Abstract
Approximately 75% of all breast cancers express the nuclear hormone receptor estrogen receptor α (ERα). However, the majority of mammary tumors from genetically engineered mouse models (GEMMs) are ERα-negative. To model ERα-positive breast cancer in mice, we exogenously introduced expression of mouse and human ERα in an existing GEMM of p53-deficient breast cancer. After initial ERα expression during mammary gland development, expression was reduced or lost in adult glands and p53-deficient mammary tumors. Chromatin immunoprecipitation (ChIP)-sequencing analysis of primary mouse mammary epithelial cells (MMECs) derived from these models, in which expression of the ERα constructs was induced in vitro, confirmed interaction of ERα with the DNA. In human breast and endometrial cancer, and also in healthy breast tissue, DNA binding of ERα is facilitated by the pioneer factor FOXA1. Surprisingly, the ERα binding sites identified in primary MMECs, but also in mouse mammary gland and uterus, showed an high enrichment of ERE motifs, but were devoid of Forkhead motifs. Furthermore, exogenous introduction of FOXA1 and GATA3 in ERα-expressing MMECs was not sufficient to promote ERα-responsiveness of these cells. Together, this suggests that species-specific differences in pioneer factor usage between mouse and human are dictated by the DNA sequence, resulting in ERα-dependencies in mice that are not FOXA1 driven. These species-specific differences in ERα-biology may limit the utility of mice for in vivo modeling of ERα-positive breast cancer.





Similar content being viewed by others
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Perou CM, Sørlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, et al. Molecular portraits of human breast tumours. Nature. 2000;406:747–52.
Sørlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H, et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci U S A. 2001;98:10869–74.
Brünner N, Bronzert D, Vindeløv LL, Rygaard K, Spang-Thomsen M, Lippman ME. Effect on growth and cell cycle kinetics of estradiol and tamoxifen on MCF-7 human breast cancer cells grown in vitro and in nude mice. Cancer Res. 1989;49:1515–20.
Arnal J-F, Lenfant F, Metivier R, Flouriot G, Henrion D, Adlanmerini M, et al. Membrane and nuclear estrogen receptor alpha actions: from tissue specificity to medical implications. Physiol Rev. 2017;97:1045–87.
Gruber CJ, Tschugguel W, Schneeberger C, Huber JC. Production and actions of estrogens. N Engl J Med. 2002;346:340–52.
Kumar R, Zakharov MN, Khan SH, Miki R, Jang H, Toraldo G, et al. The dynamic structure of the estrogen receptor. J Amino Acids. 2011;2011:812540.
Zaret KS, Carroll JS. Pioneer transcription factors: establishing competence for gene expression. Genes Dev. 2011;25:2227–41.
Hurtado A, Holmes KA, Ross-Innes CS, Schmidt D, Carroll JS. FOXA1 is a key determinant of estrogen receptor function and endocrine response. Nat Genet. 2011;43:27–33.
Theodorou V, Stark R, Menon S, Carroll JS. GATA3 acts upstream of FOXA1 in mediating ESR1 binding by shaping enhancer accessibility. Genome Res. 2013;23:12–22.
Kushner PJ, Agard DA, Greene GL, Scanlan TS, Shiau AK, Uht RM, et al. Estrogen receptor pathways to AP-1. J Steroid Biochem Mol Biol. 2000;74:311–7.
Safe S. Transcriptional activation of genes by 17 beta-estradiol through estrogen receptor-Sp1 interactions. Vitam Horm. 2001;62:231–52.
McKenna NJ, Lanz RB, O’Malley BW. Nuclear receptor coregulators: cellular and molecular biology. Endocr Rev. 1999;20:321–44.
Wärnmark A, Treuter E, Wright APH, Gustafsson J-Å. Activation functions 1 and 2 of nuclear receptors: molecular strategies for transcriptional activation. Mol Endocrinol. 2003;17:1901–9.
Yaşar P, Ayaz G, User SD, Güpür G, Muyan M. Molecular mechanism of estrogen–estrogen receptor signaling. Reprod Med Biol. 2017;16:4–20.
Xu J, Wu R-C, O’Malley BW. Normal and cancer-related functions of the p160 steroid receptor co-activator (SRC) family. Nat Rev Cancer. 2009;9:615–30.
Özdemir BC, Sflomos G, Brisken C. The challenges of modeling hormone receptor-positive breast cancer in mice. Endocr Relat Cancer. 2018;25:R319–30.
Osborne CK, Hobbs K, Clark GM. Effect of estrogens and antiestrogens on growth of human breast cancer cells in athymic nude mice. Cancer Res. 1985;45:584–90.
Wagner K-U. Models of breast cancer: quo vadis, animal modeling? Breast Cancer Res. 2004;6:31–8.
Vargo-Gogola T, Rosen JM. Modelling breast cancer: one size does not fit all. Nat Rev Cancer. 2007;7:659–72.
Dabydeen SA, Furth PA. Genetically engineered ERα-positive breast cancer mouse models. Endocr Relat Cancer. 2014;21:R195–208.
Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490:61–70.
Ciriello G, Gatza ML, Beck AH, Wilkerson MD, Rhie SK, Pastore A, et al. Comprehensive molecular portraits of invasive lobular breast cancer. Cell. 2015;163:506–19.
Derksen PWB, Braumuller TM, van der Burg E, Hornsveld M, Mesman E, Wesseling J, et al. Mammary-specific inactivation of E-cadherin and p53 impairs functional gland development and leads to pleomorphic invasive lobular carcinoma in mice. Dis Model Mech. 2011;4:347–58.
Jonkers J, Meuwissen R, van der Gulden H, Peterse H, van der Valk M, Berns A. Synergistic tumor suppressor activity of BRCA2 and p53 in a conditional mouse model for breast cancer. Nat Genet. 2001;29:418–25.
Huijbers IJ, Del Bravo J, Bin Ali R, Pritchard C, Braumuller TM, van Miltenburg MH, et al. Using the GEMM-ESC strategy to study gene function in mouse models. Nat Protoc. 2015;10:1755–85.
Henneman L, van Miltenburg MH, Michalak EM, Braumuller TM, Jaspers JE, Drenth AP, et al. Selective resistance to the PARP inhibitor olaparib in a mouse model for BRCA1-deficient metaplastic breast cancer. Proc Natl Acad Sci U S A. 2015;112:8409–14.
Boelens MC, Nethe M, Klarenbeek S, de Ruiter JR, Schut E, Bonzanni N, et al. PTEN loss in E-cadherin-deficient mouse mammary epithelial cells rescues apoptosis and results in development of classical invasive lobular carcinoma. Cell Rep. 2016;16:2087–101.
Kas SM, de Ruiter JR, Schipper K, Annunziato S, Schut E, Klarenbeek S, et al. Insertional mutagenesis identifies drivers of a novel oncogenic pathway in invasive lobular breast carcinoma. Nat Genet. 2017;49:1219–30.
Follenzi A, Ailles LE, Bakovic S, Geuna M, Naldini L. Gene transfer by lentiviral vectors is limited by nuclear translocation and rescued by HIV-1 pol sequences. Nat Genet. 2000;25:217–22.
Trapnell C, Pachter L, Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinforma Oxf Engl. 2009;25:1105–11.
Langmead B, Trapnell C, Pop M, Salzberg SL. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009;10:R25.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence alignment/map format and SAMtools. Bioinforma Oxf Engl. 2009;25:2078–9.
Anders S, Pyl PT, Huber W. HTSeq – a Python framework to work with high-throughput sequencing data. Bioinforma Oxf Engl. 2015;31:166–9.
Anders S, Huber W. Differential expression analysis for sequence count data. Genome Biol. 2010;11:R106.
Singh AA, Schuurman K, Nevedomskaya E, Stelloo S, Linder S, Droog M, et al. Optimized ChIP-seq method facilitates transcription factor profiling in human tumors. Life Sci Alliance. 2019;2:e201800115.
Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE, et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 2008;9:R137.
Kumar V, Muratani M, Rayan NA, Kraus P, Lufkin T, Ng HH, et al. Uniform, optimal signal processing of mapped deep-sequencing data. Nat Biotechnol. 2013;31:615–22.
Ye T, Krebs AR, Choukrallah M-A, Keime C, Plewniak F, Davidson I, et al. seqMINER: an integrated ChIP-seq data interpretation platform. Nucleic Acids Res. 2011;39:e35.
Ji X, Li W, Song J, Wei L, Liu XS. CEAS: cis -regulatory element annotation system. Nucleic Acids Res. 2006;34:W551–4.
Liu T, Ortiz JA, Taing L, Meyer CA, Lee B, Zhang Y, et al. Cistrome: an integrative platform for transcriptional regulation studies. Genome Biol. 2011;12:R83.
Battersby S, Robertson B, Anderson T, King R, McPherson K. Influence of menstrual cycle, parity and oral contraceptive use on steroid hormone receptors in normal breast. Br J Cancer. 1992;65:601–7.
Goyal R, Gupta T, Gupta R, Aggarwal A, Sahni D, Singh G. Histological and immunohistochemical study of estrogen and progesterone receptors in normal human breast tissue in adult age groups vulnerable to malignancy: histological and IHC study of ER and PR in normal breast tissue. Clin Anat. 2016;29:729–37.
Nilsson ME, Vandenput L, Tivesten Å, Norlén A-K, Lagerquist MK, Windahl SH, et al. Measurement of a comprehensive sex steroid profile in rodent serum by high-sensitive gas chromatography-tandem mass spectrometry. Endocrinology. 2015;156:2492–502.
van de Ven M, Liu X, van der Burg E, Klarenbeek S, Alexi X, Zwart W, et al. BRCA1-associated mammary tumorigenesis is dependent on estrogen rather than progesterone signaling. J Pathol. 2018;246:41–53.
Zwart W, Theodorou V, Kok M, Canisius S, Linn S, Carroll JS. Oestrogen receptor–co-factor–chromatin specificity in the transcriptional regulation of breast cancer. EMBO J. 2011;30:4764–76.
Usary J, Llaca V, Karaca G, Presswala S, Karaca M, He X, et al. Mutation of GATA3 in human breast tumors. Oncogene. 2004;23:7669–78.
Jansen MPHM, Knijnenburg T, Reijm EA, Simon I, Kerkhoven R, Droog M, et al. Hallmarks of aromatase inhibitor drug resistance revealed by epigenetic profiling in breast cancer. Cancer Res. 2013;73:6632–41.
Droog M, Nevedomskaya E, Dackus GM, Fles R, Kim Y, Hollema H, et al. Estrogen receptor α wields treatment-specific enhancers between morphologically similar endometrial tumors. Proc Natl Acad Sci U S A. 2017;114:E1316–25.
Chi D, Singhal H, Li L, Xiao T, Liu W, Pun M, et al. Estrogen receptor signaling is reprogrammed during breast tumorigenesis. Proc Natl Acad Sci U S A. 2019;116:11437–43.
Nautiyal J, Steel JH, Mane MR, Oduwole O, Poliandri A, Alexi X, et al. The transcriptional co-factor RIP140 regulates mammary gland development by promoting the generation of key mitogenic signals. Development. 2013;140:1079–89.
Hewitt SC, Li L, Grimm SA, Chen Y, Liu L, Li Y, et al. Research resource: whole-genome estrogen receptor α binding in mouse uterine tissue revealed by ChIP-seq. Mol Endocrinol Baltim Md. 2012;26:887–98.
Kong SL, Li G, Loh SL, Sung W-K, Liu ET. Cellular reprogramming by the conjoint action of ERα, FOXA1, and GATA3 to a ligand-inducible growth state. Mol Syst Biol. 2011;7:526.
Nilsson S, Mäkelä S, Treuter E, Tujague M, Thomsen J, Andersson G, et al. Mechanisms of estrogen action. Physiol Rev. 2001;81:1535–65.
Chou J, Lin JH, Brenot A, Kim J, Provot S, Werb Z. GATA3 suppresses metastasis and modulates the tumour microenvironment by regulating microRNA-29b expression. Nat Cell Biol. 2013;15:201–13.
Yan W, Cao QJ, Arenas RB, Bentley B, Shao R. GATA3 inhibits breast cancer metastasis through the reversal of epithelial-mesenchymal transition. J Biol Chem. 2010;285:14042–51.
Peterson EA, Jenkins EC, Lofgren KA, Chandiramani N, Liu H, Aranda E, et al. Amphiregulin is a critical downstream effector of estrogen signaling in ERα-positive breast cancer. Cancer Res. 2015;75:4830–8.
Altucci L, Addeo R, Cicatiello L, Dauvois S, Parker MG, Truss M, et al. 17beta-estradiol induces cyclin D1 gene transcription, p36D1-p34cdk4 complex activation and p105Rb phosphorylation during mitogenic stimulation of G(1)-arrested human breast cancer cells. Oncogene. 1996;12:2315–24.
Howell A, Osborne CK, Morris C, Wakeling AE. ICI 182,780 (Faslodex™). Cancer. 2000;89:817–25.
Tilli MT, Frech MS, Steed ME, Hruska KS, Johnson MD, Flaws JA, et al. Introduction of estrogen receptor-α into the tTA/TAg conditional mouse model precipitates the development of estrogen-responsive mammary adenocarcinoma. Am J Pathol. 2003;163:1713–9.
Torres-Arzayus MI, de Mora JF, Yuan J, Vazquez F, Bronson R, Rue M, et al. High tumor incidence and activation of the PI3K/AKT pathway in transgenic mice define AIB1 as an oncogene. Cancer Cell. 2004;6:263–74.
Torres-Arzayus MI, Zhao J, Bronson R, Brown M. Estrogen-dependent and independent-mechanisms contribute to AIB1-mediated tumor formation. Cancer Res. 2010;70:4102–11.
Chan SR, Vermi W, Luo J, Lucini L, Rickert C, Fowler AM, et al. STAT1-deficient mice spontaneously develop estrogen receptor α-positive luminal mammary carcinomas. Breast Cancer Res. 2012;14:R16.
Rose-Hellekant TA, Arendt LM, Schroeder MD, Gilchrist K, Sandgren EP, Schuler LA. Prolactin induces ERα-positive and ERα-negative mammary cancer in transgenic mice. Oncogene. 2003;22:4664–74.
Arendt LM, Rugowski DE, Grafwallner-Huseth TA, Garcia-Barchino MJ, Rui H, Schuler LA. Prolactin-induced mouse mammary carcinomas model estrogen resistant luminal breast cancer. Breast Cancer Res. 2011;13:R11.
Chen F, Li A, Gao S, Hollern D, Williams M, Liu F, et al. Tip30 controls differentiation of murine mammary luminal progenitor to estrogen receptor-positive luminal cell through regulating FoxA1 expression. Cell Death Dis. 2014;5:e1242.
Medina D, Kittrell FS, Shepard A, Stephens LC, Jiang C, Lu J, et al. Biological and genetic properties of the p53 null preneoplastic mammary epithelium. FASEB J. 2002;16:881–3.
Medina D, Kittrell FS, Hill J, Shepard A, Thordarson G, Brown P. Tamoxifen inhibition of estrogen receptor-α–negative mouse mammary tumorigenesis. Cancer Res. 2005;65:3493–6.
Mazumdar A, Medina D, Kittrell FS, Zhang Y, Hill JL, Edwards DE, et al. The combination of tamoxifen and the rexinoid LG100268 prevents ER-positive and ER-negative mammary tumors in p53-null mammary gland mice. Cancer Prev Res (Phila Pa). 2012;5:1195–202.
Andò S, Malivindi R, Catalano S, Rizza P, Barone I, Panza S, et al. Conditional expression of Ki-RasG12V in the mammary epithelium of transgenic mice induces estrogen receptor alpha (ERα)-positive adenocarcinoma. Oncogene. 2017;36:6420–31.
Bernardo GM, Lozada KL, Miedler JD, Harburg G, Hewitt SC, Mosley JD, et al. FOXA1 is an essential determinant of ERα expression and mammary ductal morphogenesis. Development. 2010;137:2045–54.
Besnard V, Wert SE, Hull WM, Whitsett JA. Immunohistochemical localization of Foxa1 and Foxa2 in mouse embryos and adult tissues. Gene Expr Patterns GEP. 2004;5:193–208.
Odom DT, Dowell RD, Jacobsen ES, Gordon W, Danford TW, MacIsaac KD, et al. Tissue-specific transcriptional regulation has diverged significantly between human and mouse. Nat Genet. 2007;39:730–2.
Wilson MD, Barbosa-Morais NL, Schmidt D, Conboy CM, Vanes L, Tybulewicz VLJ, et al. Species-specific transcription in mice carrying human chromosome 21. Science. 2008;322:434–8.
Villar D, Berthelot C, Aldridge S, Rayner TF, Lukk M, Pignatelli M, et al. Enhancer evolution across 20 mammalian species. Cell. 2015;160:554–66.
Lin S-CJ, Lee K-F, Nikitin AY, Hilsenbeck SG, Cardiff RD, Li A, et al. Somatic mutation of p53 leads to estrogen receptor α-positive and -negative mouse mammary tumors with high frequency of metastasis. Cancer Res. 2004;64:3525–32.
Wijnhoven SWP, Zwart E, Speksnijder EN, Beems RB, Olive KP, Tuveson DA, et al. Mice expressing a mammary gland–specific R270H mutation in the p53 tumor suppressor gene mimic human breast cancer development. Cancer Res. 2005;65:8166–73.
Díaz-Cruz ES, Furth PA. Deregulated estrogen receptor α and p53 heterozygosity collaborate in the development of mammary hyperplasia. Cancer Res. 2010;70:3965–74.
Stratikopoulos EE, Kiess N, Szabolcs M, Pegno S, Kakit C, Wu X, et al. Mouse ER+/PIK3CA H1047R breast cancers caused by exogenous estrogen are heterogeneously dependent on estrogen and undergo BIM-dependent apoptosis with BH3 and PI3K agents. Oncogene. 2019;38:47–59.
Annunziato S, Kas SM, Nethe M, Yücel H, Bravo JD, Pritchard C, et al. Modeling invasive lobular breast carcinoma by CRISPR/Cas9-mediated somatic genome editing of the mammary gland. Genes Dev. 2016;30:1470–80.
Clarke RB, Howell A, Potten CS, Anderson E. Dissociation between steroid receptor expression and cell proliferation in the human breast. Cancer Res. 1997;57:4987–91.
Clarke RB. Complementary yet distinct roles for oestrogen receptor-α and oestrogen receptor-β in mouse mammary epithelial proliferation. Breast Cancer Res. 2004;6:135–6.
Russo J, Ao X, Grill C, Russo IH. Pattern of distribution of cells positive for estrogen receptor a and progesterone receptor in relation to proliferating cells in the mammary gland. Breast Cancer Res Treat. 1999;53:217–27.
Mallepell S, Krust A, Chambon P, Brisken C. Paracrine signaling through the epithelial estrogen receptor a is required for proliferation and morphogenesis in the mammary gland. Proc Natl Acad Sci U S A. 2006;103:2196–201.
Ciarloni L, Mallepell S, Brisken C. Amphiregulin is an essential mediator of estrogen receptor α function in mammary gland development. Proc Natl Acad Sci U S A. 2007;104:5455–60.
Bocchinfuso WP, Korach KS. Mammary gland development and tumorigenesis in estrogen receptor knockout mice. J Mammary Gland Biol Neoplasia. 1997;2:323–34.
Kouros-Mehr H, Slorach EM, Sternlicht MD, Werb Z. GATA-3 maintains the differentiation of the luminal cell fate in the mammary gland. Cell. 2006;127:1041–55.
Asselin-Labat M-L, Sutherland KD, Barker H, Thomas R, Shackleton M, Forrest NC, et al. Gata-3 is an essential regulator of mammary-gland morphogenesis and luminal-cell differentiation. Nat Cell Biol. 2007;9:201–9.
Russo J, Gusterson BA, Rogers AE, Russo IH, Wellings SR, van Zwieten MJ. Comparative study of human and rat mammary tumorigenesis. Lab Investig J Tech Methods Pathol. 1990;62:244–78.
Nandi S, Guzman RC, Yang J. Hormones and mammary carcinogenesis in mice, rats, and humans: a unifying hypothesis. Proc Natl Acad Sci U S A. 1995;92:3650–7.
Shull JD, Dennison KL, Chack AC, Trentham-Dietz A. Rat models of 17β-estradiol-induced mammary cancer reveal novel insights into breast cancer etiology and prevention. Physiol Genomics. 2018;50:215–34.
Acknowledgements
We are grateful to Eline van der Burg, Ute Boon, Renske de Korte-Grimmerink, Micha Nethe and Yongsoo Kim for their technical support with the experiments. We thank the Netherlands Cancer Institute Genomics Core Facility, Mouse Clinic for Cancer and Aging, Animal Facility, and Animal Pathology Facility for their expert technical support.
GEO accession number:
GSE127863.
Funding
This work was financially supported by the Oncode Institute; the Center for Translational Molecular Medicine (CTMM) Breast Care Project; the Netherlands Organization for Scientific Research (NWO: Cancer Genomics Netherlands (CGCNL), Cancer Systems Biology Center (CSBC), Zenith 93,512,009, Vici 91,814,643); the EU Seventh Framework Program (EurocanPlatform project 260,791); the European Research Council (ERC Synergy project CombatCancer); and a National Roadmap grant for Large-Scale Research facilities from NWO.
Author information
Authors and Affiliations
Contributions
LMC, LH, WZ and JJ designed research. LMC, LH, AD and ES performed research. LMC, LH, RB, SK, WZ and JJ analyzed data. LMC, WZ and JJ wrote the paper.
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that there are no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Fig. S1
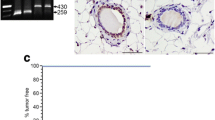
Exogenously-introduced ERα is localized to the nucleus. a PCR analysis of primary MMECs derived from Trp53F/F, Trp53F/F;mERα, Trp53F/F;hERα, Trp53F/F;HA-mERα and Trp53F/F;HA-hERα mice to detect AdCre-induced recombination of the Trp53 locus (Trp53F depicts not-recombined, Trp53Δ depicts recombined DNA) and the Col1a1 locus (lower band depicts not-recombined, upper band depicts recombined DNA). FwT1, FwT10, RvT10 refer to primers used for PCR at the Trp53 locus. FwC1, RvC1, RvC2 refer to primers used for PCR at the Col1a1 locus. Locations of the primers are indicated in Fig. 1a. b Subcellular fractionation of AdCre-induced recombined primary MMECs derived from Trp53F/F, Trp53F/F;mERα and Trp53F/F;HA-mERα (top) and Trp53F/F, Trp53F/F;hERα and Trp53F/F;HA-hERα mice (bottom) shows enrichment of mouse and human ERα mostly in the soluble-nuclear fraction and some in the chromatin-bound fraction in full medium conditions. Tubulin was used as a marker for the cytoplasmic fraction, Topoisomerase 1 (TOP1) for the soluble-nuclear fraction and histone H3 for the chromatin-bound fraction. (PNG 825 kb)
Supplementary Fig. S2
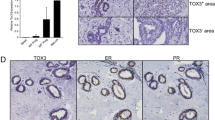
Tumor latency of WP-HA-mERα and WP-HA-hERα mice is affected to a similar extent as WP control mice by 17β-estradiol stimulation due to methylation-mediated silencing. a Kaplan-Meier analysis of mammary tumor-specific survival reflecting modest differences between the ERα overexpressing models compared to WP, split between WP-mERα and WPHA-mERα (left) and WP-hERα and WP-HA-hERα (right) compared to WP. Mantel-Cox: WP (n = 11, 214 days) versus WP-mERα (n = 20, 228 days, p = 0.0053); WP-HA-mERα (n = 20, 233 days, p = 0.0008); WP-hERα (n = 20, 240 days, p = 0.0029); WP-HA-hERα (n = 20, 215 days, p = 0.1610). *** p < 0.001, ** p < 0.01, ns p > 0.05. b Distribution of histological subtypes of tumors derived from the different models (left) and the luciferase positive tumors that are either ERα-positive or ERα-negative (right). The tumors are either >90% sarcomatoid, >90% carcinomatoid or consists of both histological subtypes (mixed). The absolute number of tumors analyzed are indicated. c Representative images of bioluminescence imaging of luciferase expression in cell lines derived from WP-mERα, WP-hERα and WP-HA-hERα luciferase-negative tumors, treated with DMSO or 5-Aza-2’-deoxycytidine. d Kaplan-Meier analysis of mammary tumor-specific survival of WP, WP-HA-mERα and WP-HA-hERα mice treated with 17β-estradiol slow-release pellets (E2; left) and with vehicle (veh; right). Mantel-Cox: E2: WP (n = 11, 165 days) versus WP-HA-mERα (n = 10, 200.5 days, p = 0.0090); WP-HA-hERα (n = 11, 190 days, p = 0.0671). Veh: WP (n = 9, 224 days) versus WP-HA-mERα (n = 9, 244 days, p = 0.0324); WP-HA-hERα (n = 9, 243 days, p = 0.0067). ** p < 0.01, * p < 0.05, ns p > 0.05. e Distribution of all palpable tumors, identified in vehicle- and E2-treated mice, separated in luciferase signal being negative, weakly positive and positive. WP-HA-mERα veh, 18 tumors; WP-HA-mERα E2, 10 tumors; WP-HA-hERα veh, 14 tumors; WP-HA-hERα E2, 10 tumors. (PNG 719 kb)
Supplementary Fig. S3
The FOXA1 motif is absent at ERα binding sites in mouse cells, while present in 820 human cells. Expression levels of mouse Esr1 and human ESR1 transcripts in primary MMECs derived from AdCre-induced recominbed Trp53F/F, Trp53F/F;HA-mERα and Trp53F/F;HA-hERα primary MMECs upon DMSO or E2 stimulation, derived from RNA-seq data. b Heatmap illustrating raw peak intensity of HA-tag ChIP-seq in primary MMECs derived from Trp53F/F, Trp53F/F;HA-mERα and Trp53F/F;HA-hERα mice upon AdCre induced recombination, separated in sites shared by both HA-mERα and HA-hERα, sites specific for HA-hERα and sites specific for HA-mERα. A window of 5kb around the peak is shown. c Average read count profiles of the HA-tag ChIP-seq peak signal at shared sites, HA-hERα specific sites and HA-mERα specific sites. d Spider plot representing the differential enrichment of the fraction of DNA motifs identified in ERα binding sites (present in at least 10% of binding sites) of human MCF7 and mouse HA-mERα and HA-hERα MMECs (as shown in Fig. 4b.) compared to publicly available datasets. ERα binding motifs in human MCF7 are compared to motifs identified in human breast tumors (GSE40867), human endometrial tumors (GSE94524), and human healthy breast tissue (GSE99680) (left). ERα binding motifs in mouse HA-mERα and HA-hERα MMECs are compared to motifs identified in mouse mammary gland (GSE43415) and E2 stimulated mouse uterus (GSE36455) (right). e mRNA expression levels of ERα and the transcription factors FOXA1 and GATA3, that are expressed relatively high in MCF7 and low in primary MMECs, derived from RNA-seq data. f mRNA expression levels of three additional co-factors; SRC1, SRC2 and SRC3. SRC3 also shows relatively high expression in MCF7 compared to MMECs. Data derived from RNA-seq data. (PNG 2612 kb)
Supplementary Fig. S4
Expression of human ERα in combination with additional transcription factors does not sensitize MMECs to E2 stimulation. a RT-qPCR analysis using primers specific for human ERα, Flag-tagged SRC1, Flag-tagged SRC2, SRC3 and FOXA1 and primers recognizing both mouse and human GATA3 in MMECs transduced with lentiviral constructs containing the indicated cDNAs. Data represent mean + SD, n = 3. b RT-qPCR analysis using primers specific for mouse Areg (left) and mouse Ccnd1 (right) in MMECs transduced with lentiviral constructs containing the indicated cDNAs after stimulation with DMSO, E2 or ICI. Data represent mean + SD, n = 2. Two-way ANOVA: Areg: Empty vector DMSO versus ICI, p = 0.0052; hERα DMSO versus E2, p = 0.0156; hERα/hSRC1/2/3/hFOXA1 DMSO versus ICI, p = 0.0264; hERα/hSRC1/2/3/hFOXA1/HA-hGATA3 DMSO versus ICI, p = 0.0065. Ccnd1: hERα DMSO versus E2, p = 0.0025; hERα/hSRC1/2/3/hFOXA1/HA-hGATA3 DMSO versus ICI, p = 0.0103. ** p < 0.01, * p < 0.05, ns p > 0.05. c,d Representative images (c) and quantification (d) of clonogenic assays of MMECs transduced with lentiviral constructs containing the indicated cDNAs and treated with DMSO, E2 or ICI. Data represent mean + SD, n = 2. Two-way ANOVA: hERα DMSO versus E2, p = 0.0001; hERα/hSRC1/2/3 DMSO versus E2, p = 0.0114; Empty vector DMSO versus hERα/hSRC1/2/3 DMSO, p = 0.0020; Empty vector DMSO versus hERα/hSRC1/2/3/hFOXA1 DMSO, p = 0.0001; Empty vector DMSO versus hERα/hSRC1/2/3/hFOXA1/HA-hGATA3 DMSO, p = 0.0001. *** p < 0.001, ** p < 0.01, * p < 0.05, ns p > 0.05. (PNG 892 kb)
Supplementary Table S1
Primer sequences. (DOCX 21 kb)
Supplementary Table S2
Antibodies. (DOCX 16 kb)
Rights and permissions
About this article
Cite this article
Cornelissen, L.M., Henneman, L., Drenth, A.P. et al. Exogenous ERα Expression in the Mammary Epithelium Decreases Over Time and Does Not Contribute to p53-Deficient Mammary Tumor Formation in Mice. J Mammary Gland Biol Neoplasia 24, 305–321 (2019). https://doi.org/10.1007/s10911-019-09437-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10911-019-09437-z