Ther 26, 389–403 (2020). https://doi.org/10.1007/s10989-019-09845-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10989-019-09845-z