Abstract
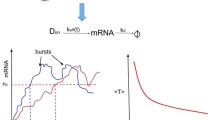
Transcription bursting creates variation among the individuals of a given population. Bursting emerges as the consequence of turning on and off the transcription process randomly. There are at least three sub-processes involved in the bursting phenomenon with different timescale regimes viz. flipping across the on–off state channels, microscopic transcription elongation events and the mesoscopic transcription dynamics along with the mRNA recycling. We demonstrate that when the flipping dynamics is coupled with the microscopic elongation events, then the distribution of the resultant transcription rates will be over-dispersed. This in turn reflects as the transcription bursting with over-dispersed non-Poisson type distribution of mRNA numbers. We further show that there exist optimum flipping rates (αC, βC) at which the stationary state Fano factor and variance associated with the mRNA numbers attain maxima. These optimum points are connected via \( \alpha_{C} = \sqrt {\beta_{C} \left( {\beta_{C} + \gamma_{r} } \right)} \). Here α is the rate of flipping from the on-state to the off-state, β is the rate of flipping from the off-state to the on-state and γr is the decay rate of mRNA. When α = β = χ with zero rate in the off-state channel, then there exist optimum flipping rates at which the non-stationary Fano factor and variance attain maxima. Here \( \chi_{C,v} \simeq {{3k_{r}^{ + } } \mathord{\left/ {\vphantom {{3k_{r}^{ + } } {2\left( {1 + k_{r}^{ + } t} \right)}}} \right. \kern-0pt} {2\left( {1 + k_{r}^{ + } t} \right)}} \) (here \( k_{r}^{ + } \) is the rate of transcription purely through the on-state elongation channel) is the optimum flipping rate at which the variance of mRNA attains a maximum and \( \chi_{C,\kappa } \simeq {{1.72} \mathord{\left/ {\vphantom {{1.72} t}} \right. \kern-0pt} t} \) is the optimum flipping rate at which the Fano factor attains a maximum. Close observation of the transcription mechanism reveals that the RNA polymerase performs several rounds of stall-continue type dynamics before generating a complete mRNA. Based on this observation, we model the transcription event as a stochastic trajectory of the transcription machinery across these on–off state elongation channels. Each mRNA transcript follows different trajectory. The total time taken by a given trajectory is the first passage time (FPT). Inverse of this FPT is the resultant transcription rate associated with the particular mRNA. Therefore, the time required to generate a given mRNA transcript will be a random variable. For a stall-continue type dynamics of RNA polymerase, we show that the overall average transcription rate can be expressed as \( k_{r} \simeq h_{\infty }^{ + } k_{r}^{ + } \) where \( k_{r}^{ + } \simeq {{\lambda_{r}^{ + } } \mathord{\left/ {\vphantom {{\lambda_{r}^{ + } } L}} \right. \kern-0pt} L} \), λ+r is the microscopic transcription elongation rate in the on-state channel and L is the length of a complete mRNA transcript and h+∞ = [β/(α + β)] is the stationary state probability of finding the transcription machinery in the on-state channel.












Similar content being viewed by others
Abbreviations
- MFPT:
-
Mean first passage time. It is the average time required to generate a full mRNA transcript with a given elongation speed of RNA polymerase (s)
- FPT (T):
-
First passage time. It is the time required to generate a given mRNA transcript. Clearly this quantity will vary from transcript to transcript. This is denoted by the random variable T (s)
- m :
-
Concentration of the mRNA molecules or the number of mRNA molecules (M, or number)
- RNAP:
-
RNA polymerase complex
- Fano factor:
-
Variance/mean (units of mean)
- CV:
-
Coefficient of variation = standard deviation/mean (dimensionless)
- k r :
-
Overall average or effective transcription rate (M/s or 1/s)
- k + r :
-
Transcription rate in the on-state channel of transcription (M/s or 1/s)
- k − r :
-
Transcription rate in the off-state channel of transcription (M/s or 1/s)
- L :
-
Length of the complete mRNA transcript (bp)
- λ + r :
-
Microscopic transcription elongation rate constant associated with the addition of single bp with already emerging mRNA in the on-state channel, k+r = λ+r /L (bp/s)
- λ − r :
-
Microscopic transcription elongation rate constant associated with the addition of single bp with already emerging mRNA in the off-state channel, k−r = λ−r /L (bp/s)
- γ r :
-
Decay rate constant associated with the mRNA molecules. By definition 1/γr is the lifetime of mRNA (1/s)
- α :
-
Rate of flipping from the on-state to off-state channel of the transcription (1/s)
- β :
-
Rate of flipping from the off-state to the on-state channel of the transcription (1/s)
- χ :
-
When α = β then α = β = χ is the rate of flipping across on–off states of transcription (1/s)
- χ C,v :
-
Optimum on–off flipping rate at which the variance associated with the mRNA number fluctuations attains a maximum. When k−r = 0 and γr = 0, then one finds that χC,v ≃ 3k+r /2(1 + k+r t) (1/s)
- χ C,κ :
-
Optimum on–off flipping rate at which the Fano factor associated with the mRNA number fluctuations attains a maximum. When k−r = 0 and γr = 0, then one finds that χC,κ ≃ 1.72/t (1/s)
- σ :
-
kr/α, is the transcription efficiency or the burst size (molecules)
- σ C :
-
kr/αC, is the optimum transcription efficiency or the burst size at which the steady state Fano factor attains a maximum (molecules or dimensionless number)
- η m,t :
-
Mean value associated with the mRNA number fluctuations at time t (M or number)
- μ m,t :
-
Coefficient of variation (= variance/square of mean) associated with the mRNA number fluctuations at time t, μm,t = vm,t/η2m,t (dimensionless)
- κ m,t :
-
Fano factor (= variance/mean) associated with the mRNA number fluctuations at time t, κm,t = vm,t/ηm,t (M or number)
- v m,t :
-
Variance associated with the mRNA fluctuations at time t (M2 or number)
- η m,∞ :
-
Mean associated with the mRNA number fluctuations at the steady state (M or number)
- μ m,∞ :
-
Coefficient of variation associated with the mRNA number fluctuations at the steady state, μm,∞ = vm,∞/η2m,∞ (dimensionless)
- κ m,∞ :
-
Fano factor associated with the mRNA number fluctuations at the steady state. κm,∞ = vm,∞/ηm,∞ (M or number)
- v m,∞ :
-
Steady state variance associated with the mRNA numbers (M2 or number)
- α C :
-
When β is fixed, then the optimum value of α at which the stationary state Fano factor attains a maximum. This can be obtained by solving [∂κm,∞/∂α] = 0 for α. Explicitly, when k−r = 0 then one finds that \( \alpha_{C} = \sqrt {\beta \left( {\beta + \gamma_{r} } \right)} \) (1/s)
- β C :
-
When α is fixed, then iteration over β shows an optimum point βC at which the steady state Fano factor associated with the mRNA number fluctuations attain a maximum. Explicitly one finds that \( \beta_{C} = \sqrt {\alpha^{2} + {{\gamma_{r}^{2} } \mathord{\left/ {\vphantom {{\gamma_{r}^{2} } 4}} \right. \kern-0pt} 4}} - {{\gamma_{r} } \mathord{\left/ {\vphantom {{\gamma_{r} } 2}} \right. \kern-0pt} 2} \). Both these optimum flipping rates αC, βC are connected via \( \alpha_{C} = \sqrt {\beta_{C} \left( {\beta_{C} + \gamma_{r} } \right)} \) (1/s)
- p ±m,t :
-
Probability of finding m number of mRNAs at time t in the respective on (+) and off (−) channels of transcription (dimensionless)
- p ±m,∞ :
-
Probability of finding m number of mRNAs in the respective on (+) and off (−) channels of transcription at the steady state (dimensionless)
- p m,t :
-
p+m,t + p−m,t , Total probability of finding m number of mRNA molecules at time t (dimensionless)
- p m,∞ :
-
p+m,∞ + p−m,∞ , total stationary state probability of finding m number of mRNA molecules at time t (dimensionless)
- T ± n :
-
Mean first passage times associated with the generation of a complete mRNA transcript starting from n number of bases to n = L bp over on–off state channels Tn = T+n + T−n (s)
- T 0 :
-
Mean first passage time associated with the generation of a complete mRNA transcript starting from n = 0 number of bases to n = L bp. By definition the overall transcription rate is kr = 1/T0 (s)
- ξ :
-
Transcription rate associated with an individual mRNA synthesis. Clearly, ξ is a random variable. It is the inverse of the FPT required to generate a complete mRNA transcript for a given elongation speed of RNA polymerase. Its ensemble average kr = 〈ξ〉 is the overall average transcription rate (M/s or 1/s)
- χ C,T :
-
On–off flipping rate at which the Fano factor of FPTs associated with the generation of a complete mRNA transcript attains the maximum point (1/s)
- q ±n,t :
-
Probability of finding n number of bases in the elongating mRNA at time t in the respective on (+) or off (−) channel of transcription (dimensionless)
- G ±s,t :
-
∑∞m=0 smp±m,t , is the generating function associated with the probability density p±m,t (dimensionless)
- R 0 :
-
Second moment of the distribution of FPTs required to generate a complete mRNA transcript starting from n = 0 (s2)
- v T :
-
R0 − T20 , variance associated with the distribution of FPTs required to generate a complete mRNA transcript starting from n = 0 number of bases (s2)
- κ T :
-
vT/ηT, Fano factor associated with the distribution of FPTs required to generate a complete mRNA transcript starting from n = 0 number of bases (s)
- μ T :
-
vT/η2T , coefficient of variation associated with the distribution of FPTs required to generate a complete mRNA transcript starting from n = 0 number bases (dimensionless)
- η T :
-
T0, mean associated with the distribution of FPTs required to generate a complete mRNA transcript starting from n = 0 (s)
- α S , β S :
-
Critical values of the flipping rates (α, β) such that when α < αS or β < βS, then the Fano factor κT associated with the distribution of FPTs becomes as κT < 1 (sub-Poisson type). When α > αS or β > βS, then κT > 1 (super-Poisson type distribution). When α = αS or β = βS, then κT = 1 as described in Eq. (42) (1/s)
- w(T)dT :
-
Distribution function of FPTs. When χ → 0 and χ → ∞, as well as for the case where α ≠ β, one can approximate this as, \( w\left( T \right) \simeq {{\exp \left( { - {{\left( {T - \mu_{T} } \right)^{2} } \mathord{\left/ {\vphantom {{\left( {T - \mu_{T} } \right)^{2} } {2v_{T} }}} \right. \kern-0pt} {2v_{T} }}} \right)} \mathord{\left/ {\vphantom {{\exp \left( { - {{\left( {T - \mu_{T} } \right)^{2} } \mathord{\left/ {\vphantom {{\left( {T - \mu_{T} } \right)^{2} } {2v_{T} }}} \right. \kern-0pt} {2v_{T} }}} \right)} {\sqrt {2\pi v_{T} } }}} \right. \kern-0pt} {\sqrt {2\pi v_{T} } }} \) (dimensionless)
References
B. Alberts, Molecular Biology of the Cell (Garland Science, New York, 2002)
B. Lewin, J.E. Krebs, S.T. Kilpatrick, E.S. Goldstein, B. Lewin, Lewin’s Genes X (Jones and Bartlett, Sudbury, 2011)
F. Kepes, Periodic transcriptional organization of the E. coli genome. J. Mol. Biol. 340(5), 957–964 (2004). https://doi.org/10.1016/j.jmb.2004.05.039
T.E. Kuhlman, E.C. Cox, Gene location and DNA density determine transcription factor distributions in Escherichia coli. Mol. Syst. Biol. 8, 610 (2012). https://doi.org/10.1038/msb.2012.42
W.J. Blake, M. Ka, C.R. Cantor, J.J. Collins, Noise in eukaryotic gene expression. Nature 422(6932), 633–637 (2003). https://doi.org/10.1038/nature01546
M.C. Mackey, M. Santillán, M. Tyran-Kamińska, E.S. Zeron, The utility of simple mathematical models in understanding gene regulatory dynamics. Silico Biol. 12, 23–53 (2012). https://doi.org/10.3233/ISB-140463
E.M. Ozbudak, M. Thattai, I. Kurtser, A.D. Grossman, A. van Oudenaarden, Regulation of noise in the expression of a single gene. Nat. Genet. 31(1), 69–73 (2002). https://doi.org/10.1038/ng869. PubMed PMID: 11967532
H.B. Fraser, A.E. Hirsh, G. Giaever, J. Kumm, M.B. Eisen, Noise minimization in eukaryotic gene expression. PLoS Biol. 2(6), e137 (2004). https://doi.org/10.1371/journal.pbio.0020137
A. Becskei, B.B. Kaufmann, A. van Oudenaarden, Contributions of low molecule number and chromosomal positioning to stochastic gene expression. Nat. Genet. 37(9), 937–944 (2005). https://doi.org/10.1038/ng1616
A. Raj, A. van Oudenaarden, Nature, nurture, or chance: stochastic gene expression and its consequences. Cell 135(2), 216–226 (2008). https://doi.org/10.1016/j.cell.2008.09.050
M.B. Elowitz, A.J. Levine, E.D. Siggia, P.S. Swain, Stochastic gene expression in a single cell. Science 297(5584), 1183–1186 (2002). https://doi.org/10.1126/science.1070919
N. Mitarai, I.B. Dodd, M.T. Crooks, K. Sneppen, The generation of promoter-mediated transcriptional noise in bacteria. PLoS Comput. Biol. 4(7), e1000109 (2008). https://doi.org/10.1371/journal.pcbi.1000109
H.H. McAdams, A. Arkin, Stochastic mechanisms in gene expression. Proc. Natl. Acad. Sci. U.S.A. 94(3), 814–819 (1997). https://doi.org/10.1073/pnas.94.3.814
J. Paulsson, Summing up the noise in gene networks. Nature 427(6973), 415–418 (2004). https://doi.org/10.1038/nature02257
I. Golding, J. Paulsson, S.M. Zawilski, E.C. Cox, Real-time kinetics of gene activity in individual bacteria. Cell 123(6), 1025–1036 (2005). https://doi.org/10.1016/j.cell.2005.09.031
A.M. Corrigan, E. Tunnacliffe, D. Cannon, J.R. Chubb, A continuum model of transcriptional bursting. Elife (2016). https://doi.org/10.7554/eLife.13051
S. Chong, C. Chen, H. Ge, X.S. Xie, Mechanism of transcriptional bursting in bacteria. Cell 158(2), 314–326 (2014). https://doi.org/10.1016/j.cell.2014.05.038
V. Shahrezaei, P.S. Swain, Analytical distributions for stochastic gene expression. Proc. Natl. Acad. Sci. U.S.A. 105(45), 17256–17261 (2008). https://doi.org/10.1073/pnas.0803850105
J. Yu, J. Xiao, X. Ren, K. Lao, X.S. Xie, Probing gene expression in live cells, one protein molecule at a time. Science 311(5767), 1600–1603 (2006). https://doi.org/10.1126/science.1119623
N. Kumar, A. Singh, R.V. Kulkarni, Transcriptional bursting in gene expression: analytical results for general stochastic models. PLoS Comput. Biol. 11(10), e1004292 (2015). https://doi.org/10.1371/journal.pcbi.1004292
V. Shahrezaei, J.F. Ollivier, P.S. Swain, Colored extrinsic fluctuations and stochastic gene expression. Mol. Syst. Biol. 4, 196 (2008). https://doi.org/10.1038/msb.2008.31
A. Raj, C.S. Peskin, D. Tranchina, D.Y. Vargas, S. Tyagi, Stochastic mRNA synthesis in mammalian cells. PLoS Biol. 4(10), e309 (2006). https://doi.org/10.1371/journal.pbio.0040309
A. Klindziuk, A.B. Kolomeisky, Theoretical investigation of transcriptional bursting: a multistate approach. J. Phys. Chem. B 122(50), 11969–11977 (2018). https://doi.org/10.1021/acs.jpcb.8b09676
L.H. So, A. Ghosh, C. Zong, L.A. Sepulveda, R. Segev, I. Golding, General properties of transcriptional time series in Escherichia coli. Nat. Genet. 43(6), 554–560 (2011). https://doi.org/10.1038/ng.821
K. Fujita, M. Iwaki, T. Yanagida, Transcriptional bursting is intrinsically caused by interplay between RNA polymerases on DNA. Nat. Commun. 7, 13788 (2016). https://doi.org/10.1038/ncomms13788
O. Pulkkinen, R. Metzler, Distance matters: the impact of gene proximity in bacterial gene regulation. Phys. Rev. Lett. 110(19), 198101 (2013). https://doi.org/10.1103/PhysRevLett.110.198101
D. Levens, D.R. Larson, A new twist on transcriptional bursting. Cell 158(2), 241–242 (2014). https://doi.org/10.1016/j.cell.2014.06.042
T. Fukaya, B. Lim, M. Levine, Enhancer control of transcriptional bursting. Cell 166(2), 358–368 (2016). https://doi.org/10.1016/j.cell.2016.05.025
E. Tunnacliffe, A.M. Corrigan, J.R. Chubb, Promoter-mediated diversification of transcriptional bursting dynamics following gene duplication. Proc. Natl. Acad. Sci. U.S.A. 115(33), 8364–8369 (2018). https://doi.org/10.1073/pnas.1800943115
J. Ma, M.D. Wang, DNA supercoiling during transcription. Biophys. Rev. 8(Suppl 1), 75–87 (2016). https://doi.org/10.1007/s12551-016-0215-9
C.A. Brackley, J. Johnson, A. Bentivoglio, S. Corless, N. Gilbert, G. Gonnella et al., Stochastic model of supercoiling-dependent transcription. Phys. Rev. Lett. 117(1), 018101 (2016). https://doi.org/10.1103/PhysRevLett.117.018101
N.S. Gerasimova, N.A. Pestov, O.I. Kulaeva, D.J. Clark, V.M. Studitsky, Transcription-induced DNA supercoiling: new roles of intranucleosomal DNA loops in DNA repair and transcription. Transcription 7(3), 91–95 (2016). https://doi.org/10.1080/21541264.2016.1182240
Kim S, Beltran B, Irnov I, Jacobs-Wagner C. Long-Distance Cooperative and Antagonistic RNA Polymerase Dynamics via DNA Supercoiling. Cell. 2019;179(1):106-19 e16. https://doi.org/10.1016/j.cell.2019.08.033. PubMed PMID: 31539491
K. Tantale, F. Mueller, A. Kozulic-Pirher, A. Lesne, J.M. Victor, M.C. Robert et al., A single-molecule view of transcription reveals convoys of RNA polymerases and multi-scale bursting. Nat. Commun. 7, 12248 (2016). https://doi.org/10.1038/ncomms12248
T. Heberling, L. Davis, J. Gedeon, C. Morgan, T. Gedeon, A mechanistic model for cooperative behavior of co-transcribing RNA polymerases. PLoS Comput. Biol. 12(8), e1005069 (2016). https://doi.org/10.1371/journal.pcbi.1005069
I. Jonkers, J.T. Lis, Getting up to speed with transcription elongation by RNA polymerase II. Nat. Rev. Mol. Cell Biol. 16(3), 167–177 (2015). https://doi.org/10.1038/nrm3953
P. Maiuri, A. Knezevich, A. De Marco, D. Mazza, A. Kula, J.G. McNally et al., Fast transcription rates of RNA polymerase II in human cells. EMBO Rep. 12(12), 1280–1285 (2011). https://doi.org/10.1038/embor.2011.196
S. Smith, R. Grima, Single-cell variability in multicellular organisms. Nat. Commun. 9(1), 345 (2018). https://doi.org/10.1038/s41467-017-02710-x
J. Peccoud, B. Ycart, Markovian modeling of gene-product synthesis. Theor. Popul. Biol. 48(2), 222–234 (1995). https://doi.org/10.1006/tpbi.1995.1027
S. Iyer-Biswas, F. Hayot, C. Jayaprakash, Stochasticity of gene products from transcriptional pulsing. Phys. Rev. E Stat. Nonlinear Soft Matter Phys. 79(3 Pt 1), 031911 (2009). https://doi.org/10.1103/PhysRevE.79.031911
C.W. Gardiner, Handbook of Stochastic Methods for Physics, Chemistry, and the Natural Sciences (Springer, Berlin, 1985)
Kampen NGv. Stochastic Processes in Physics and Chemistry. Amsterdam; New York; New York: North-Holland; Sole Distributors for the USA and Canada. North-Holland: Elsevier; 1981
H. Risken, The Fokker–Planck Equation: Methods of Solution and Applications (Springer, Berlin, 1989)
E.T. Whittaker, G.N. Watson, A Course of Modern Analysis (University Press, Cambridge, 1969)
M. Abramowitz, I.A. Stegun, Handbook of Mathematical Functions, with Formulas, Graphs, and Mathematical Tables (Dover, New York, 1965)
D.T. Gillespie, Stochastic simulation of chemical kinetics. Annu. Rev. Phys. Chem. 58, 35–55 (2007). https://doi.org/10.1146/annurev.physchem.58.032806.104637
D.T. Gillespie, Exact stochastic simulation of coupled chemical reactions. J. Phys. Chem. 81(25), 2340–2361 (1977). https://doi.org/10.1021/j100540a008
R. Murugan, Stochastic transcription initiation: time dependent transcription rates. Biophys. Chem. 121(1), 51–56 (2006). https://doi.org/10.1016/j.bpc.2005.12.010
D.S. Grebenkov, R. Metzler, G. Oshanin, Strong defocusing of molecular reaction times results from an interplay of geometry and reaction control. Commun. Chem. 1(1), 96 (2018). https://doi.org/10.1038/s42004-018-0096-x
M. Bauer, E.S. Rasmussen, M.A. Lomholt, R. Metzler, Real sequence effects on the search dynamics of transcription factors on DNA. Sci. Rep. 5, 10072 (2015). https://doi.org/10.1038/srep10072
A. Godec, R. Metzler, Universal proximity effect in target search kinetics in the few-encounter limit. Phys. Rev. X 6(4), 041037 (2016). https://doi.org/10.1103/PhysRevX.6.041037
Y. Wang, T. Ni, W. Wang, F. Liu, Gene transcription in bursting: a unified mode for realizing accuracy and stochasticity. Biol. Rev. Camb. Philos. Soc. (2018). https://doi.org/10.1111/brv.12452
B. Munsky, G. Neuert, A. van Oudenaarden, Using gene expression noise to understand gene regulation. Science 336(6078), 183–187 (2012). https://doi.org/10.1126/science.1216379
M. Dobrzynski, F.J. Bruggeman, Elongation dynamics shape bursty transcription and translation. Proc. Natl. Acad. Sci. U.S.A. 106(8), 2583–2588 (2009). https://doi.org/10.1073/pnas.0803507106
R. Zwanzig, Rate processes with dynamical disorder. Acc. Chem. Res. 23(5), 148–152 (1990). https://doi.org/10.1021/ar00173a005
R. Zwanzig, Dynamical disorder: passage through a fluctuating bottleneck. J. Chem. Phys. 97(5), 3587–3589 (1992). https://doi.org/10.1063/1.462993
Acknowledgements
This work was supported by DST SERB grants CRG/2019/001208 and MTR/2019/00002.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendices
Appendix 1
Consider the following backward type coupled master equations along with the boundary conditions (Eq. 39 along with Eq. 37 of the main text) corresponding to the second moments of the first passage times (FPTs) required to generate a complete mRNA transcript starting from an arbitrary n size of mRNA [41, 43].
In this equation, the first moments of the FPTs are defined as follows (as described in Eq. 37 of the main text).
The set of difference Eq. (80) is exactly solvable using standard summing technique [41] and the final expression corresponding to the variance (vT), Fano factor (κT) and coefficient of variation (μT) of FPTs associated with the generation of a complete mRNA transcript of length L starting from n = 0 can be written as follows.
Here y and W are defined as in Eq. (81) and various other terms A1, A2 and A3 in Eq. (82) are defined as follows.
In Eqs. (83)–(85), W is defined as in Eq. (81). When α = β = χ, then the expression corresponding to the variance of FPTs given in Eq. (82) simplifies to the following form.
The terms Y, B1, B2 and B3 in Eq. (86) are defined as follows.
Numerical analysis of Eq. (86) reveals the existence of maximum vT, κT and μT at the optimum flipping rate χC,T which is demonstrated in Fig. 8b, d. Equation (81) clearly suggest that when the microscopic transcription elongation rate associated with the off-state channel is close to zero i.e. λ−r → 0 then the overall average transcription rate scales with the on–off flipping rates as kr = [1/T0] ≃ k+r [β/(α + β)] where k+r = λ+r /L is the overall transcription rate of the pure on-state channel and λ+r is the elongation rate of the on-state channel.
Appendix 2
Let us consider the uncoupled on–off state transcription elongation channels. When there is no flipping across the on and off state channels, then Eq. (1) will be uncoupled as follows.
Here u±m,t is the probability of finding m number of mRNAs at time t in the respective independent transcription channels. Upon defining the generating functions as G±s,t = ∑∞m=0 smu±m,t one can transform Eq. (89) into the following partial differential equations.
Upon solving Eq. (89) for the for the appropriate boundary conditions one finds the expression for the generating functions as follows.
Using this generating function one can derive various statistical properties of the mRNA number fluctuations corresponding to on–off channels as follows.
Upon expanding the generating function into a Macularin series with respect to variable s and then setting s = 1, one finally obtains the probability density functions as follows.
Using Eq. (92) one can directly obtain the steady state probability density functions as follows.
When the on–off state flipping is uncoupled from the transcription elongation, then the flipping dynamics can be described by the following set of coupled differential equations.
Here we have the normalization condition h+t + h−t = 1. Upon solving Eq. (94) with the appropriate initial conditions, one obtains the expression for the time dependent probability h±t of finding the transcription state in the respective on–off channels as follows.
Upon combining Eq. (92) with Eq. (95), one can derive p±m,t ≃ u±m,t h±t , which is the probability of finding m number of mRNAs in the respective on–off state channels. Explicitly one can write the following expression.
In this equation, we have defined the function φ±t = (k±r /γr)[1 − exp (− γrt)]. When the timescale associated with the on–off state flipping dynamics is much lower than the timescale associated with the mRNA synthesis and decay dynamics so that (α + β) ≫ γr, then Eq. 96 reduces to pm,t ≃ u+m,t h+∞ + u−m,t h−∞ . Under complete steady state conditions pm,∞ ≃ u+m,∞ h+∞ + u−m,∞ h−∞ and one obtains the following bimodal Poisson type expression.
When the transcription rate associated with the off-state channel tends toward zero, then one recovers the Poisson density function with zero spike [17] as follows.
Rights and permissions
About this article
Cite this article
Murugan, R. Theory of transcription bursting: stochasticity in the transcription rates. J Math Chem 58, 2140–2187 (2020). https://doi.org/10.1007/s10910-020-01166-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10910-020-01166-7