Abstract
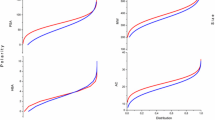
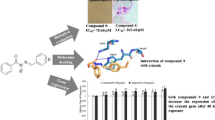
Chagas’s is a neglected tropical disease caused by the protozoan parasite Trypanosoma cruzi. According to the World Health Organization, 7 million people are infected worldwide leading to 7000 deaths per year. Drugs available, nifurtimox and benzimidazole, are limited due to low efficacy and high toxicity. As a validated target, cruzain represents a major front in drug discovery attempts for Chagas disease. Herein, we describe the development of 2D QSAR (\(r_{{{\text{pred}}}}^{2}\) = 0.81) and a 3D-QSAR-based pharmacophore (\(r_{{{\text{pred}}}}^{2}\) = 0.82) from a series of non-covalent cruzain inhibitors represented mostly by oxadiazoles (lead compound, IC50 = 200 nM). Both models allowed us to map key intermolecular interactions in S1′, S2 and S3 cruzain sub-sites (including halogen bond and C‒H/π). To probe the predictive capacity of obtained models, inhibitors available in the literature from different classes displaying a range of scaffolds were evaluate achieving mean absolute deviation of 0.33 and 0.51 for 2D and 3D models, respectively. CoMFA revealed an unexplored region where addition of bulky substituents to produce new compounds in the series could be beneficial to improve biological activity.















Similar content being viewed by others
Abbreviations
- AutoQSAR:
-
Automated QSAR
- BSSE:
-
Basis set superposition error
- B3LYP-D3:
-
Becke, three-parameter hybrid functional combined with Lee–Yang–Parr correlation functional including Grimme’s D3 dispersion scheme
- CoMFA:
-
Comparative molecular field analysis
- GA:
-
Genetic algorithm
- GALAHAD:
-
Genetic algorithm with linear assignment for the hypermolecular alignment of datasets
- HTS:
-
High-throughput screening
- KPLS:
-
Kernel-based partial least square
- LMP2:
-
Localized Møller–Plesset perturbation theory
- MLR:
-
Multiple linear regression
- OPLS3:
-
Optimized potentials for liquid simulations
- PCR:
-
Principal component analysis
- pIC50 :
-
Logarithm of the inverse of compound concentration that reduces the enzyme activity by 50% [log (1/IC50)]
- PLS:
-
Partial least square
- q 2 :
-
Leave-one-out cross-validated correlation coefficient
- QSAR:
-
Quantitative structure–activity relationship
- r 2 :
-
Non-cross-validated correlation coefficient
- RMSE:
-
Root-mean-square error of test set predictions
- SAR:
-
Structure–activity relationship
- SD:
-
Standard deviation
- SDC:
-
Standard deviation coefficient
- SEE:
-
Standard error of estimate
- SEP:
-
Standard error prediction
- X3LYP:
-
Extended hybrid functional combined with Lee–Yang–Parr correlation functional
References
Rassi A Jr, Rassi A, Marin-Neto JA (2010) Chagas disease. Lancet 375:1388–1402
Bern C (2015) Chagas’ disease. N Engl J Med 373:456–466
World Health Organization (2016) Investing to overcome the global impact of neglected tropical diseases: third WHO report on neglected tropical diseases. http://www.who.int/neglected_diseases/9789241564540$4en/. Accessed 9 Oct 2016
Bern C, Montgomery SP (2009) An estimate of the burden of Chagas disease in the United States. Clin Infect Dis 49:e52‒e54
Castro JA, de Mecca MM, Bartel LC (2006) Toxic side effects of drugs used to treat Chagas’ disease (American trypanosomiasis). Hum Exp Toxicol 25:471–479
Campos MC et al (2014) Benznidazole-resistance in Trypanosoma cruzi: evidence that distinct mechanism can act in concert. Mol Biochem Parasitol 193:17–19
Doyle PS et al (2011) The Trypanosoma cruzi protease cruzain mediates immune evasion. PLoS Pathog 7:e1002139
Engel JC et al (1998) Cysteine protease inhibitors alter Golgi complex ultrastructure and function in Trypanosoma cruzi. J Cell Sci 111:597–606
Doyle PS et al (2007) A cysteine protease inhibitor cures Chagas’ disease in an immunodeficient-mouse model of infection. Antimicrob Agents Chemother 51:3932–3939
Martinez-Mayorga K et al (2015) Cruzain inhibitors: efforts made, current leads and a structural outlook of new hits. Drug Discov Today 20:890–899
Ferreira RS et al (2009) Divergent modes of enzyme inhibition in a homologous structure-activity series. J Med Chem 52:5005–5008
Filho JMS et al (2009) Design, synthesis and cruzain docking of 3-(4-substituted-aryl)-1,2,4-oxadiazole-N-acylhydrazones as anti-Trypanosoma cruzi agents. Bioorg Med Chem 17:6682–6691
Ferreira RS et al (2010) Complementarity between a docking and a high-throughput screen in discovering new cruzain inhibitors. J Med Chem 53:4891–4905
Filho JMS et al (2012) Optimization of anti-Trypanosoma cruzi oxadiazoles leads to identification of compounds with efficacy in infected mice. Bioorg Med Chem 20:6423–6433
Patani GA, LaVoie EJ (1996) Bioisosterism: a rational approach in drug design. Chem Rev 96:3147–3176
Kaserer T et al (2015) Pharmacophore models and pharmacophore-based virtual screening: concepts and application exemplified on hydroxysteroid dehydrogenases. Molecules 20:22799–22832
Malvezzi A et al (2009) Pharmacophore model of cruzain inhibitors. QSAR Comb Sci 28:781–784
Cherkasov A et al (2014) QSAR modeling: where have you been? Where are you going to? J Med Chem 57:4977–5010
Scotti M T et al (2016) Variable-selection approaches to generate QSAR models for a set of antichagasic semicarbazones and analogues. Chemometr Intell Lab 154:137–149
Belaidi S et al (2015) Structure activity relationship and quantitative structure-activity relationships modeling of antitrypanosomal activities of alkyldiamine cryptolepine derivatives. J Comput Theor Nanosci 12:2421–2427
Mendez-Lucio O et al (2011) 3D-QSAR studies on purine-carbonitriles as cruzain inhibitors: comparative molecular field analysis (CoMFA) and comparative molecular similarity indices analysis (CoMSIA). MedChemComm 2:1058–1065
Freitas RF, Oprea TI, Montanari CA (2008) 2D QSAR and similarity studies on cruzain inhibitors aimed at improving selectivity over cathepsin L. Bioorg Med Chem 16:838–853
Guido RVC et al (2008) Structure-activity relationships for a class of selective inhibitors of the major cysteine protease from Trypanosoma cruzi. J Enzym Inhib Med Chem 23:964–973
Rodrigues CR et al (2011) CoMFA and HQSAR of acylhydrazide cruzain inhibitors. Bioorg Med Chem Lett 12:1537–1541
Leonard JT, Roy K (2006) On selection of training and test sets for the development of predictive QSAR models. QSAR Comb Sci 25:235–251
Wood DJ, Davis AM (2013) Quantitative structure-activity relationship models that stand the test of time. Mol Pharm 10:1183–1190
Dixon SJ et al (2016) AutoQSAR: an automated machine learning tool for best-practice. QSAR modeling. Future Med Chem 8:1825–1839
Schrödinger Release 2016-2 (2016) Small-Molecule Drug Discovery suite, Schrödinger, LLC, New York
Duan J et al (2010) Analysis and comparison of 2D fingerprints: insights into database screening performance using eight fingerprint methods. J Mol Graph Model 29:157–170
Sastry M et al (2010) Large-scale systematic analysis of 2D fingerprint methods and parameters to improve virtual screening enrichments. J Chem Inf Model 50:771–784
An Y, Sherman W, Dixon SL (2013) Kernel-based partial least squares: application to fingerprint-based QSAR with model visualization. J Chem Inf Model 53:2312–2321
Richmond NJ et al (2006) GALAHAD: 1. Pharmacophore identification by hypermolecular alignment of ligands in 3D. J Comput Aided Mol Des 20:567–587
Yang SY (2010) Pharmacophore modeling and applications in drug discovery: challenges and recent advances. Drug Discov Today 15:444–450
Long W et al (2008) 3D-QSAR studies on a class of IKK-2 inhibitors with GALAHAD used to develop molecular alignment models. QSAR Comb Sci 27:1113–1119
Harder E et al (2015) OPLS3: a force field providing broad coverage of drug-like small molecules and proteins. J Chem Theory Comput 12:281–296
Sastry GM et al (2013) Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J Comput Aided Mol Des 27:221–234
Friesner RA et al (2006) Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J Med Chem 49:6177–6196
Halgren TA et al (2004) Glide: a new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J Med Chem 47:1750–1759
Friesner RA et al (2004) Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J Med Chem 47:1739–1749
Pettersen EF et al (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
SYBYL®-X Suite Molecular Modeling from Sequence through Lead Optimization, version 2.0, Certara USA, Inc
Melo-Filho CC, Braga RC, Andrade CH (2014) 3D-QSAR approaches in drug design: perspectives to generate reliable CoMFA models. Curr Comput Aided Drug Des 10:148–159
Bochevarov AD et al (2013) Jaguar: a high-performance quantum chemistry software program with strengths in life and materials sciences. Int J Quantum Chem 113:2110–2142
Kaminski GA et al (2005) Pseudospectral local second-order Mollet-Plesset methods for computation of hydrogen bonding energies of molecular pairs. J Chem Theory Comput 1:248–254
Walker JD et al (2003) Guidelines for developing and using quantitative structure-activity relationships. Environ Toxicol Chem 22:1653–1665
Desiraju GR et al (2013) Definition of the halogen bond (IUPAC recommendations 2013). Pure Appl Chem 85:1711–1713
Cavallo G et al (2016) The halogen bond. Chem Rev 116:2478–2601
Wilckern R et al (2012) Principles and applications of halogen bonding in medicinal chemistry and chemical biology. J Med Chem 56:1363–1388
Ferreira RS et al (2014) Synthesis, biological evaluation, and structure–activity relationships of potent noncovalent and nonpeptidic cruzain inhibitors as anti-Trypanosoma cruzi agents. J Med Chem 57:2380–2392
Tsuzuki S (2000) Origin of the attraction and directionality of the NH/π interaction: comparison with OH/π and CH/π interactions. J Am Chem Soc 122:11450–11458
Tsuzuki S (2006) Magnitude and directionality of the interaction energy and aliphatic CH/π interaction: significant difference from hydrogen bond. J Phys Chem A 110:10163–10168
Barreiro EJ, Kummerle AE, Fraga CAM (2011) The methylation effect in medicinal chemistry. Chem Rev 11:5215–5246
Wiggers HJ (2013) Non-peptidic cruzain inhibitors with Trypanocidal activity discovered by virtual screening and in vitro assay. PLoS Negl Trop Dis 7:e2370
Carvalho SA et al (2012) Design and synthesis of new (E)-cinnamic N-acylhydrazones as potent antirypanosomal agents. Eur J Med Chem 54:512–521
Trossini GH et al (2010) Cruzain inhibition by hydroxymethylnitrofurazone and nitrofurazone: investigation of a new target in Trypanosoma cruzi. J Enzym Inhib Med Chem 25(1):62–67
Aguilera E et al (2016) Potent and selective inhibitors of Trypanosoma cruzi triosephosphate isomerase with concomitant inhibition of cruzipain: inhibition of parasite growth through multitarget activity. ChemMedChem 11(12):1328–1338
Sassano MF et al (2103) Colloidal aggregation causes inhibition of G protein-coupled receptors. J Med Chem 56(6):2406–2414
Braga SFP et al (2017) Synthesis and biological evaluation of potential inhibitors of the cysteine proteases cruzain and rhodesain designed by molecular simplification. Bioorg Med Chem 25(6):1889–1890
Couto M et al (2015) 2-H-[1,2]Dithiole as a new anti-Trypanosoma cruzi chemotype: biological and mechanism of action studies. Molecules 20(8):14595–14610
Burger, MCM et al (2014) Structures and bioactivities of dihydrochalcones from Metrodorea stipularis. J Nat Prod 77(11):2418–2422
Serafim RAM et al (2017) Molecular modeling and structure-activity relationships studies of bioisoster hybrids on N-acylhydrazone. Med Chem Res 26(4):760–769
Silva-Junior EF et al (2016) Design, synthesis, molecular docking and biological evaluation of thiophen-2-iminothiazolidine derivatives for use against Trypanosoma cruzi. Bioorg Med Chem 24(18):4228–4240
Bellara CL et al (2013) Application of computer-aided drug repurposing in the search of new cruzipain inhibitors: discovery of amiodarone and bromocriptine inhibitory effects. J Chem Inf Model 53(9):2402–2408
Schonherr H, Cernak T (2013) Profound methyl effects in drug discovey and a call for new C–H methylation reactions. Angew Chem Int Ed 52:12256–12267
Funding
Financial support was provided by the State of São Paulo Research Foundation (FAPESP, Fundação de Amparo à Pesquisa do Estado de São Paulo), Grant 2013/07600-3. A.S.S. and M.T.O acknowledge the Coordination for the Improvement of Higher Education Personnel (CAPES, Coordenação de Aperfeiçoamento de Pessoal de Nível Superior) for institutional grants.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
de Souza, A.S., de Oliveira, M.T. & Andricopulo, A.D. Development of a pharmacophore for cruzain using oxadiazoles as virtual molecular probes: quantitative structure–activity relationship studies. J Comput Aided Mol Des 31, 801–816 (2017). https://doi.org/10.1007/s10822-017-0039-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-017-0039-0