Abstract
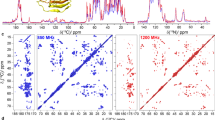
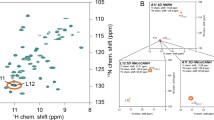
A procedure is presented for the substantial simplification of 2D constant-time 13C−1H heteronuclear single-quantum correlation (HSQC) spectra of 13C-enriched proteins. In this approach, a single pulse sequence simultaneously records eight sub-spectra wherein the phases of the NMR signals depend on spin topology. Signals from different chemical groups are then stratified into different sub-spectra through linear combination based on Hadamard encoding of 13CHn multiplicity (n = 1, 2, and 3) and the chemical nature of neighboring 13C nuclei (aliphatic, carbonyl/carboxyl, aromatic). This results in five sets of 2D NMR spectra containing mutually exclusive signals from: (i) 13Cβ−1Hβ correlations of asparagine and aspartic acid, 13Cγ−1Hγ correlations of glutamine and glutamic acid, and 13Cα−1Hα correlations of glycine, (ii) 13Cα−1Hα correlations of all residues but glycine, and (iii) 13Cβ−1Hβ correlations of phenylalanine, tyrosine, histidine, and tryptophan, and the remaining (iv) aliphatic 13CH2 and (v) aliphatic 13CH/13CH3 resonances. As HSQC is a common element of many NMR experiments, the spectral simplification proposed in this article can be straightforwardly implemented in experiments for resonance assignment and structure determination and should be of widespread utility.







Similar content being viewed by others
References
Bax A, Ikura M, Kay LE, Torchia DA, Tschudin R (1990) Comparison of different modes of two-dimensional reverse-correlation NMR for the study of proteins. J Magn Reson 86:304–318. https://doi.org/10.1016/0022-2364(90)90262-8
Bendall MR, Doddrell DM, Pegg DT (1981) Editing of carbon-13 NMR spectra. 1. A pulse sequence for the generation of subspectra. J Am Chem Soc 103:4603–4605. https://doi.org/10.1021/ja00405a062
Bodenhausen G, Ruben DJ (1980) Natural abundance nitrogen-15 NMR by enhanced heteronuclear spectroscopy. Chem Phys Lett 69:185–189. https://doi.org/10.1016/0009-2614(80)80041-8
Boyer RD, Johnson R, Krishnamurthy K (2003) Compensation of refocusing inefficiency with synchronized inversion sweep (CRISIS) in multiplicity-edited HSQC. J Magn Reson 165:253–259. https://doi.org/10.1016/j.jmr.2003.08.009
Brenner AK, Frøystein NÅ (2014) Using MUSIC and CC(CO)NH for backbone assignment of two medium-sized proteins not fully accessible to standard 3D NMR. Molecules 19:8890–8903. https://doi.org/10.3390/molecules19078890
Brutscher B (2001) Accurate measurement of small spin-spin couplings in partially aligned molecules using a novel J-mismatch compensated spin-state-selection filter. J Magn Reson 151:332–338. https://doi.org/10.1006/jmre.2001.2375
Brutscher B (2004) Combined frequency- and time-domain NMR spectroscopy. Application to fast protein resonance assignment. J Biomol NMR 29:57–64. https://doi.org/10.1023/B:JNMR.0000019501.21697.34
Bystrov VF (1976) Spin-spin coupling and the conformational states of peptide systems. Prog Nucl Magn Reson Spectrosc 10:41–82. https://doi.org/10.1016/0079-6565(76)80001-5
Chen K, Freedberg DI, Keire DA (2015) NMR profiling of biomolecules at natural abundance using 2D 1H–15N and 1H–13C multiplicity-separated (MS) HSQC spectra. J Magn Reson 251:65–70. https://doi.org/10.1016/j.jmr.2014.11.011
Davis DG (1990) Simplification of proton-detected, natural abundance carbon-13 correlation spectra of proteins via multiplet editing. J Magn Reson 90:589–596. https://doi.org/10.1016/0022-2364(90)90067-J
Davis DG (1991) Improved multiplet editing of proton-detected, heteronuclear shift-correlation spectra. J Magn Reson 91:665–672. https://doi.org/10.1016/0022-2364(91)90398-D
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293. https://doi.org/10.1007/BF00197809
Doddrell DM, Pegg DT, Bendall MR (1982) Distortionless enhancement of NMR signals by polarization transfer. J Magn Reson 48:323–327. https://doi.org/10.1016/0022-2364(82)90286-4
Dubey A, Mondal S, Chandra K, Atreya HS (2016) Rapid identification of amino acid types in proteins using phase modulated 2D HN(CACB) and 2D HN(COCACB). J Magn Reson 267:22–29. https://doi.org/10.1016/j.jmr.2016.04.004
Emsley L, Bodenhausen G (1992) Optimization of shaped selective pulses for NMR using a quaternion description of their overall propagators. J Magn Reson 97:135–148. https://doi.org/10.1016/0022-2364(92)90242-Y
Feng W, Rios CB, Montelione GT (1996) Phase labeling of C-H and C-C spin-system topologies: application in PFG-HACANH and PFG-HACA(CO)NH triple-resonance experiments for determining backbone resonance assignments in proteins. J Biomol NMR 8:98–104. https://doi.org/10.1007/BF00198144
Feuerstein S, Plevin MJ, Willbold D, Brutscher B (2012) iHADAMAC: a complementary tool for sequential resonance assignment of globular and highly disordered proteins. J Magn Reson 214:329–334. https://doi.org/10.1016/j.jmr.2011.10.019
Freeman R, Kempsell SP, Levitt MH (1980) Radiofrequency pulse sequences which compensate their own imperfections. J Magn Reson 38:453–479. https://doi.org/10.1016/0022-2364(80)90327-3
Grzesiek S, Bax A (1993) Amino acid type determination in the sequential assignment procedure of uniformly 13C/15N-enriched proteins. J Biomol NMR 3:185–204. https://doi.org/10.1007/BF00178261
Heikkinen S, Toikka MM, Karhunen PT, Kilpeläinen IA (2003) Quantitative 2D HSQC (Q-HSQC) via suppression of J-dependence of polarization transfer in NMR spectroscopy: application to wood lignin. J Am Chem Soc 125:4362–4367. https://doi.org/10.1021/ja029035k
Hoffmann F, Xue M, Schafer LV, Mulder FAA (2018) Narrowing the gap between experimental and computational determination of methyl group dynamics in proteins. Phys Chem Chem Phys 20:24577–24590. https://doi.org/10.1039/C8CP03915A
Hwang T-L, van Zijl PCM, Garwood M (1997) Broadband adiabatic refocusing without phase distortion. J Magn Reson 124:250–254. https://doi.org/10.1006/jmre.1996.1049
Kay LE, Bax A (1989) Separation of NH and NH2 resonances in 1H-detected heteronuclear multiple-quantum correlation spectra. J Magn Reson 84:598–603. https://doi.org/10.1016/0022-2364(89)90125-X
Kessler H, Schmieder P, Kurz M (1989) Implementation of the DEPT sequence in inverse shift correlation; the DEPT-HMQC. J Magn Reson 85:400–405. https://doi.org/10.1016/0022-2364(89)90153-4
Kupče E, Nishida T, Freeman R (2003) Hadamard NMR spectroscopy. Prog Nucl Magn Reson Spectrosc 42:95–122. https://doi.org/10.1016/S0079-6565(03)00022-0
Lescop E, Rasia R, Brutscher B (2008) Hadamard amino-acid-type edited NMR experiment for fast protein resonance assignment. J Am Chem Soc 130:5014–5015. https://doi.org/10.1021/ja800914h
Levitt MH, Freeman R (1979) NMR population inversion using a composite pulse. J Magn Reson 33:473–476. https://doi.org/10.1016/0022-2364(79)90265-8
Marion D, Ikura M, Tschudin R, Bax A (1989) Rapid recording of 2D NMR spectra without phase cycling. Application to the study of hydrogen exchange in proteins. J Magn Reson 85:393–399. https://doi.org/10.1016/0022-2364(89)90152-2
Markley JL, Bax A, Arata Y, Hilbers CW, Kaptein R, Sykes BD, Wright PE, Wüthrich K (1998) Recommendations for the presentation of NMR structures of proteins and nucleic acids. IUPAC-IUBMB-IUPAB inter-union task group on the standardization of data bases of protein and nucleic acid structures determined by NMR spectroscopy. J Biomol NMR 12:1–23. https://doi.org/10.1023/a:1008290618449
Morris GA, Freeman R (1979) Enhancement of nuclear magnetic resonance signals by polarization transfer. J Am Chem Soc 101:760–762. https://doi.org/10.1021/ja00497a058
Mulder FAA (2009) Leucine side-chain conformation and dynamics in proteins from 13C NMR chemical shifts. ChemBioChem 10:1477–1479. https://doi.org/10.1002/cbic.200900086
Nagana Gowda GA (2002) Improved sensitivity and gradient-enhanced multiplicity edited two-dimensional heteronuclear shift correlation technique. Chem Phys Lett 353:49–54. https://doi.org/10.1016/S0009-2614(01)01480-4
Otting G, Wüthrich K (1988) Efficient purging scheme for proton-detected heteronuclear two-dimensional NMR. J Magn Reson 76:569–574. https://doi.org/10.1016/0022-2364(88)90361-7
Pantoja-Uceda D, Santoro J (2008) Amino acid type identification in NMR spectra of proteins via b- and g-carbon edited experiments. J Magn Reson 195:187–195. https://doi.org/10.1016/j.jmr.2008.09.010
Rios CB, Feng W, Tashiro M, Shang Z, Montelione GT (1996) Phase labeling of C-H and C-C spin-system topologies: application in constant-time PFG-CBCA(CO)NH experiments for discriminating amino acid spin-system types. J Biomol NMR 8:345–350. https://doi.org/10.1007/BF00410332
Sakhaii P, Bermel W (2015) A different approach to multiplicity-edited heteronuclear single quantum correlation spectroscopy. J Magn Reson 259:82–86. https://doi.org/10.1016/j.jmr.2015.07.006
Santoro J, King GC (1992) A constant-time 2D overbodenhausen experiment for inverse correlation of isotopically enriched species. J Magn Reson 97:202–207. https://doi.org/10.1016/0022-2364(92)90250-B
Schmidt JM, Rueterjans H (1990) Proton-detected 2D heteronuclear shift correlation via multiple-quantum coherences of the type I2S. J Am Chem Soc 112:1279–1280. https://doi.org/10.1021/ja00159a077
Schubert M, Labudde D, Leitner D, Oschkinat H, Schmieder P (2005) A modified strategy for sequence specific assignment of protein NMR spectra based on amino acid type selective experiments. J Biomol NMR 31:115–128. https://doi.org/10.1007/s10858-004-8263-z
Schubert M, Oschkinat H, Schmieder P (2001a) MUSIC, selective pulses, and tuned delays: amino acid type-selective 1H–15N correlations, II. J Magn Reson 148:61–72. https://doi.org/10.1006/jmre.2000.2222
Schubert M, Oschkinat H, Schmieder P (2001b) MUSIC and aromatic residues: amino acid type-selective 1H–15N correlations, III. J Magn Reson 153:186–192. https://doi.org/10.1006/jmre.2001.2447
Schubert M, Smalla M, Schmieder P, Oschkinat H (1999) MUSIC in triple-resonance experiments: amino acid type-selective 1H–15N correlations. J Magn Reson 141:34–43. https://doi.org/10.1006/jmre.1999.1881
Shaka AJ, Barker PB, Freeman R (1985) Computer-optimized decoupling scheme for wideband applications and low-level operation. J Magn Reson 64:547–552. https://doi.org/10.1016/0022-2364(85)90122-2
Tashiro M, Rios CB, Montelione GT (1995) Classification of amino acid spin systems using PFG HCC(CO)NH-TOCSY with constant-time aliphatic 13C frequency labeling. J Biomol NMR 6:211–216. https://doi.org/10.1007/BF00211785
Tate S, Masui Y, Inagaki F (1991) Application of the DEPT sequence to the separation of 15NH and 15NH2 resonances in 1H-detected 15N single-quantum coherence spectroscopy. J Magn Reson 94:625–630. https://doi.org/10.1016/0022-2364(91)90152-J
van de Ven FJM, Philippens MEP (1992) Optimization of constant-time evolution in multidimensional NMR experiments. J Magn Reson 97:637–644. https://doi.org/10.1016/0022-2364(92)90045-9
Van Melckebeke H, Simorre JP, Brutscher B (2004) Amino acid-type edited NMR experiments for methyl-methyl distance measurement in 13C-labeled proteins. J Am Chem Soc 126:9584–9591. https://doi.org/10.1021/ja0489644
Vuister GW, Bax A (1992) Resolution enhancement and spectral editing of uniformly 13C-enriched proteins by homonuclear broadband 13C decoupling. J Magn Reson 98:428–435. https://doi.org/10.1016/0022-2364(92)90144-V
Walinda E, Morimoto D, Shirakawa M, Sugase K (2017) F1F2-selective NMR spectroscopy. J Biomol NMR 68:41–52. https://doi.org/10.1007/s10858-017-0113-x
Williamson MP (2013) Using chemical shift perturbation to characterise ligand binding. Prog Nucl Magn Reson Spectrosc 73:1–16. https://doi.org/10.1016/j.pnmrs.2013.02.001
Zuiderweg ERP (2002) Mapping protein-protein interactions in solution by NMR spectroscopy. Biochemistry 41:1–7. https://doi.org/10.1021/bi011870b
Zwahlen C, Legault P, Vincent SJF, Greenblatt J, Konrat R, Kay LE (1997) Methods for measurement of intermolecular NOEs by multinuclear NMR Spectroscopy: Application to a bacteriophage l N-peptide/boxB RNA complex. J Am Chem Soc 119:6711–6721. https://doi.org/10.1021/ja970224q
Acknowledgements
This work was performed under the Cooperative Research Programs at the Institute for Protein Research, Osaka University (NMRCR-17-05 and NMRCR-18-05). We thank Dr. Yohei Miyanoiri (Institute for Protein Research, Osaka University) for help with NMR analysis. This work was supported by JSPS KAKENHI (Grant Number JP 19K14677) and the Uehara Memorial Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yoshimura, Y., Mulder, F.A.A. Sensitive and simplified: a combinatorial acquisition of five distinct 2D constant-time 13C−1H NMR protein correlation spectra. J Biomol NMR 74, 695–706 (2020). https://doi.org/10.1007/s10858-020-00341-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-020-00341-x