Abstract
Circadian rhythms are ubiquitous phenomena that recur daily in a self-sustaining, entrainable, and oscillatory manner, and orchestrate a wide range of molecular, physiological, and behavioral processes. Circadian clocks are comprised of a hierarchical network of central and peripheral clocks that generate, sustain, and synchronize the circadian rhythms. The functioning of the peripheral clock is regulated by signals from autonomic innervation (from the central clock), endocrine networks, feeding, and other external cues. The critical role played by circadian rhythms in maintaining both systemic and tissue-level homeostasis is well established, and disruption of the rhythm has direct consequence for human health, disorders, and diseases. Circadian oscillations in both pharmacokinetics and pharmacodynamic processes are known to affect efficacy and toxicity of several therapeutic agents. A variety of modeling approaches ranging from empirical to more complex systems modeling approaches have been applied to characterize circadian biology and its influence on drug actions, optimize time of dosing, and identify opportunities for pharmacological modulation of the clock mechanisms and their downstream effects. In this review, we summarize current understanding of circadian rhythms and its influence on physiology, pharmacology, and therapeutic interventions, and discuss the role of chronopharmacometrics in gaining new insights into circadian rhythms and its applications in chronopharmacology.
Similar content being viewed by others
Introduction
Circadian biorhythms are ubiquitous phenomena that recur daily in a self-sustaining, entrainable, and oscillatory manner in most organisms. Circadian rhythms orchestrate a wide array of complex physiological processes at all levels from molecular interactions and expression to phenotypic and behavioral responses. These biological rhythms synchronize endogenous, genetically based circadian ‘clock’ or ‘pacemaker’ mechanisms with changes in environmental cues (e.g. light–dark cycles) thus enabling the organism to adapt, anticipate, and respond to changes and maintain homeostatic periodicity in body functions. Numerous physiological processes and parameters such as core body temperature, sleep–wake cycles, cardiovascular function, feeding, endocrine secretions (e.g. melatonin, cortisol, growth hormone, prolactin), hepatic metabolism, renal function, and several others exhibit rhythmicity [1, 2]. Diverse rhythmic patterns of circadian gene expression occur in virtually every cell, tissue, and organ system in the body [3, 4], and nearly half of the mammalian protein-coding genome is expressed with a circadian rhythm with tissue specificity [5]. Biorhythms in gene regulation, biochemical processes, and physiological functions bear important implications in health, disease, and pharmacotherapy [3, 6,7,8,9].
The body’s circadian clock operates even under constant environmental conditions with an approximate 24h period. Thus, it is important to recognize that sleep–wake cycles or environmental light–dark cues do not cause biological rhythms per se, but rather serve as entrainers or zeitgebers of the period and phase of the endogenous molecular clock and their downstream oscillatory processes. Both basic [10, 11] and clinical [12, 13] studies have demonstrated that disruption of circadian rhythms can be either the cause or the effect of disease including metabolic syndrome [14], inflammation [15], and cancer [16]. Furthermore, increased severity in some disease symptoms and ‘flares’ (e.g. asthma, rheumatoid arthritis) or acute events (e.g. myocardial infarction) can present at specific times of the day. These observations provide daily ‘windows of opportunity’ to effectively align treatments with disease symptoms.
For many drugs, circadian variability in their pharmacokinetics and pharmacodynamic actions are two major sources of time-varying effects. Circadian rhythms influence both the disposition and actions of various drugs and can affect therapeutic efficacy and toxicity based on dosing time. All four processes (absorption, distribution, metabolism, and elimination) influencing drug pharmacokinetics show circadian rhythms. For many drugs, significant differences in their effects can occur with different dosing times despite similar drug exposure, indicative of other pharmacodynamic mechanisms. Chronopharmacology (or chronomedicine) aims to incorporate knowledge of circadian (and/or other) biological rhythms to improve pharmacotherapy using two broad approaches: (1) direct targeting and modulation of the molecular clock (e.g. light therapy, melatonin administration), or (2) understanding and exploiting the rhythmic physiology and downstream outputs of the internal clock [17]. To maximize therapeutic benefit and/or lower adverse effects, such approaches aim to recommend an optimal time for drug administration in populations with well-synchronized circadian physiology, especially for new and existing drugs with narrow therapeutic indices. To better understand and integrate circadian physiology, aid in the analysis and interpretation of chronopharmacologic data, and guide time-based dosing, use of mathematical models and chronopharmacometric methodologies have gained importance in chronotherapy.
In the opening review of this special issue on Chronopharmacometrics, we summarize current knowledge on the biology of circadian rhythms and highlight their influence on body physiology and pharmacology. The convergence of chronopharmacology and pharmacometrics to quantify biorhythms in physiology, disease, drug biomarkers, and therapeutic effects are summarized with selected examples.
The circadian clock system
Circadian clocks comprise a hierarchical network of central and peripheral clocks which generate, sustain, and synchronize circadian rhythms. The clock system broadly comprises four major components: (1) a biochemical central clock with a period of about 24 h, (2) signaling input pathways to the central clock that enable its entrainment with the environment, (3) diverse output pathways, including hormones and the autonomic nervous system, which are linked with specific phases of the central clock, and (4) molecular clocks present in all cells of peripheral tissues that regulate rhythms in global transcriptomic expression, and thus, physiology, in a tissue-specific manner.
The suprachiasmatic nucleus (SCN)—master circadian synchronizer
In mammals, the brain plays an important role in controlling and coordinating circadian rhythms. The circadian or “master” clock resides in the suprachiasmatic nuclei (SCN), a pair of distinct neuronal clusters located in the anterior hypothalamus. The SCN is necessary for circadian function, and bilateral ablation of the SCN in rodents and pituitary tumors or vascular disease in humans has been shown to compromise circadian control of behavioral, metabolic, and endocrine rhythms [18, 19]. Transplantation of the region into a SCN-lesioned animal from another donor animal restores circadian behaviors, suggesting that the SCN region is sufficient for circadian function [19]. In addition, the SCN is capable of autonomous timekeeping: isolated neurons in primary cultures sustain circadian oscillations in gene expression and neuronal firing rates for weeks in the absence of external cues, although their rhythmic phases scatter from one another [20]. Such cell-autonomous rhythms are even more robust in organotypic SCN slices [2]. In the intact organism, the circadian clock is entrained with light/dark cues by direct retinal innervation. Specialized photosensitive retinal ganglion cells transmit photic (light/dark) input to the SCN via the retinohypothalamic tract, which entrains and synchronizes the circadian clock to the 24-h light/dark period. Some non-retinal inputs are also received by the SCN from other brain regions and external cues (e.g. feeding, temperature); these mechanisms have been reviewed elsewhere [21, 22]. Outputs from the SCN are directed to other parts of the hypothalamus and other brain regions that control both anterior and posterior pituitary hormones as well as the autonomic nervous system [23, 24]. These hormonal and autonomic outputs to a large extent convey the rhythmicity in light–dark cycles to the rest of the body.
The molecular clock – a transcriptionally-regulated oscillator
The molecular clockwork of the mammalian central pacemaker has been reviewed extensively [9, 21, 25, 26]. In vitro and ex vivo studies demonstrating cell-autonomy in clock oscillations, coupled with many genetic studies in Drosophila and mice have led to the clarification of basic molecular mechanisms underlying the circadian clock (Fig. 1). It is now established that central clock regulation involves a transcriptional, post-transcriptional, and translational auto-regulatory negative feedback loop with a period of approximately 24 h (Table 1). A pair of transcription factors encoded by the Clock (Circadian Locomotor Output Cycles Kaput) and Bmal1 (Brain-muscle Arnt like 1) genes comprise positive regulators of the feedback loop. During the early light period, CLOCK and BMAL1 heterodimerize, translocate to the nucleus, bind specific DNA elements, E-boxes, in the promoter region of target genes including Per1, Per2, and Per 3 (Period) and Cry1 and Cry2 (Cryptochrome), and stimulate their transcription. PER and CRY form the negative arm of the feedback loop. PER and CRY proteins accumulate in the cell over time (over several hours), form heterotypic complexes, and then translocate into the nucleus to repress the transcriptional activity of the CLOCK:BMAL1 complexes. During the dark period, the PER:CRY complexes degrade to low concentrations, which releases its inhibitory action on CLOCK:BMAL1 and initiates a new cycle of transcription/translation within the feedback loop. The CLOCK:BMAL1 dimers also initiate the transcription of a second feedback loop involving the Ebox-driven accessory orphan receptors REV-ERBα/β (NR1D1/2) and the RORs (retinoic acid related-orphan receptors). REV-ERBα down regulates whereas ROR up-regulates the transcription of Bmal1 by competing for ROR binding element (RRE) binding sites within the Bmal1 promoter. The casein kinase 1 epsilon and delta (CK1ε and CK1δ) also regulate PER turnover in the cytoplasm by phosphorylating PER, tagging it for ubiquitination by βTrCP and degradation by the 26S proteasome [21, 26]. This phosphorylation activity is counterbalanced by PP1 (protein phosphatase 1), producing a more gradual increase in phosphorylated PER [27]. Further, there exists a separate set of Dbox-containing clock output genes (e.g. Dbp, Hlf, Tef, and E4bp4) under the direct control of the core circulatory loop. These clock-controlled output transcription factors, along with core clock genes, control the circadian transcriptomic expression of numerous genes harboring specific regulatory motifs in the proximity of their promoters, such as E-boxes and RREs, or cAMP response elements (CREs) and D-boxes. Although the accessory genes are not required for rhythm generation [21], these interconnected feedback loops shape a robust cellular oscillator that fine-tune the phase and amplitude of core-clock gene expression.
The molecular mechanism of the circadian clock in mammals—an autoregulatory transcriptional feedback loop involving core activators, core negative feedback repressors, accessory feedback loops, additional regulatory proteins, and tissue-specific clock-controlled output genes. Clock, Circadian Locomotor Output Cycles Kaput; Bmal, Brain-muscle Arnt like 1; Per, Period; Cry, Cryptochrome; ROR, retinoic acid related-orphan receptor; CK1, casein kinase 1; PP1, protein phosphatase 1; CCG, clock-controlled genes
The peripheral clocks
Early studies document the existence of rhythmic metabolic patterns in cultured explants of the heart and adrenal gland [28]. In addition, rhythms were found to persist in isolated non-neural cultures for several cell divisions, indicating that non-SCN cells also contain endogenous circadian oscillators [29]. Following the cloning of the core clock genes, rhythms in clock mRNA and/or protein expression have been observed in various cells and tissues throughout the body [29, 30]. Transcriptomic studies have revealed that circadian regulation is highly tissue-specific [31,32,33] and more recently that nearly 50% of all protein coding genes are transcribed in a circadian manner in mice and in humans [5, 34]. Although there are thousands of genes in tissues expressed in a circadian manner, it does not mean they are all clock genes. Rather, the daily transcriptional activities of various cellular pathways and gene families, which vary greatly by cell type, are intricately coordinated by the phases of the core clock components in each cell.
Although peripheral clocks produce self-sustaining circadian oscillations in gene expression and organ function, the SCN plays an important role in coordinating rhythmic behavior in the periphery [35]. Although the core clock machinery is largely conserved within different organs [31], tissue entrainment by circadian changes of hormones such as melatonin and cortisol, rhythmic autonomic control, and indirect cues such as body temperature or feeding cycles can differentially regulate the phases and amplitudes of peripheral clock components, and consequently, patterns of the output pathways they control. Peripheral clocks play an important and unique role in each tissue, driving the circadian expression of specific genes involved in a variety of physiological functions, including xenobiotic metabolism, glucose and lipid homeostasis, cellular repair, and inflammation [30, 36]. This rhythmic variability in the periphery appears to be an essential characteristic of homeostasis, designed to optimally accommodate each tissue’s function throughout a circadian cycle.
Circadian organization: integration of central and peripheral clocks
Tissue-specific gene expression patterns are regulated by both ‘local’ as well as central mechanisms [30]. Conditional transgene experiments in mice have demonstrated that liver-specific disruption of CLOCK: BMAL1 abrogate rhythmicity in most hepatic transcripts, indicating that most circadian oscillations of hepatic function rely on an intact liver clock [37]. However, a subset of transcripts continued to cycle robustly even in the absence of a functional liver clock [37], underscoring the importance of extracellular cues. The SCN is considered to regulate peripheral tissue clocks via a combination of (1) autonomic innervation of peripheral tissues, (2) endocrine signaling, (3) temperature, and (4) behavioral signals (e.g. feeding) (Fig. 2) [3, 30]. The two primary mechanisms, autonomic nerve control and endocrine signaling, are reviewed here. Interested readers are referred to more comprehensive reviews on the roles of temperature, feeding, and other cues [21, 30, 38].
Integrative regulation of central and peripheral circadian clocks at the systemic level via (1) autonomic innervation of peripheral tissues, (2) endocrine signaling, and (3) environmental/behavioral signals. SCN suprachiasmatic nucleus, CRF corticotropin-releasing factor, GnRH gonadotropin-releasing hormone, FSH follicle stimulating hormone, LH luteinizing hormone, ACTH adrenocorticotropic hormone
Both sympathetic and parasympathetic neurons which receive inputs from the SCN and hypothalamus innervate almost all peripheral organs including the liver, muscle, adipose tissue, intestine, heart, and ovary. Photic information from the SCN regulates autonomic nerve activity with sympathetic innervations conveying light/dark signals from the SCN to the pineal gland, a small, highly vascularized, secretory neuroendocrine organ attached to the roof of the third ventricle [39]. Through synaptic transmission involving glutamate, gamma-amino butyric acid, and norepinephrine, sympathetic inputs from the SCN regulate the output of melatonin [39] (Fig. 2). Melatonin, secreted from the pineal gland, plays an important role in the regulation of sleep–wake cycles and is an established marker of the phase of the internal circadian clock [40]. Light suppresses the output of melatonin, producing peaks during the dark period and very low concentrations of melatonin during the light period. Melatonin receptors are expressed in the cells of the SCN, and melatonin plays a feedback role on the SCN [3]. It is well accepted that exogenous melatonin administration can modulate circadian rhythmicity by inducing phase shifts in clock function [1]. Melatonin membrane receptors are also present in various peripheral cells and interact with downstream messengers such as adenylyl cyclase, phospholipases, and calcium and potassium channels [41].
Like the pineal gland, autonomic pathways from the SCN directly innervate the adrenal glands (Fig. 2). By regulating epinephrine release from the adrenal medulla, sympathetic innervation via the splanchnic nerve influences a second major neuroendocrine system imparting circadian rhythmicity to peripheral tissues, the hypothalamic–pituitary–adrenal (HPA) axis [42]. Sympathoadrenomedullary release of epinephrine modulates the sensitivity of the adrenal cortex to adrenocorticotropic hormone (ACTH) influencing rhythmic glucocorticoid release. Hypothalamic paraventricular nuclei receiving photic inputs from the SCN produce corticotropin-releasing hormone (CRH) which in turn stimulates the production of ACTH from the anterior pituitary. In addition to the modulatory effect of epinephrine, it is ACTH that imparts robust circadian rhythmicity to adrenocortical glucocorticoid hormone release. Further, free circulating concentrations of glucocorticoids act in a rapid negative feedback manner to limit hypothalamic CRF and pituitary ACTH release (Fig. 2) thereby maintaining systemic homeostasis within the HPA [43]. Whereas melatonin strictly peaks in the dark period and is low during the light period independent of species, patterns in the HPA axis are essentially reversed in diurnal and nocturnal animals. In nocturnal animals such as mice and rats, corticosterone reaches peak concentrations during the transition from the light/inactive period to the dark/active period [44]. Conversely, the cortisol peak occurs during the transition from the dark/inactive period to the light/active period in diurnal animals (e.g. humans) [45]. In essence, the circadian rhythm of glucocorticoid secretion prepares the animal for the upcoming period of activity and feeding [3]. Thus, feeding patterns and sleep behavior are important aspects to consider when evaluating and comparing circadian experiments conducted in animals versus human studies.
The first definitive evidence that glucocorticoids entrained peripheral oscillators in vivo came upon the observation that administering the exogenous synthetic glucocorticoid, dexamethasone, produced transient phase shifts in circadian gene expression in peripheral tissues (liver, kidney, and heart) [46]. It is now recognized that, in virtually all tissues, glucocorticoids transcriptionally regulate the expression of both core clock genes (Bmal1, Cry1, Per1, and Per2) and a plethora of clock-controlled genes via glucocorticoid-response elements (GREs) present in the regulatory regions of these genes [47].
Akin to their influence on the HPA, both central and peripheral molecular clocks contribute to circadian rhythms within the reproductive hypothalamic-pituitary–gonadal (HPG) axis (Fig. 2). At the level of the hypothalamus, SCN neurons project directly onto gonadotropin-releasing hormone (GnRH) neurons to modulate their activity. GnRH regulates secretion of luteinizing hormone (LH) and follicle stimulating hormone (FSH) from the anterior pituitary. In females, LH and FSH stimulate ovarian follicular development and estrogen synthesis. Circulating free estrogen in turn modulates FSH and LH release through negative feedback mechanisms, completing the HPG loop (Fig. 2). The gonadotropins, FSH and LH, through regulation of the core clock genes, Per and Bmal1, regulate the ovarian circadian clock which in turn controls the timing and extent of expression of clock-controlled genes critical for normal ovarian physiology [48]. Estradiol acts in an estrogen receptor (ER) dependent manner to modulate the rhythm of PER2 in the uterus [49, 50]. Conversely, sex steroids can directly modulate the central clock by acting on ER and progesterone receptor expressing neurons in the SCN [51] (Fig. 2). The complex interactions of central and peripheral circadian clock genes, sex hormone release, and reproductive tissue sensitivity have been reviewed recently [48].
Influence on human physiology and disease
The fact that nearly half of the mammalian protein-coding genome is expressed with a circadian rhythm with tissue specificity [5], speaks by itself to the pronounced effect of circadian rhythms in influencing almost all key biochemical and physiological processes. Given the critical role played by circadian rhythm in maintaining both the systemic and tissue-level homeostasis, disruption of the rhythm has direct consequence for human health, disorders and diseases. Figure 3 summarizes the peak time for some of the key processes or biological mediators that controls physiological functions such as energy metabolism, fluid balance, inflammation, cellular turnover, and cognition/neurological responses. Core-clock genes are directly or indirectly implicated in controlling these processes, and a summary of key physiological functions affected by disrupting these core-clock genes in experimental models is described in previous publications [3, 4]. Some of the early understanding of circadian disruption on health were obtained from studies in jet lagged subjects. It is a well-known fact that changes in time zone can result in minor disruption of sleep/wake cycles. Studies have shown that frequent travels across time zones can result in not only temporary cognitive defects, but also structural brain changes such as reduction in the temporal lobe volume [12]. Multiple studies in shift workers suggests that night shifts are associated with increase in the incidence of multiple diseases such as neurological disorders, diabetes, cancer, and cardiovascular disorders [52, 53]. Just like the impact of disruption in circadian rhythms in shift workers, the ubiquitous use of light-emitting electronic devices could pose an increased risk for several circadian related disorders to a much larger segment of the population [54].

Adapted from Smolensky and Peppas (2007) and Scherholz et al. (2019). TNF-α, tumor necrosis factor-α; IFN-γ, interferon- γ, IL-6, interleukin-6; TSH, thyroid stimulating hormone; ACTH, adrenocorticotropic hormone
Diversity of circadian rhythms in human physiology. The peak time or acrophase of several biological processes in humans are shown relative to the sleep–wake cycle.
Several endocrine outputs (that control key physiological processes) such as cortisol, Adrenocorticotropic hormone (ACTH), epinephrine, testosterone, Thyroid Stimulating Hormone (TSH), Leptin, and Growth Hormone (GH) are closely regulated by circadian rhythms, and serve as a link between the central and the peripheral clocks, and their zeitgebers [43, 55, 56]. Physiological processes controlling blood flow and fluid balance like cardiac output, lymphatic flow, vascular resistance, and glomerular filtration rate follow a strong circadian fluctuation. Circadian expression of metabolic enzymes and ion channels enables the optimal metabolic and cardiac output that allows for efficient use of nutrients to match the diurnal variation in energy demand. Several lines of incidence suggest that circadian effects are likely to impact myocardial infarct incidence [57]. In the control of energy homeostasis, several of the mediators involved in regulating the energy homeostatic pathways like glycolysis, gluconeogenesis, lipid biosynthesis and metabolism, and oxidative phosphorylation are either directly or indirectly under the control of core clock genes [3, 4]. Imbalances in the timely coordination between these processes can disrupt the energy homeostasis, and could lead to disorders such as metabolic syndrome, obesity, and diabetes [53].
Decades of research in characterizing immune-regulatory and inflammatory processes suggests that these processes are intricately connected with all the key biological processes. Studies show that the core clock genes are rhythmically expressed in cells regulating immunological processes, including the functions of innate and adaptive immunity [58]. Several of the key immune cells such as T-lymphocytes, B-lymphocytes, monocytes, neutrophils, and natural killer cells show circadian rhythmicity in maturation and trafficking [59]. Furthermore, expression of several key cytokines such as TNF-α, interferon γ, IL-6, and IL-2 also show strong circadian fluctuations [59]. Multiple inflammatory and auto-immune diseases such as rheumatoid arthritis, asthma and systemic lupus show a clear circadian fluctuation in symptoms and disease severity. Furthermore, host circadian clock disruption can lead to increased pathogen replication and dissemination, which indicates that the severity of acute infections can be influenced by circadian rhythms [60]. Such complex interactions have been of particular relevance during the coronavirus disease 2019 (COVID-19) pandemic and has been reviewed [60, 61]. A recent publication summarizing an NIH workshop on circadian rhythms, sleep and immunity provides the current understanding and future opportunities in understanding the complex interactions between circadian rhythms and immune-modulatory processes, and its implications in understanding the disease pathophysiology and prevention/treatment of multiple diseases [62].
Influence on drug disposition and actions
Early experiments in mice models revealed that a drug, given at the same dose, can be lethal when administered at certain times of day or night, but had little adverse effect when administered at other times [63, 64]. Today, it is recognized that time-dependent toxicity and/or efficacy of medications can reflect predictable circadian variability in all four processes governing their pharmacokinetics (absorption, distribution, metabolism, and excretion), or drug sensitivity, or both. Further, rhythmic variations in physiology and tissue gene expression can influence drug actions, including pharmacodynamic responses, based on the availability or functioning of target proteins [3]. Time of drug dosing could play an important role in determining key pharmacokinetic parameters including overall exposure (AUC), bioavailability (F), clearance (CL), peak concentration (Cmax) and time of peak exposure (tmax), and half-life (t1/2). For many drugs, significant differences in their effects can occur with different dosing times despite similar drug exposure, indicative of other pharmacodynamic mechanisms. In this section, the effects of circadian rhythms on key determinants of drug pharmacokinetics and pharmacodynamics are summarized.
Absorption
Upon extravascular dosing, drugs must undergo passive and/or active transport from the site of administration to enter the systemic circulation. The absorption of orally administered drugs is dependent on both physiochemical properties of the compound (e.g. lipophilicity, charge/ionization, molecular weight) as well as physiological factors including gastrointestinal (GI) blood flow, pH, gastric emptying rate, and the expression and function of drug uptake and efflux transporters in the gut. Studies show that circadian rhythms are prevalent within all these physiological processes. The existence of circadian clocks within the gut, and the importance of these clocks in the timing of gut gene expression, physiology, and xenobiotic handling have been established [65,66,67].
Circadian regulation of both physiologic parameters and specific transport proteins provide a mechanistic basis for understanding observed time-of-day variability in the absorption of many drugs. Both nutrient and drug transporters in the GI tract show circadian rhythms; with many under direct core-clock control. For instance, the intestinal absorption of lipids shows a circadian rhythm, where the rate of absorption peaks during the active period of the organism [3]. Furthermore, diurnal patterns of lipid absorption and its mediators [e.g. microsomal triglyceride transfer protein (MTP)] are disrupted in Clock mutant mice [68, 69]. Consistently, circadian patterns of absorption occur for some commonly used lipophilic drugs (e.g. cyclosporine, tacrolimus, and propranolol) [70], with greater absorption occurring during the day than at night. Circadian differences in the absorption of NSAIDs have been documented in humans. Statistically significant differences in Cmax, which were higher after morning drug administration (+ 32% [diclofenac], + 52% [indomethacin], + 50% [ketoprofen]) have been shown [71]. The increased rate of absorption may be explained by daily variations in the gastric transit time and/or in intestinal blood flow [72].
Several influx and efflux transporters, including Mdr1, Mct1, Mrp2, Pept1, and Bcrp1 show circadian variation in their mRNA expression in the jejunal mucosa [73]. Recent studies by Yu et al. showed that the intestine-specific knockout of Bmal1 in mice significantly blunted the diurnal rhythm of MRP2 mRNA and protein, which in turn abrogated circadian time-dependency of MDR2-dependent methotrexate pharmacokinetics and toxicity [74]. Further, their studies revealed that Bmal1 generates diurnal MRP2 expression by temporally coordinating the expression of DBP (a MRP2 activator), REV-ERBα (an E4BP4 repressor), and E4BP4 (a MRP2 repressor) [74]. Similarly, the circadian expression of other ATP binding cassette (ABC) efflux transporters including BCRP (breast cancer resistance protein) and MDR1 (P-glycoprotein) are regulated by specific clock-controlled transcription factor(s) [75, 76]. Several chemotherapeutic agents such as methotrexate, topotecan, irinotecan and its metabolite SN-38, and doxorubicin are substrates of BCRP whereas P-glycoprotein is responsible for the efflux of small hydrophobic compounds such as digoxin, tacrolimus, and fexofenadine [77]. The proton-coupled peptide transporter, PEPT1, which actively absorbs oral β-lactam antibiotics, ester prodrugs (e.g. ACE inhibitors, protease inhibitors, valacyclovir, L-DOPA), and artificial di- and tripeptides, shows a circadian rhythm which is directly controlled by the clock-output transcription factor DBP [78]. Together, these findings indicate that the circadian molecular clock plays an important role in regulating the absorption of many drugs.
Although a few time of day dependencies in drug absorption via non-oral routes have been reported (e.g. intranasal salmon calcitonin) [79], the exact contributory mechanisms are unclear. Another relevant and emerging area in drug absorption and metabolism involves the gut microbiome [80, 81], which deserves further investigation within the context of circadian-gut microbe interactions [82, 83].
Distribution
The volume of distribution of a drug in plasma/blood and different tissues is determined largely by physiological factors such as the physical tissue volumes, blood flow rates to the organ/tissue, presence of drug transporters, red blood cell partitioning, and protein binding (binding affinity and the abundance of proteins) in plasma and tissues. There is no evidence of circadian changes in the actual tissue volumes. However, the other four factors are subject to circadian changes. Diurnal variations in both cardiac output and fractional blood flow rates to organs (e.g. brain, liver, skin, muscle) have been documented in both animals and humans [84,85,86]. Both cardiac output and organ flow rates tend to be higher during periods of activity. Therefore, for small molecule drugs with high diffusion rates, their transport out of capillaries can be limited by circadian variations in cardiac output and blood flow rates to different organs. It is well documented that various ion channels, transporters, and efflux pumps show circadian variations in tissue expression [3], which can potentially impact the transport and efflux of drug substrates in a tissue-specific manner. As factored within the Gillette equation [87], protein binding in both plasma and tissue is another major determinant of drug distribution. Indeed, albumin and other specific plasma binding proteins (e.g. transcortin and sex-hormone binding globulin) show circadian variations in systemic circulation, which can alter the plasma free fraction of highly bound drugs, and consequently impact their distribution. Transcortin, which shows low capacity, high binding for selected glucocorticoids, produces a circadian rhythm in binding to synthetic corticosteroids (e.g. prednisolone, hydrocortisone) and natural hormone (e.g. cortisol in humans) [88, 89]. The lowest binding capacity occurs during the night, around 4 AM, and the highest around 5 PM [90]. However, the clinical consequences of such variation in drug binding on their actions have not been demonstrated yet. A recent population-based PK analysis by Melin et al. [91] showed that a 23% diurnal variation in plasma transcortin from baseline, as observed in healthy volunteers [90], produced modest differences in the median total AUC (≤ 12.2%) and free fractions at higher concentrations (10.3% vs 11.7%) of hydrocortisone between times of highest and lowest transcortin concentrations. It was concluded that circadian variations in transcortin are unlikely of clinical consequence.
Drugs that bind to their pharmacological targets with high affinity and to a significant extent (relative to dose) can exhibit target-mediated drug disposition (TMDD), where the drug-target interaction can be reflected in their dose-dependent PK profiles [92]. Many such receptors and drug targets can exhibit circadian rhythms at their sites of expression in tissues and cells. For instance, Volz et al. recently developed a pharmacokinetic TMDD model, which corroborated the hypothesis that the nonlinear PK behavior of bosentan, a small molecule endothelin receptor antagonist, is mediated by its high receptor binding affinity coupled with time-varying (diurnal) receptor synthesis [93].
Metabolism
The optimization of metabolism and energy expenditure are key tasks of the circadian timing system. Importantly, fasting-feeding cycles accompanying rest-activity rhythms are major timing cues that synchronize rhythmic processes in the body. Within this context, the evolution of a time-dependent mechanism for the detoxification of noxious components ingested during food consumption is particularly relevant in mammals. Thus, it can be reconciled that the same underlying xenobiotic detoxification system plays an important role in the timing of drug metabolism, manifesting itself in circadian pharmacokinetics and pharmacodynamics, hence producing circadian changes in drug efficacy and toxicity [63].
The liver is the body’s major organ that is involved in the biotransformation of drugs. In accordance with the well-stirred model of hepatic clearance, factors affecting the metabolism of a drug in liver include the hepatic blood flow rate, specific enzymatic activity (intrinsic clearance), and plasma protein binding (as only the free unbound drug can be metabolized) [94]. Both the blood flow rate to the liver and plasma protein binding show circadian rhythms [86]. It is also well established that several enzymes involved drug metabolism also show circadian rhythms in gene expression and activity [3]. There is strong evidence that the transcription of many such enzymes is, at least in part, regulated by core clock genes. Although circadian variations are observed in nearly all factors controlling hepatic metabolism, the relative contribution to the circadian metabolism of a drug can vary. For a high extraction drug (intrinsic clearance > > hepatic blood flow), the circadian variation in the hepatic blood flow rate is the major determinant of hepatic clearance. For example, nicotine metabolism occurs rapidly and extensively in liver; thus, its clearance is likely dependent on hepatic blood flow. As such, nicotine clearance displays an average daily variation of 17% (from peak to through) [95]. In contrast, for a low extraction drug (intrinsic clearance < < hepatic blood flow) the circadian variation in both protein binding and enzymatic activity will be the major factors that determine the circadian rhythmicity in their metabolism. This phenomenon is commonly documented upon monitoring blood drug concentrations during a continuous infusion. For example, significant circadian variations occurred in the plasma concentrations of 5-fluorouracil (5-FU) in patients administered a continuous infusion at 300 mg/m2/day [96]. In peripheral blood mononuclear cells obtained the same patients, circadian variations in the activity of dihydropyrimidine dehydrogenase, the major catabolizing enzyme of 5-FU were noted. This rhythm shared an inverse relationship with the variation in plasma 5-FU [96]. Circadian variation in the hepatic elimination rate of 5-FU in isolated perfused rat liver has also been reported [97].
Numerous cytochrome P450 genes in liver, including Cyp2b10, Cyp2e1, Cyp4a14, Cyp2a4, Cypa5, Cyp2c22, Cyp2e1, Cyp3a, Cyp4a3, Cyp7, and Cyp17 show circadian rhythmicity in expression [3, 98]. Enzymes involved in phase two metabolism including glutathione S-transferase, carboxylesterase, cysteine dioxygenase, UDP glucuronosyltransferase, and sulfotransferase are also expressed in a circadian manner [98]. The diverse roles, drug substrates, and site(s) of expression of such circadian-driven metabolizing enzymes have been the subject of recent and extensive review [99]. Rapid advances in molecular biology techniques [e.g., molecular cloning, gene editing, and chromatin immunoprecipitation (ChIP) analysis] have revealed both direct and indirect mechanisms of core clock gene regulation of circadian drug metabolizing enzyme transcription. Bmal1 and Clock generate circadian rhythmicity in Cyp2a4/5, Ugt1a1, Fmo5, and Sult1a1 expression by direct transcriptional activation upon binding to E-box elements in the promoter regions of these target genes [100, 101]. In addition, Bmal1/Clock also regulate expression of drug-metabolizing enzymes through indirect mechanisms. For example, Bmal1 regulates diurnal expression of Cyp3a11 through DBP and HNF4α, two direct targets of Bmal1 and activators of Cyp3a11 [102]. Lu and colleagues have recently published an excellent review on this topic and summarize mechanistic studies that provide a strong link between molecular clock output genes and hepatic drug metabolism [100].
Excretion
The excretion of drug either unchanged or as a metabolite is the last step in its disposition. For most drugs, this process is primarily mediated by the kidney whereas biliary excretion and subsequent fecal elimination occur with some drugs. The circadian presence of efflux pumps in the bile canalicular membranes may influence the excretion of certain types of drugs from the hepatocytes into bile. Circadian rhythms have been documented in all three contributory processes of kidney excretion—glomerular filtration, tubular secretion, and tubular reabsorption [103]. Studies evaluating daily glomerular filtration rate (GFR) using inulin as a marker suggest that GFR shows a circadian variation which peaks during the daytime in humans and is lower at night [104]. For drugs which are primarily eliminated through glomerular filtration and show very low protein binding, their circadian variation in renal clearance would depend primarily on daily GFR fluctuation. For example, the circadian variation of renal excretion of amikacin, an aminoglycoside antibiotic, shows a pattern like that of GFR [3]. Consistently, the antibiotic gentamycin displayed the best renal tolerability following afternoon dosing instead of dosing during the rest period (midnight to 7:30 AM) in patients with severe infections [105]. However, for highly bound drugs, circadian variation in plasma protein binding can also affect circadian changes in renal excretion.
Similar to other tissues, the kidney also possesses an intrinsic circadian timing system with clock-controlled tissue-specific output genes [106]. Many output genes are involved in the control of sodium ions, water balance, and nutrient/xenobiotic transport. Specific transporters in the kidney proximal tubule including P-glycoprotein, various organic acid and base transporters, and proteins of the solute carrier families, which are involved in the tubular secretion of drugs display circadian rhythms [107]. Ampicillin, a drug primarily excreted by tubular secretion shows circadian variation in clearance, which indicates a possible circadian rhythm in the tubular secretion process [108]. Of note, nephron-specific Bmal1 knockout mice showed an 80% reduction in organic anion transporter 3 (OAT3) expression with an impaired natriuretic response to furosemide [109]. These findings point to a central role of core circadian clock function in renal physiology and pharmacology. Numerous drugs and essential solutes are reabsorbed by either an active process or simple passive diffusion. There is no documented evidence for circadian variation in active reabsorption thus far. However, the passive reabsorption of a drug depends upon its lipophilicity and pKa as well as the urinary pH and flow rate. Both the urinary pH and urinary flow rate display diurnal rhythms, with urinary pH being more acidic during the inactive period and urinary flow rate peaking during the active period [63].
Drug actions (chronopharmacodynamics)
Chronopharmacology studies have revealed significant differences in drug effects with different timing of dosing, despite showing the same concentration–time profiles in plasma and/or at their site(s) of action. Rhythmic changes in free-to-bound drug, target or receptor availability, downstream mediators, and activity levels of rate-limiting steps in pharmacologic signaling pathways in drug-targeted tissues are some factors which can produce pharmacodynamic effects of varying intensities based on dosing time. This phenomenon is particularly well-documented for diverse anticancer agents. The antitumor effect of interferon- β (IFN-β; 0.5 MI.U./kg, intratumoral) was more efficient upon 9:00 AM dosing versus 9:00 PM dosing in melanoma bearing mice [110]. Mechanistically, this dosing time-dependent IFN-β antitumor effect was attributed to a change in drug sensitivity with increased IFN receptor synthesis and specific receptor binding by IFN-β on tumor cells in S phase [110]. Furthermore, IFN-β concentrations were increased in tumor, presumably due to increased IFN receptor binding [111], upon morning dosing [110]. An antiangiogenic agent, SU1498 (VEGFR-2 tyrosine kinase inhibitor), showed increased antitumor efficacy when administered during the early light phase compared with the early dark phase [112]. This differential effect based on dosing time was related to the robust circadian variation in vascular endothelial growth factor (VEGF) production, which occurs under direct regulation of the molecular clock [112]. A similar mechanism involving the circadian availability of platelet-derived growth factor (PDGF) has been shown to explain the chrono-efficacy (antitumor effect) of imatinib mesylate (Gleevec) [113]. Mechanisms contributing to time-dependent efficacy can involve feedback from secondary signaling proteins, hormones, or regulatory networks. Epidermal growth factor receptor (EGFR) signaling, an essential pathway for cell growth and cell–cell interactions, is suppressed by high glucocorticoid concentrations that occur during the active phase (night in rodents), while EGFR signals are enhanced during the resting phase [114]. Consistent with this pattern, treatment of animals bearing EGFR-driven tumors with lapatinib (40 mg/kg per day orally), an inhibitor of ErbB1 and ErbB2 tyrosine kinases, was more effective when administered during the resting phase of the day, when glucocorticoids are low [114]. Collectively, such findings support a circadian clock-based paradigm in cancer therapy.
Chronoefficacy is also relevant to drugs administered for other indications, including neuropathic pain, cardiovascular disease, and metabolic disease. Using a mouse model of neuropathic pain, Kusunose et al. elucidated the molecular basis for the dosing time-dependency of anti-allodynic effects of gabapentin [115]. Specifically, the authors demonstrated that circadian oscillations in the protein abundance of the calcium channel α2δ-1 subunit, a target of gabapentin, correlated with fluctuations in the maximal binding capacity of gabapentin in the dorsal root ganglion; and that the anti-allodynic effect of gabapentin was attenuated at the times of the day when α2δ-1 subunit protein was abundant [115]. Serum gabapentin concentrations did not significantly differ between the two dosing times tested (5:00 AM vs. 5:00 PM) [115]. The anticoagulant effect of rivaroxaban, a direct inhibitor of activated factor X (FX), is influenced by the dosing time [116]. In rats, FX activity follows a 24-h rhythm with a peak in the middle of the light phase and a trough at the beginning of the dark phase. Consistently, a single dose of rivaroxaban (1 mg/kg or 10 mg/kg) inhibited FX activity more strongly after dosing at the beginning of the light phase than after dosing than in the dark phase, with minimal influence of dosing times on rivaroxaban pharmacokinetics [116]. However, additional differences in rivaroxaban pharmacokinetics related to dosing time cannot be ruled out. A human trial demonstrated > twofold drug concentrations after intake and therefore more pronounced FX inhibition and suppression of in vivo thrombin formation in the first 12 h after administration in the evening as compared to morning intake [117].
Unsurprisingly, the core proteins of the circadian clock, such as Per, Rev-erbα/β, Rorα/γ, and Cry have and continue to be targeted by investigational small molecule drugs for the treatment of certain diseases. The advances, challenges, and opportunities for ‘drugging the clock’ to manage and treat diverse disorders have been discussed elsewhere [65, 118, 119].
Chronopharmacometrics and systems chronotherapeutics
The key theme of this issue of the Journal of Pharmacokinetics & Pharmacodynamics – Chronopharmacometrics – encompasses pharmacometric methods relevant to quantifying chronopharmacology and chronotherapeutics. Over the past three decades, multiple chronopharmacometric methodologies have evolved within the broader context of general pharmacokinetic-pharmacodynamic modeling. Spanning the spectrum of empirical, pharmaco-statistical methods of analysis to more complex mechanistic and systems modeling approaches, the general goals of modeling circadian phenomena are to (1) better understand circadian systems biology and mechanism(s) of drug action, (2) aid in the appropriate quantitative analysis and interpretation of chronopharmacologic time course data, and (3) guide time-based dosing using modeling and simulation.
Diverse mathematical or statistical models can be incorporated within established pharmacometric models to characterize circadian rhythmicity in experimental (placebo plus drug treatment) data. To jointly model circadian rhythms in endogenous cortisol in placebo-treated volunteers along with the time course endogenous cortisol suppression by administration of inhaled fluticasone propionate, Chakraborty et al. implemented indirect response models (IDR) using various biorhythmic functions (single cosine, dual ramps, dual zero-order, dual cosines, and Fourier-based L2-Norm methods) for the production rate [120]. It was shown that the Fourier series with L2-Norm best captured the placebo and drug treatment data, offering a flexible and accurate method to capture periodic rhythms. However, appropriate selection of a suitable rhythmic function or model will be case-specific and highly dependent on the experimental data available. A general methodological review on procedures for numerical analysis of circadian rhythms—from visual inspection of time plots and actograms to several mathematical methods of time series modeling—has been published [121].
To delineate circadian effects and interactions impacting both the pharmacokinetics and pharmacodynamics (cortisol suppression and lymphocyte trafficking) of total plus free prednisolone and prednisone, Derendorf and colleagues employed IDR models to comprehensively capture the joint effects of free prednisolone and cortisol on inhibition of the production rate for lymphocyte suppression, as a function of dosing time [122]. Moving beyond these efforts, studies from the Jusko laboratory have evolved more mechanistic systems pharmacokinetic-pharmacodynamic models incorporating circadian rhythmicity in hormonal concentrations, tissue mRNA biomarker expression, and protein concentrations to fully decipher the physiologic and pharmacologic control of endogenous and exogenous corticosteroid responses in a quantitative manner [123,124,125,126]. The integrated effects of circadian rhythms, methylprednisolone kinetics, and the glucose/free fatty acids/insulin system on the regulation of two important adipokines, adiponectin and leptin, were evaluated using systems modeling [125]. The model simultaneously captured the complex response dynamics of adiponectin and leptin mRNA and protein concentrations as well as plasma glucose, insulin, and free fatty acid concentrations, under homeostatic (circadian) and perturbed (steroid-dosed) conditions. Another study [127] experimentally examined the concerted, multi-tissue regulation of the glucocorticoid induced leucine zipper (GILZ), a novel and sensitive anti-inflammatory biomarker of corticosteroids, in rat lung, skeletal muscle, and fat in vivo. By incorporating Fourier-based harmonics to describe rhythms in both GILZ and glucocorticoid receptor dynamics within the fifth-generation model framework for corticosteroids [128], the model captured the tissue-specific dynamics of GILZ mRNA under baseline (circadian) and steroid-treated conditions in all tissues examined [123]. Scherholz et al. provide a sweeping overview of recent advances in corticosteroid therapy that have further improved patient safety and efficacy, with an emphasis on chronopharmacology [129].
Recently, the IDR paradigm with circadian input rates was extended by developing indirect pharmacodynamic models with periodic removal of the response variable [130]. The developed model captured applicable pharmacodynamic biomarkers such as plasma uric acid, brain amyloid β, and dopamine; providing a mechanistic basis to model drug responses displaying nonstationary baselines controlled by removal mechanisms [130]. In addition, Koch and Schropp have proposed IDR-based models that include a delay process to realize oscillating response behaviors [131]. In this type of model, production rate is first-order whereas loss is replaced by a second-order term with delayed control by the response variable. Interpretations posited for the delay include lifespan and maturation time [131].
In complement and in parallel to advancements in chronopharmacometrics, the field of systems biology has offered considerable and longstanding contributions to better understand circadian phenomena. In general, systems biology comprises a broad spectrum of computational methods to understand physiology and disease at levels of molecular pathways, regulatory networks, cells, tissues, organs, and, ultimately, the whole organism [132]. Long before molecular genetics led to the identification of the transcriptional and translational feedback loops governing the core clock that underlies the mechanism of circadian oscillation in organisms, theoretical assessments, dating as early as 1965 [133], have sought to understand how circadian properties of oscillations, periodicity, and entrainment emerge. For example, systems modelers have shown, based on a hypothetical biochemical network with negative feedback, there are necessary constraints on reaction rates for the generation of instability at steady state [134, 135]. Assuming this structural model and parameter constraints, it was shown that the rate of transcription and translation are not vital for setting period length, but instead a critical feature is the degradation rate of the repressor mediating feedback [134, 136]. These studies highlight fundamental contributions of systems modeling, even prior to any knowledge of the molecular network underpinning circadian rhythms.
Subsequent identification of the molecular components of circadian rhythms have led to a large array of more elaborate models incorporating intricate gene loops of core clock regulation, including their mRNA and proteins, downstream functions, and clock-controlled genes. This expansion is also largely attributed to advances in transcriptomics and proteomics methods, which have enabled systems biology-derived discoveries of intrinsic clock phenomena (reviewed extensively in [134]). Circadian ‘signatures’ for numerous genes derived from such large ‘omics’ datasets have also been incorporated with mathematical models of circadian rhythms. For example, Mavroudis et al. utilized a mathematical model in conjunction with large-scale, multi-tissue microarray data to explore the dynamics of core-clock and clock-controlled genes measured in four tissues of the rat (liver, muscle, adipose, and lung). Their analysis revealed inter-tissue circadian variability among the same clock-controlled genes, attributable to differing tissue-specific rates of transcription factor activities on Ebox/RRE/Dbox elements [31, 137]. Efforts in quantitative systems modeling of chronotherapy within the context of molecular-to-whole body integration have expanded over the past two decades. The Androulakis lab have assembled complex models of systems biology for understanding circadian rhythms within various relevant contexts such as inflammation [138], circadian-immune interactions [139], and seasonal impacts on synchronization of the circadian clock and cell cycle [140].
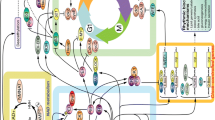
There is growing interest in the complementary adoption of PK/PD and systems biology approaches toward the development of integrative quantitative systems pharmacology (QSP) models to understand more fully the complex interactions of drugs and disease biology and consequently improve drug development and pharmacotherapy [141]. As a multidisciplinary science, QSP promises a framework for integrating information obtained from studying biologic (normal and aberrant) pathways and pharmacological targets to predict clinical efficacy and adverse events through iterations between mathematical modeling and experimentation. As conceptualized in Fig. 4, there remains considerable opportunity to further integrate key aspects of physiology and disease, circadian biology, and therapeutic interventions (i.e. drug disposition and pharmacology) towards the development of circadian systems pharmacology models. Such models should seek to integrate the time-course of drug pharmacokinetics, target engagement and downstream effects (pharmacodynamics), interactions of the molecular components of the circadian clock and/or other relevant circadian physiology that manifest as time-varying drug concentrations or responses, and last, the processes reflecting disease and its perturbation on normal body function. Relevant information to inform and integrate each component of the systems model may rely on existing experimental data, published mathematical models, and prospective fit-for-purpose experiments designed to calibrate or qualify the model (Fig. 4). To quantitatively predict circadian modulation by the light–dark cycle as well as the by the CK1δ/ε inhibitor, PF-670462, Kim et al. developed a systems pharmacology model, which successfully validated experimental data indicating that chronic CK1δ/ε inhibition during the earlier hours of the light–dark cycle can produce a constant stable delay of rhythm whereas chronic dosing later during the day, or in the presence of longer light intervals, would not yield an entrained rhythm [142]. Ballesta et al. extensively review recent advances in multiscale systems chronopharmacology approaches toward the design of patient-tailored chronotherapies, with an with emphasis on both cancer management and circadian timing system–resetting strategies [7]. In their review, the development of multiscale systems chronopharmacology models, which integrate data from in vitro cells systems to in vivo animal models, are described for chemotherapeutic agents such as irinotecan, oxaliplatin, and 5-flurouracil [7]. For other drugs and hormones with complex multifactorial effects, such as corticosteroids, there remains opportunities and challenges to further evolve more global and mechanistic models that jointly consider (1) the circadian physiology of endogenous glucocorticoid hormones, including HPA feedback and adrenal suppression [120], (2) possible PK complexities arising from circadian time-varying transcortin concentrations [91], (3) tissue-specific peripheral clock gene regulation [137], (4) effects of steroid on the peripheral clocks and circadian glucocorticoid receptor concentrations in tissues [46], and (5) the tissue-specific receptor-mediated genomic actions of corticosteroids [123, 124]. It is anticipated that continued evolution of such systems models could eventually lead to a highly predictive framework, reducing need for empiricism (e.g., use of data-driven cosine fitting) within current pharmacokinetic-pharmacodynamic models.
Schematic representing a systems pharmacology approach in integrating circadian biology, physiology and PK/PD. Conceptualization of relevant information use and integration of knowledge is shown using broken arrows. See Figs. 1 and 2 for full details contained within the boxes shown on the top-right (‘Circadian Biology’) and top-left (‘Physiology and Disease’)
Concluding perspectives
Circadian rhythms and the internal clock are elegant products of evolution perfected over millions of years to aid in the adaptation of the living beings to periodic fluctuations in the external world. Owing to several decades (if not centuries) of research into understanding the molecular, systemic, and behavioral components of circadian rhythms, and the intricate interactions between the central and the peripheral clocks and the external cues that entrains them, we now have a robust understanding on how circadian functioning maintains health, or when disrupted, can lead to disease. There has been some success in translating this understanding into identifying pharmacological targets and molecular therapeutics. However, owing to the complexity of how the internal clock mechanisms work, and the extensive crosstalk and feedback loops that maintain homeostasis, a large portion of the internal clock mechanisms are still undruggable and under-exploited. Recent advances made in applying systems biology and systems pharmacology approaches can help in gaining a more in-depth understanding of the circadian circuitry and its down-stream effects, which could translate into identifying molecular targets for the treatment of different diseases. In addition, this in-depth understanding is important for making relevant life-style interventions that modulate the external cues capable of regulating the internal clock, and hence help prevent or manage circadian disruption related disorders.
References
Skene DJ, Arendt J (2006) Human circadian rhythms: physiological and therapeutic relevance of light and melatonin. Ann Clin Biochem 43(Pt 5):344–353. https://doi.org/10.1258/000456306778520142
Maywood ES, O’Neill JS, Chesham JE, Hastings MH (2007) Minireview: the circadian clockwork of the suprachiasmatic nuclei—analysis of a cellular oscillator that drives endocrine rhythms. Endocrinology 148(12):5624–5634. https://doi.org/10.1210/en.2007-0660
Sukumaran S, Almon RR, DuBois DC, Jusko WJ (2010) Circadian rhythms in gene expression: Relationship to physiology, disease, drug disposition and drug action. Adv Drug Deliv Rev 62(9–10):904–917. https://doi.org/10.1016/j.addr.2010.05.009
Wang H, Yang Z, Li X, Huang D, Yu S, He J, Li Y, Yan J (2020) Single-cell in vivo imaging of cellular circadian oscillators in zebrafish. PLoS Biol 18(3):e3000435. https://doi.org/10.1371/journal.pbio.3000435
Zhang R, Lahens NF, Ballance HI, Hughes ME, Hogenesch JB (2014) A circadian gene expression atlas in mammals: implications for biology and medicine. Proc Natl Acad Sci U S A 111(45):16219–16224. https://doi.org/10.1073/pnas.1408886111
Adam D (2019) Core Concept: emerging science of chronotherapy offers big opportunities to optimize drug delivery. Proc Natl Acad Sci U S A 116(44):21957–21959. https://doi.org/10.1073/pnas.1916118116
Ballesta A, Innominato PF, Dallmann R, Rand DA, Lévi FA (2017) Systems chronotherapeutics. Pharmacol Rev 69(2):161. https://doi.org/10.1124/pr.116.013441
Brainard J, Gobel M, Scott B, Koeppen M, Eckle T (2015) Health implications of disrupted circadian rhythms and the potential for daylight as therapy. Anesthesiology 122(5):1170–1175. https://doi.org/10.1097/ALN.0000000000000596
Patke A, Young MW, Axelrod S (2020) Molecular mechanisms and physiological importance of circadian rhythms. Nat Rev Mol Cell Biol 21(2):67–84. https://doi.org/10.1038/s41580-019-0179-2
Quagliarini F, Mir A, Balazs K, Wierer M, Dyar K, Jouffe C, Makris K, Hawe J, Heinig M, Filipp F, Barish G, Uhlenhaut H (2019) Cistromic Reprogramming of the diurnal glucocorticoid hormone response by high-fat diet. Mol Cell. https://doi.org/10.1016/j.molcel.2019.10.007
Hadadi E, Taylor W, Li X-M, Aslan Y, Villote M, Rivière J, Duvallet G, Auriau C, Dulong S, Raymond-Letron I, Provot S, Bennaceur-Griscelli A, Acloque H (2020) Chronic circadian disruption modulates breast cancer stemness and immune microenvironment to drive metastasis in mice. Nat Commun 11(1):3193. https://doi.org/10.1038/s41467-020-16890-6
Cho K (2001) Chronic “jet lag” produces temporal lobe atrophy and spatial cognitive deficits. Nat Neurosci 4(6):567–568. https://doi.org/10.1038/88384
Biggi N, Consonni D, Galluzzo V, Sogliani M, Costa G (2008) Metabolic syndrome in permanent night workers. Chronobiol Int 25(2):443–454. https://doi.org/10.1080/07420520802114193
Maury E, Ramsey KM, Bass J (2010) Circadian rhythms and metabolic syndrome: from experimental genetics to human disease. Circ Res 106(3):447–462. https://doi.org/10.1161/CIRCRESAHA.109.208355
Castanon-Cervantes O, Wu M, Ehlen JC, Paul K, Gamble KL, Johnson RL, Besing RC, Menaker M, Gewirtz AT, Davidson AJ (2010) Dysregulation of inflammatory responses by chronic circadian disruption. J Immunol 185(10):5796. https://doi.org/10.4049/jimmunol.1001026
Masri S, Sassone-Corsi P (2018) The emerging link between cancer, metabolism, and circadian rhythms. Nat Med 24(12):1795–1803. https://doi.org/10.1038/s41591-018-0271-8
Ruben MD, Smith DF, FitzGerald GA, Hogenesch JB (2019) Dosing time matters. Science 365(6453):547–549. https://doi.org/10.1126/science.aax7621
Weaver DR (1998) The suprachiasmatic nucleus: a 25-year retrospective. J Biol Rhythms 13(2):100–112. https://doi.org/10.1177/074873098128999952
Herzog ED, Hermanstyne T, Smyllie NJ, Hastings MH (2017) Regulating the suprachiasmatic nucleus (SCN) Circadian Clockwork: Interplay between cell-autonomous and circuit-level mechanisms. Cold Spring Harb Perspect Biol 9(1):a027706. https://doi.org/10.1101/cshperspect.a027706
Webb AB, Angelo N, Huettner JE, Herzog ED (2009) Intrinsic, nondeterministic circadian rhythm generation in identified mammalian neurons. Proc Natl Acad Sci U S A 106(38):16493–16498. https://doi.org/10.1073/pnas.0902768106
Buhr ED, Takahashi JS (2013) Molecular components of the mammalian circadian clock. Handb Exp Pharmacol 217:3–27. https://doi.org/10.1007/978-3-642-25950-0_1
Hastings MH, Maywood ES, Brancaccio M (2018) Generation of circadian rhythms in the suprachiasmatic nucleus. Nat Rev Neurosci 19(8):453–469. https://doi.org/10.1038/s41583-018-0026-z
Buijs RM, Scheer FA, Kreier F, Yi C, Bos N, Goncharuk VD, Kalsbeek A (2006) Organization of circadian functions: interaction with the body. Prog Brain Res 153:341–360. https://doi.org/10.1016/s0079-6123(06)53020-1
Kalsbeek A, Palm IF, La Fleur SE, Scheer FAJL, Perreau-Lenz S, Ruiter M, Kreier F, Cailotto C, Buijs RM (2006) SCN outputs and the hypothalamic balance of life. J Biol Rhythms 21(6):458–469. https://doi.org/10.1177/0748730406293854
Chang DC, Reppert S (2001) The circadian clocks of mice and men. Neuron 29:555–558
Ko CH, Takahashi JS (2006) Molecular components of the mammalian circadian clock. Human Mol Genet 15(2):R271-277. https://doi.org/10.1093/hmg/ddl207
Lee HM, Chen R, Kim H, Etchegaray JP, Weaver DR, Lee C (2011) The period of the circadian oscillator is primarily determined by the balance between casein kinase 1 and protein phosphatase 1. Proc Natl Acad Sci U S A 108(39):16451–16456. https://doi.org/10.1073/pnas.1107178108
Andrews RV, Edgar Folk G (1964) Circadian metabolic patterns in cultured hamster adrenal glands. Comp Biochem Physiol 11(4):393–409. https://doi.org/10.1016/0010-406X(64)90006-4
Balsalobre A, Damiola F, Schibler U (1998) A serum shock induces circadian gene expression in mammalian tissue culture cells. Cell 93(6):929–937. https://doi.org/10.1016/S0092-8674(00)81199-X
Mohawk JA, Green CB, Takahashi JS (2012) Central and peripheral circadian clocks in mammals. Annu Rev Neurosci 35:445–462. https://doi.org/10.1146/annurev-neuro-060909-153128
Mavroudis PD, DuBois DC, Almon RR, Jusko WJ (2018) Daily variation of gene expression in diverse rat tissues. PLoS ONE 13(5):e0197258. https://doi.org/10.1371/journal.pone.0197258
Panda S, Antoch MP, Miller BH, Su AI, Schook AB, Straume M, Schultz PG, Kay SA, Takahashi JS, Hogenesch JB (2002) Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 109(3):307–320. https://doi.org/10.1016/s0092-8674(02)00722-5
Storch K-F, Lipan O, Leykin I, Viswanathan N, Davis FC, Wong WH, Weitz CJ (2002) Extensive and divergent circadian gene expression in liver and heart. Nature 417(6884):78–83. https://doi.org/10.1038/nature744
Ruben MD, Wu G, Smith DF, Schmidt RE, Francey LJ, Lee YY, Anafi RC, Hogenesch JB (2018) A database of tissue-specific rhythmically expressed human genes has potential applications in circadian medicine. Sci Translat Med 10(458):eaat8806. https://doi.org/10.1126/scitranslmed.aat8806
Akhtar RA, Reddy AB, Maywood ES, Clayton JD, King VM, Smith AG, Gant TW, Hastings MH, Kyriacou CP (2002) Circadian cycling of the mouse liver transcriptome, as revealed by cDNA microarray, is driven by the suprachiasmatic nucleus. Curr Biol 12(7):540–550. https://doi.org/10.1016/s0960-9822(02)00759-5
Lamia KA, Storch K-F, Weitz CJ (2008) Physiological significance of a peripheral tissue circadian clock. Proc Natl Acad Sci 105(39):15172. https://doi.org/10.1073/pnas.0806717105
Kornmann B, Schaad O, Bujard H, Takahashi JS, Schibler U (2007) System-driven and oscillator-dependent circadian transcription in mice with a conditionally active liver clock. PLoS Biol 5(2):e34. https://doi.org/10.1371/journal.pbio.0050034
Buijs RM, Kalsbeek A (2001) Hypothalamic integration of central and peripheral clocks. Nat Rev Neurosci 2(7):521–526. https://doi.org/10.1038/35081582
Aulinas A (Last Updated Dec 10 2019) Physiology of the Pineal Gland and Melatonin. Endotext.org. https://www.ncbi.nlm.nih.gov/books/NBK550972/?report=classic.
Lewy AJ, Cutler NL, Sack RL (1999) The endogenous melatonin profile as a marker for circadian phase position. J Biol Rhythms 14(3):227–236. https://doi.org/10.1177/074873099129000641
Cipolla-Neto J, Amaral FGd (2018) Melatonin as a hormone: new physiological and clinical insights. Endocr Rev 39(6):990–1028. https://doi.org/10.1210/er.2018-00084
Kalsbeek A, van der Spek R, Lei J, Endert E, Buijs RM, Fliers E (2012) Circadian rhythms in the hypothalamo-pituitary-adrenal (HPA) axis. Mol Cell Endocrinol 349(1):20–29. https://doi.org/10.1016/j.mce.2011.06.042
Evanson NK, Tasker JG, Hill MN, Hillard CJ, Herman JP (2010) Fast feedback inhibition of the hpa axis by glucocorticoids is mediated by endocannabinoid signaling. Endocrinology 151(10):4811–4819. https://doi.org/10.1210/en.2010-0285
Yao Z, DuBois DC, Almon RR, Jusko WJ (2006) Modeling circadian rhythms of glucocorticoid receptor and glutamine synthetase expression in rat skeletal muscle. Pharm Res 23(4):670–679. https://doi.org/10.1007/s11095-005-9608-3
Challet E (2007) Minireview: entrainment of the suprachiasmatic clockwork in diurnal and nocturnal mammals. Endocrinology 148(12):5648–5655. https://doi.org/10.1210/en.2007-0804
Balsalobre A, Brown SA, Marcacci L, Tronche F, Kellendonk C, Reichardt HM, Schütz G, Schibler U (2000) Resetting of circadian time in peripheral tissues by glucocorticoid signaling. Science 289(5488):2344–2347. https://doi.org/10.1126/science.289.5488.2344
Dickmeis T (2009) Glucocorticoids and the circadian clock. J Endocrinol 200(1):3–22. https://doi.org/10.1677/joe-08-0415
Sen A, Hoffmann HM (2020) Role of core circadian clock genes in hormone release and target tissue sensitivity in the reproductive axis. Mol Cell Endocrinol 501:110655. https://doi.org/10.1016/j.mce.2019.110655
Gery S, Virk RK, Chumakov K, Yu A, Koeffler HP (2007) The clock gene Per2 links the circadian system to the estrogen receptor. Oncogene 26(57):7916–7920. https://doi.org/10.1038/sj.onc.1210585
Nakamura TJ, Sellix MT, Menaker M, Block GD (2008) Estrogen directly modulates circadian rhythms of PER2 expression in the uterus. Am J Physiol Endocrinol Metab 295(5):E1025-1031. https://doi.org/10.1152/ajpendo.90392.2008
Kruijver FP, Swaab DF (2002) Sex hormone receptors are present in the human suprachiasmatic nucleus. Neuroendocrinology 75(5):296–305. https://doi.org/10.1159/000057339
Woo JM, Postolache TT (2008) The impact of work environment on mood disorders and suicide: evidence and implications. Int J Disabil Hum Dev 7(2):185–200. https://doi.org/10.1515/ijdhd.2008.7.2.185
Esquirol Y, Bongard V, Mabile L, Jonnier B, Soulat JM, Perret B (2009) Shift work and metabolic syndrome: respective impacts of job strain, physical activity, and dietary rhythms. Chronobiol Int 26(3):544–559. https://doi.org/10.1080/07420520902821176
Chang AM, Aeschbach D, Duffy JF, Czeisler CA (2015) Evening use of light-emitting eReaders negatively affects sleep, circadian timing, and next-morning alertness. Proc Natl Acad Sci U S A 112(4):1232–1237. https://doi.org/10.1073/pnas.1418490112
Wittert G (2014) The relationship between sleep disorders and testosterone in men. Asian J Androl 16(2):262–265. https://doi.org/10.4103/1008-682X.122586
Brandenberger G, Weibel L (2004) The 24-h growth hormone rhythm in men: sleep and circadian influences questioned. J Sleep Res 13(3):251–255. https://doi.org/10.1111/j.1365-2869.2004.00415.x
Virag JA, Lust RM (2014) Circadian influences on myocardial infarction. Front Physiol 5:422. https://doi.org/10.3389/fphys.2014.00422
Boivin DB, James FO, Wu A, Cho-Park PF, Xiong H, Sun ZS (2003) Circadian clock genes oscillate in human peripheral blood mononuclear cells. Blood 102(12):4143–4145. https://doi.org/10.1182/blood-2003-03-0779
Cuesta M, Boudreau P, Dubeau-Laramée G, Cermakian N, Boivin DB (2016) Simulated night shift disrupts circadian rhythms of immune functions in humans. J Immunol 196(6):2466–2475. https://doi.org/10.4049/jimmunol.1502422
Ray S, Reddy AB (2020) COVID-19 management in light of the circadian clock. Nat Rev Mol Cell Biol 21(9):494–495. https://doi.org/10.1038/s41580-020-0275-3
Morin CM, Carrier J, Bastien C, Godbout R, Canadian S, Circadian N (2020) Sleep and circadian rhythm in response to the COVID-19 pandemic. Can J Public Health 111(5):654–657. https://doi.org/10.17269/s41997-020-00382-7
Haspel JA, Anafi R, Brown MK, Cermakian N, Depner C, Desplats P, Gelman AE, Haack M, Jelic S, Kim BS, Laposky AD, Lee YC, Mongodin E, Prather AA, Prendergast BJ, Reardon C, Shaw AC, Sengupta S, Szentirmai É, Thakkar M, Walker WE, Solt LA (2020) Perfect timing: circadian rhythms, sleep, and immunity - an NIH workshop summary. JCI Insight. https://doi.org/10.1172/jci.insight.131487
Levi F, Schibler U (2007) Circadian rhythms: mechanisms and therapeutic implications. Annu Rev Pharmacol Toxicol 47:593–628. https://doi.org/10.1146/annurev.pharmtox.47.120505.105208
Haus E, Halberg F, Pauly JE, Cardoso S, Kuhl JF, Sothern RB, Shiotsuka RN, Hwang DS (1972) Increased tolerance of leukemic mice to arabinosyl cytosine with schedule adjusted to circadian system. Science 177(4043):80–82. https://doi.org/10.1126/science.177.4043.80
Musiek ES, Fitzgerald GA (2013) Molecular clocks in pharmacology. Handb Exp Pharmacol 217(217):243–260. https://doi.org/10.1007/978-3-642-25950-0_10
Konturek PC, Brzozowski T, Konturek SJ (2011) Gut clock: implication of circadian rhythms in the gastrointestinal tract. J Physiol Pharmacol 62(2):139–150
Bron R, Furness JB (2009) Rhythm of digestion: keeping time in the gastrointestinal tract. Clin Exp Pharmacol Physiol 36(10):1041–1048. https://doi.org/10.1111/j.1440-1681.2009.05254.x
Pan X, Hussain MM (2009) Clock is important for food and circadian regulation of macronutrient absorption in mice. J Lipid Res 50(9):1800–1813. https://doi.org/10.1194/jlr.M900085-JLR200
Pan X, Zhang Y, Wang L, Hussain MM (2010) Diurnal regulation of MTP and plasma triglyceride by CLOCK is mediated by SHP. Cell Metab 12(2):174–186. https://doi.org/10.1016/j.cmet.2010.05.014
Park SI, Felipe CR, Pinheiro-Machado PG, Garcia R, Tedesco-Silva H Jr, Medina-Pestana JO (2007) Circadian and time-dependent variability in tacrolimus pharmacokinetics. Fundam Clin Pharmacol 21(2):191–197. https://doi.org/10.1111/j.1472-8206.2007.00468.x
Baraldo M (2008) The influence of circadian rhythms on the kinetics of drugs in humans. Expert Opin Drug Metab Toxicol 4(2):175–192. https://doi.org/10.1517/17425255.4.2.175
Goo RH, Moore JG, Greenberg E, Alazraki NP (1987) Circadian variation in gastric emptying of meals in humans. Gastroenterology 93(3):515–518. https://doi.org/10.1016/0016-5085(87)90913-9
Stearns AT, Balakrishnan A, Rhoads DB, Ashley SW, Tavakkolizadeh A (2008) Diurnal rhythmicity in the transcription of jejunal drug transporters. J Pharmacol Sci 108(1):144–148. https://doi.org/10.1254/jphs.08100sc
Yu F, Zhang T, Zhou C, Xu H, Guo L, Chen M, Wu B (2019) The circadian clock gene Bmal1 controls intestinal exporter MRP2 and drug disposition. Theranostics 9(10):2754–2767. https://doi.org/10.7150/thno.33395
Murakami Y, Higashi Y, Matsunaga N, Koyanagi S, Ohdo S (2008) Circadian clock-controlled intestinal expression of the multidrug-resistance gene mdr1a in mice. Gastroenterology 135(5):1636–1644. https://doi.org/10.1053/j.gastro.2008.07.073
Hamdan AM, Koyanagi S, Wada E, Kusunose N, Murakami Y, Matsunaga N, Ohdo S (2012) Intestinal expression of mouse Abcg2/breast cancer resistance protein (BCRP) gene is under control of circadian clock-activating transcription factor-4 pathway. J Biol Chem 287(21):17224–17231. https://doi.org/10.1074/jbc.M111.333377
Dietrich CG, Geier A, Oude Elferink RP (2003) ABC of oral bioavailability: transporters as gatekeepers in the gut. Gut 52(12):1788–1795. https://doi.org/10.1136/gut.52.12.1788
Saito H, Terada T, Shimakura J, Katsura T, Inui K (2008) Regulatory mechanism governing the diurnal rhythm of intestinal H+/peptide cotransporter 1 (PEPT1). Am J Physiol Gastrointest Liver Physiol 295(2):G395-402. https://doi.org/10.1152/ajpgi.90317.2008
Tarquini B, Cavallini V, Cariddi A, Checchi M, Sorice V, Cecchettin M (1988) Prominent circadian absorption of intranasal salmon calcitonin (SCT) in healthy subjects. Chronobiol Int 5(2):149–152. https://doi.org/10.3109/07420528809079555
Malfatti MA, Kuhn EA, Murugesh DK, Mendez ME, Hum N, Thissen JB, Jaing CJ, Loots GG (2020) Manipulation of the gut microbiome alters acetaminophen biodisposition in mice. Sci Rep 10(1):4571. https://doi.org/10.1038/s41598-020-60982-8
Clarke G, Sandhu KV, Griffin BT, Dinan TG, Cryan JF, Hyland NP (2019) Gut reactions: breaking down xenobiotic-microbiome interactions. Pharmacol Rev 71(2):198–224. https://doi.org/10.1124/pr.118.015768
Frazier K, Chang EB (2020) Intersection of the gut microbiome and circadian rhythms in metabolism. Trends Endocrinol Metab 31(1):25–36. https://doi.org/10.1016/j.tem.2019.08.013
Liang X, FitzGerald GA (2017) Timing the microbes: the circadian rhythm of the gut microbiome. J Biol Rhythms 32(6):505–515. https://doi.org/10.1177/0748730417729066
Yosipovitch G, Sackett-Lundeen L, Goon A, Yiong Huak C, Leok Goh C, Haus E (2004) Circadian and ultradian (12 h) variations of skin blood flow and barrier function in non-irritated and irritated skin—effect of topical corticosteroids. J Investig Dermatol 122(3):824–829. https://doi.org/10.1111/j.0022-202X.2004.22313.x
Conroy DA, Spielman AJ, Scott RQ (2005) Daily rhythm of cerebral blood flow velocity. J Circadian Rhythms 3(1):3–3. https://doi.org/10.1186/1740-3391-3-3
Lemmer B, Nold G (1991) Circadian changes in estimated hepatic blood flow in healthy subjects. Br J Clin Pharmacol 32(5):627–629. https://doi.org/10.1111/j.1365-2125.1991.tb03964.x
Gillette JR (1971) Factors affecting drug metabolism. Ann N Y Acad Sci 179(1):43–66. https://doi.org/10.1111/j.1749-6632.1971.tb46890.x
Angeli A, Frajria R, De Paoli R, Fonzo D, Ceresa F (1978) Diurnal variation of prednisolone binding to serum corticosteroid-binding globulin in man. Clin Pharmacol Ther 23(1):47–53. https://doi.org/10.1002/cpt197823147
Rocci ML, D’Ambrosio R, Johnson NF, JusKo WJ (1982) Prednisolone binding to albumin and transcortin in the presence of cortisol. Biochem Pharmacol 31(3):289–292. https://doi.org/10.1016/0006-2952(82)90172-1
Debono M, Harrison RF, Whitaker MJ, Eckland D, Arlt W, Keevil BG, Ross RJ (2016) Salivary cortisone reflects cortisol exposure under physiological conditions and after hydrocortisone. J Clin Endocrinol Metab 101(4):1469–1477. https://doi.org/10.1210/jc.2015-3694
Melin J, Hartung N, Parra-Guillen ZP, Whitaker MJ, Ross RJ, Kloft C (2019) The circadian rhythm of corticosteroid-binding globulin has little impact on cortisol exposure after hydrocortisone dosing. Clin Endocrinol 91(1):33–40. https://doi.org/10.1111/cen.13969
Levy G (1994) Pharmacologic target-mediated drug disposition. Clin Pharmacol Ther 56(3):248–252. https://doi.org/10.1038/clpt.1994.134
Volz A-K, Dingemanse J, Krause A, Lehr T (2019) Target-mediated population pharmacokinetic modeling of endothelin receptor antagonists. Pharm Res 37(1):2. https://doi.org/10.1007/s11095-019-2723-3
Wilkinson GR, Shand DG (1975) Commentary: a physiological approach to hepatic drug clearance. Clin Pharmacol Ther 18(4):377–390. https://doi.org/10.1002/cpt1975184377
Gries JM, Benowitz N, Verotta D (1996) Chronopharmacokinetics of nicotine. Clin Pharmacol Ther 60(4):385–395. https://doi.org/10.1016/s0009-9236(96)90195-2
Harris BE, Song R, Soong SJ, Diasio RB (1990) Relationship between dihydropyrimidine dehydrogenase activity and plasma 5-fluorouracil levels with evidence for circadian variation of enzyme activity and plasma drug levels in cancer patients receiving 5-fluorouracil by protracted continuous infusion. Cancer Res 50(1):197–201
Harris BE, Song RL, Soong SJ, Diasio RB (1989) Circadian variation of 5-fluorouracil catabolism in isolated perfused rat liver. Cancer Res 49(23):6610–6614
Zhang YK, Yeager RL, Klaassen CD (2009) Circadian expression profiles of drug-processing genes and transcription factors in mouse liver. Drug Metab Dispos 37(1):106–115. https://doi.org/10.1124/dmd.108.024174
Dong D, Yang D, Lin L, Wang S, Wu B (2020) Circadian rhythm in pharmacokinetics and its relevance to chronotherapy. Biochem Pharmacol 178:114045. https://doi.org/10.1016/j.bcp.2020.114045
Lu D, Zhao M, Chen M, Wu B (2020) Circadian clock-controlled drug metabolism Implications for chronotherapeutics. Drug Metabol Disposit. https://doi.org/10.1124/dmd.120.090472
Guo L, Yu F, Zhang T, Wu B (2018) The clock protein bmal1 regulates circadian expression and activity of sulfotransferase 1a1 in Mice. Drug Metab Dispos 46(10):1403. https://doi.org/10.1124/dmd.118.082503
Lin Y, Wang S, Zhou Z, Guo L, Yu F, Wu B (2019) Bmal1 regulates circadian expression of cytochrome P450 3a11 and drug metabolism in mice. Commun Biol 2:378. https://doi.org/10.1038/s42003-019-0607-z
Firsov D, Bonny O (2018) Circadian rhythms and the kidney. Nat Rev Nephrol 14(10):626–635. https://doi.org/10.1038/s41581-018-0048-9
Koopman MG, Koomen GC, Krediet RT, de Moor EA, Hoek FJ, Arisz L (1989) Circadian rhythm of glomerular filtration rate in normal individuals. Clin Sci (Lond) 77(1):105–111. https://doi.org/10.1042/cs0770105
Prins JM, Weverling GJ, van Ketel RJ, Speelman P (1997) Circadian variations in serum levels and the renal toxicity of aminoglycosides in patients. Clin Pharmacol Ther 62(1):106–111. https://doi.org/10.1016/s0009-9236(97)90156-9
Stow LR, Gumz ML (2011) The circadian clock in the kidney. J Am Soc Nephrol 22(4):598–604. https://doi.org/10.1681/ASN.2010080803
Zuber AM, Centeno G, Pradervand S, Nikolaeva S, Maquelin L, Cardinaux L, Bonny O, Firsov D (2009) Molecular clock is involved in predictive circadian adjustment of renal function. Proc Natl Acad Sci 106(38):16523. https://doi.org/10.1073/pnas.0904890106
Mesnard-Ricci B, White CA (1998) Chronokinetics of active biliary ampicillin secretion in rats. Chronobiol Int 15(4):309–321. https://doi.org/10.3109/07420529808998692
Nikolaeva S, Ansermet C, Centeno G, Pradervand S, Bize V, Mordasini D, Henry H, Koesters R, Maillard M, Bonny O, Tokonami N, Firsov D (2016) Nephron-specific deletion of circadian clock gene bmal1 alters the plasma and renal metabolome and impairs drug disposition. J Am Soc Nephrol 27(10):2997–3004. https://doi.org/10.1681/asn.2015091055
Takane H, Ohdo S, Yamada T, Yukawa E, Higuchi S (2000) Chronopharmacology of antitumor effect induced by interferon-beta in tumor-bearing mice. J Pharmacol Exp Ther 294(2):746–752
Mager DE, Neuteboom B, Efthymiopoulos C, Munafo A, Jusko WJ (2003) Receptor-mediated pharmacokinetics and pharmacodynamics of interferon-beta1a in monkeys. J Pharmacol Exp Ther 306(1):262–270. https://doi.org/10.1124/jpet.103.049502
Koyanagi S, Kuramoto Y, Nakagawa H, Aramaki H, Ohdo S, Soeda S, Shimeno H (2003) A molecular mechanism regulating circadian expression of vascular endothelial growth factor in tumor cells. Can Res 63(21):7277
Nakagawa H, Takiguchi T, Nakamura M, Furuyama A, Koyanagi S, Aramaki H, Higuchi S, Ohdo S (2006) Basis for dosing time-dependent change in the anti-tumor effect of imatinib in mice. Biochem Pharmacol 72(10):1237–1245. https://doi.org/10.1016/j.bcp.2006.08.002
Lauriola M, Enuka Y, Zeisel A, D’Uva G, Roth L, Sharon-Sevilla M, Lindzen M, Sharma K, Nevo N, Feldman M, Carvalho S, Cohen-Dvashi H, Kedmi M, Ben-Chetrit N, Chen A, Solmi R, Wiemann S, Schmitt F, Domany E, Yarden Y (2014) Diurnal suppression of EGFR signalling by glucocorticoids and implications for tumour progression and treatment. Nat Commun 5(1):5073. https://doi.org/10.1038/ncomms6073
Kusunose N, Koyanagi S, Hamamura K, Matsunaga N, Yoshida M, Uchida T, Tsuda M, Inoue K, Ohdo S (2010) Molecular basis for the dosing time-dependency of anti-allodynic effects of gabapentin in a mouse model of neuropathic pain. Mol Pain 6:83. https://doi.org/10.1186/1744-8069-6-83
Fujiwara Y, Ando H, Ushijima K, Horiguchi M, Yamashita C, Fujimura A (2017) Dosing-time-dependent effect of rivaroxaban on coagulation activity in rats. J Pharmacol Sci 134(4):234–238. https://doi.org/10.1016/j.jphs.2017.08.001
Brunner-Ziegler S, Jilma B, Schörgenhofer C, Winkler F, Jilma-Stohlawetz P, Koppensteiner R, Quehenberger P, Seger C, Weigel G, Griesmacher A, Brunner M (2016) Comparison between the impact of morning and evening doses of rivaroxaban on the circadian endogenous coagulation rhythm in healthy subjects. J Thromb Haemost 14(2):316–323. https://doi.org/10.1111/jth.13213
Antoch MP, Kondratov RV (2013) Pharmacological modulators of the circadian clock as potential therapeutic drugs: focus on genotoxic/anticancer therapy. Handb Exp Pharmacol 217:289–309. https://doi.org/10.1007/978-3-642-25950-0_12
Sulli G, Manoogian ENC, Taub PR, Panda S (2018) Training the circadian clock, clocking the drugs, and drugging the clock to prevent, manage, and treat chronic diseases. Trends Pharmacol Sci 39(9):812–827. https://doi.org/10.1016/j.tips.2018.07.003
Chakraborty A, Krzyzanski W, Jusko WJ (1999) Mathematical modeling of circadian cortisol concentrations using indirect response models: comparison of several methods. J Pharmacokinet Biopharm 27(1):23–43. https://doi.org/10.1023/A:1020678628317
Refinetti R, Lissen GC, Halberg F (2007) Procedures for numerical analysis of circadian rhythms. Biol Rhythm Res 38(4):275–325. https://doi.org/10.1080/09291010600903692
Xu J, Winkler J, Sabarinath SN, Derendorf H (2008) Assessment of the impact of dosing time on the pharmacokinetics/pharmacodynamics of prednisolone. AAPS J 10(2):331–341. https://doi.org/10.1208/s12248-008-9038-3
Ayyar VS, DuBois DC, Almon RR, Jusko WJ (2017) Mechanistic multi-tissue modeling of glucocorticoid-induced leucine zipper regulation: Integrating circadian gene expression with receptor-mediated corticosteroid pharmacodynamics. J Pharmacol Exp Ther 363(1):45–57. https://doi.org/10.1124/jpet.117.242990
Ayyar VS, DuBois DC, Almon RR, Jusko WJ (2019) Modeling corticosteroid pharmacokinetics and pharmacodynamics, part III: estrous cycle and estrogen receptor-dependent antagonism of glucocorticoid-induced leucine zipper (GILZ) enhancement by corticosteroids. J Pharmacol Exp Ther 370(2):337–349. https://doi.org/10.1124/jpet.119.257543
Sukumaran S, Jusko WJ, DuBois DC, Almon RR (2011) Mechanistic modeling of the effects of glucocorticoids and circadian rhythms on adipokine expression. J Pharmacol Exp Ther 337(3):734–746. https://doi.org/10.1124/jpet.111.179960
Hazra A, Pyszczynski N, DuBois DC, Almon RR, Jusko WJ (2007) Modeling receptor/gene-mediated effects of corticosteroids on hepatic tyrosine aminotransferase dynamics in rats: dual regulation by endogenous and exogenous corticosteroids. J Pharmacokinet Pharmacodyn 34(5):643–667. https://doi.org/10.1007/s10928-007-9063-3
Ayyar VS, Almon RR, Jusko WJ, DuBois DC (2015) Quantitative tissue-specific dynamics of in vivo GILZ mRNA expression and regulation by endogenous and exogenous glucocorticoids. Physiol Rep. https://doi.org/10.14814/phy2.12382
Ramakrishnan R, DuBois DC, Almon RR, Pyszczynski NA, Jusko WJ (2002) Fifth-generation model for corticosteroid pharmacodynamics: application to steady-state receptor down-regulation and enzyme induction patterns during seven-day continuous infusion of methylprednisolone in rats. J Pharmacokinet Pharmacodyn 29(1):1–24. https://doi.org/10.1023/a:1015765201129
Scherholz ML, Schlesinger N, Androulakis IP (2019) Chronopharmacology of glucocorticoids. Adv Drug Deliv Rev 151–152:245–261. https://doi.org/10.1016/j.addr.2019.02.004
Ayyar VS, Krzyzanski W, Jusko WJ (2019) Indirect pharmacodynamic models for responses with circadian removal. J Pharmacokinet Pharmacodyn 46(1):89–101. https://doi.org/10.1007/s10928-019-09620-z
Koch G, Schropp J (2018) Delayed logistic indirect response models: realization of oscillating behavior. J Pharmacokinet Pharmacodyn 45(1):49–58. https://doi.org/10.1007/s10928-017-9563-8
Butcher EC, Berg EL, Kunkel EJ (2004) Systems biology in drug discovery. Nat Biotechnol 22(10):1253–1259. https://doi.org/10.1038/nbt1017
Goodwin BC (1965) Oscillatory behavior in enzymatic control processes. Adv Enzyme Regul 3:425–437. https://doi.org/10.1016/0065-2571(65)90067-1
Millius A, Ueda H (2017) Systems biology derived discoveries of intrinsic clocks. Fronti Neurol. https://doi.org/10.3389/fneur.2017.00025
Thron CD (1991) The secant condition for instability in biochemical feedback control I—The role of cooperativity and saturability. Bull Math Biol 53(3):383–401. https://doi.org/10.1016/S0092-8240(05)80394-5
Forger DB (2011) Signal processing in cellular clocks. Proc Natl Acad Sci 108(11):4281. https://doi.org/10.1073/pnas.1004720108
Mavroudis PD, DuBois DC, Almon RR, Jusko WJ (2018) Modeling circadian variability of core-clock and clock-controlled genes in four tissues of the rat. PLoS ONE 13(6):e0197534. https://doi.org/10.1371/journal.pone.0197534
Scheff JD, Calvano SE, Lowry SF, Androulakis IP (2010) Modeling the influence of circadian rhythms on the acute inflammatory response. J Theor Biol 264(3):1068–1076. https://doi.org/10.1016/j.jtbi.2010.03.026
Mavroudis PD, Scheff JD, Calvano SE, Androulakis IP (2013) Systems biology of circadian-immune interactions. J Innate Immun 5(2):153–162. https://doi.org/10.1159/000342427
Pierre K, Rao RT, Hartmanshenn C, Androulakis IP (2018) Modeling the influence of seasonal differences in the hpa axis on synchronization of the circadian clock and cell cycle. Endocrinology 159(4):1808–1826. https://doi.org/10.1210/en.2017-03226
Ayyar VS, Jusko WJ (2020) Transitioning from Basic toward Systems Pharmacodynamic Models: Lessons from Corticosteroids. Pharmacol Rev 72(2):414. https://doi.org/10.1124/pr.119.018101
Kim JK, Forger DB, Marconi M, Wood D, Doran A, Wager T, Chang C, Walton KM (2013) Modeling and validating chronic pharmacological manipulation of circadian rhythms. CPT Pharmacometrics Syst Pharmacol 2(7):e57. https://doi.org/10.1038/psp.2013.34
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ayyar, V.S., Sukumaran, S. Circadian rhythms: influence on physiology, pharmacology, and therapeutic interventions. J Pharmacokinet Pharmacodyn 48, 321–338 (2021). https://doi.org/10.1007/s10928-021-09751-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10928-021-09751-2