Abstract
The effect of extrapolated area (%AUCextrap) on estimating mean AUCinf in a simulated single-dose clinical trial was examined. Concentration–time (C–t) profiles from 12 to 36 subjects for 1- and 2-compartment models after single dose administration were simulated with increasing right-tailed censoring. Each subject’s %AUCextrap and AUCinf was calculated using eight different methods, including noncompartmental analysis (NCA), population-based methods, and maximum likelihood (ML) accounting for censoring. Each method’s geometric mean AUCinf and percent relative error (PRE) from the true AUCinf was calculated. This was repeated 100 times and the mean PRE (MPRE) was calculated. Mean %AUCextrap ranged from 1 to ~30 % for the 1-compartment and 2 to 32 % for the 2-compartment model at the lowest and highest degree of censoring, respectively. NCA methods using all subjects to estimate the population mean AUCinf had similar or less bias (within ± 20 %) than when those subjects with >20 % %AUCextrap were removed. Using Cpred compared to Clast in the calculation of individual AUCinf resulted in no performance improvement. Linear mixed effects models to estimate λz and ML methods accounting for censoring resulted in either no improvement or increased bias when censoring was high. Population pharmacokinetic method bias was dependent on the nature of the C–t profile. When the C–t profile declined biphasically, population models had higher bias than NCA methods but were superior when the C–t profile decline in a log-linear manner. It is recommended that subjects with high %AUCextrap should not be removed from the estimation of mean AUCinf in NCA analyses.





Similar content being viewed by others
References
Purves RD (1992) Optimum numerical integration methods for area-under-the-curve (AUC) and area-under-the-moment-curve (AUMC). J Pharmacokinet Biopharm 20:211–226
Muir KT, Gomeni R (2004) Non-compartmental analysis. In: Bonate PL, Howard DR (eds) Pharmacokinetics in drug development: clinical study design and analysis. AAPS, Arlington, pp 235–265
United States Department of Health and Human Services, Food and Drug Administration, Center for Drug Evaluation and Research, Center for Biologics Evaluation and Research (2003) Guidance for industry: bioavailability and bioequivalence studies for orally administered drug products—general considerations
European Medicines Agency (EMEA), Committee for Medicinal Products for Human Use (CHMP) (2010) Guideline on the investigation of bioequivalence
Department of Health and Human Services, Food and Drug Administration (2013) 21 CFR Parts 314 and 320; Bioavailability and bioequivalence requirements; abbreviated applications; final rule
United States Department of Health and Human Services, Food and Drug Administration, Center for Drug Evaluation and Research, Center for Biologics Evaluation and Research (2001) Guidance for industry: statistical approaches to establishing bioequivalence
Bain LJ, Engelhardt M (1987) Introduction to probability and mathematical statistics. Duxbury, Boston
Bonate PL (2011) Pharmacokinetic—pharmacodynamic modeling and simulation, 2nd edn. Springer, New York
Ahn JE, Karlsson MO, Dunne A, Ludden TM (2008) Likelihood based approaches to handling data below the quantification limit using NONMEM VI. J Pharmacokinet Pharmacodyn 35:401–421
Lawless JF (2003) Statistical models and methods for lifetime data, 2nd edn. Wiley, New York
SAS Institute (2009) SAS/STAT 9.2 user’s guide, 2nd edn. Sas Institute, Cary
Beal SL (2001) Ways to fit a PK model with some data below the quantification limit. J Pharmacokinet Pharmacodyn 28:481–504
Akaike H (1974) A new look at the statistical model identification. IEEE Trans Automat Control 19:716–723
Svensson EM, Dooley KE, Karlsson MO (2014) Prediction of pharmacokinetic interactions for drugs with a long half-life: evidence for the need of model-based analysis. Presented at the Population Analysis Group Europe 2014, Alicante, Spain
Kleinbaum DG, Klein M (2005) Survival analysis: a self-learning text, 2nd edn. Springer, New York
Lee ET, Wang JW (2003) Statistical methods for survival data analysis. John Wiley & Sons, Inc., New York
Ranganathan P, Pramesh CS (2012) Censoring in survival analysis: potential for bias. Perspect Clin Res 3:40
Karlsson MO, Savic RM (2007) Diagnosing model diagnostics. Clin Pharmacol Ther 82:1–17
Bois FY, Tozer TN, Hauck WH, Chen M-L, Patnaik R, Williams RL (1994) Bioequivalence: performance of several measures of extent of absorption. Pharm Res 11:715–722
Cawello W (1999) Parameters for compartment-free pharmacokinetics: standardisation of study design, data analysis, and reporting. Shaker, Germany
Acknowledgments
The author would like to thank Yaning Wang for his very thoughtful and helpful comments on this manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
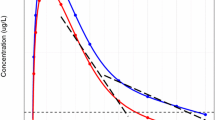
Representative concentration–time profiles on a log-linear scale for the 5 pharmacokinetic models studied. Red line is the theoretical population mean concentration–time profile. Gray lines are spaghetti plots of 10 subject’s observed concentrations with random measurement error added to the true concentration. Black line is the LLOQ. Observed concentrations below the LLOQ were censored. (JPEG 82 kb)
Supplemental Fig. 2
Representative concentration–time profiles on a linear scale for the 5 pharmacokinetic models studied. Red line is the theoretical population mean concentration–time profile. Gray lines are spaghetti plots of 10 subject’s observed concentrations with random measurement error added to the true concentration. Observed concentrations below the LLOQ were censored. Since a reference line for 0 LLOQ cannot be shown on a log-linear scale, the reference line was rescaled to the minimum axis value of 0.1. (JPEG 77 kb)
Supplemental Fig. 3
Plot of MPRE as function of LLOQ and number of subjects for the 1-compartment model with additive plus proportional residual error. Each marker represents the mean of 100 simulation replications. See Table 2 for marker definitions. Blue band is the approximate range where the mean %AUCextrap was <20 %. (JPEG 44 kb)
Supplemental Fig. 4
Plot of MPRE as function of LLOQ and number of subjects for the 2-compartment model with additive plus proportional residual error. Each marker represents the mean of 100 simulation replications. See Table 2 for marker definitions. Blue band is the approximate range where the mean %AUCextrap was <20 %. (JPEG 46 kb)
Supplemental Fig. 5
Plot of MPRE as function of LLOQ and number of subjects for the 1-compartment model with beta(0.5, 0.5) residual error. Each marker represents the mean of 100 simulation replications. See Table 2 for marker definitions. Blue band is the approximate range where the mean %AUCextrap was <20 %. (JPEG 46 kb)
Supplemental Fig. 6
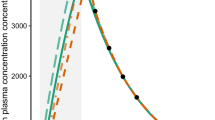
Representative concentration–time profiles for Simulation 3 where Clast was contaminated by uniform random error to ensure that Clast > Cprior. A total of 24 subjects were simulated. Subjects with contaminated Clast values are shown in red. Data shown from 2 to 8 time units post post-dose to highlight the effect of contamination. (JPEG 63 kb)
Rights and permissions
About this article
Cite this article
Bonate, P.L. Simulation studies on the estimation of total area under the curve in the presence of right-tailed censoring. J Pharmacokinet Pharmacodyn 42, 19–32 (2015). https://doi.org/10.1007/s10928-014-9395-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10928-014-9395-8