Abstract
Fucoxanthin (FX), a primary carotenoid, is associated with the fucoxanthin-chlorophyll a/c binding protein (FCP) complex integrated into the thylakoid membrane (TM) which functions as a light-harvesting complex in the diatom Phaeodactylum tricornutum. Here, we aimed to elucidate the FX production regulated by different light intensities via the correlation of FX biosynthesis and apoproteins composing of FCP complex. High light (HL) accelerated P. tricornutum growth more than low light (LL). The maximum values of FX content and productivity obtained under LL (1.7 mg g−1 and 2.12 mg L−1 day−1, respectively) were substantially higher than those obtained under HL (0.54 mg g−1 and 0.79 mg L−1 day−1, respectively). Notably, proteome and photosynthetic pigment analyses revealed the enrichment of FCP antennae in the LL culture TM fractions but not the HL culture. Semi-quantification of FCP antenna protein using LC–MS/MS and RNA transcriptome analyses revealed that PtLhcf5 and PtLhcf8 played crucial roles in FCP biosynthesis under LL. P. tricornutum cultured under light transition exhibited FCP formation only in the early growth stage to meet the increased photosynthetic activity requirements under LL. Meanwhile, FCP degradation could be triggered by HL throughout the cultivation period. Therefore, FX production was highly correlated with FCP formation, and LL conditions in the early growth stage were critical for higher FX productivity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Marine diatoms of the class Bacillariophyceae are responsible for up to 40% of the global net primary production from phytoplankton in the ocean (Field et al. 1998). Among the thousands of diatom species, Phaeodactylum tricornutum is a well-studied species which was first described as Nitzschia closterium by Bohlin (1897). Phaeodactylum tricornutum has become a robust laboratory model owing to complete whole genome sequencing (Bowler et al. 2008) and its remarkable adaptability. Phaeodactylum tricornutum can deal with stresses or unfavorable environmental conditions such as limited carbon dioxide (Heydarizadeh et al. 2017), nitrogen (Alipanah et al. 2015), and phosphorus (Huang et al. 2019) and light stress (Terry et al. 1983; Domingues et al. 2012). In response to environmental changes, P. tricornutum accumulates different bioactive compounds, including eicosapentaenoic acid, fucoxanthin (FX) (Gao et al. 2017), and triacylglycerides (Remmers et al. 2017).
FX, the most abundant xanthophyll pigment in nature, is primarily found in the chloroplasts of brown seaweeds, such as Undaria, Laminaria, and some microalgae. Notably, in vitro and in vivo experiments have demonstrated that FX exhibits numerous pharmaceutical effects, such as anticancer, anti-diabetic, anti-inflammatory, and antioxidant effects (Bae et al. 2020). For instance, FX and its derivatives can suppress weight gain in diabetic/obese mice (Gille et al. 2019), induce apoptosis in human cervical cancer cell lines (Ye et al. 2014), and alleviate colorectal cancer (Terasaki et al. 2021). Therefore, FX has attracted increasing research and social attention, leading to a higher demand for FX-based health supplements over time.
Conventional FX production using brown seaweeds has numerous disadvantages, including low yield, seasonal cultivation, and high-cost downstream processing. In contrast, microalgae, such as Chaetoceros sp., Isochrysis galbana, Nitzschia sp., Odontella aurita, and P. tricornutum, have more advantages, particularly a high cellular FX content (Petrushkina et al. 2017). The FX content in P. tricornutum ranges from 0.49 to 5.92% of dry biomass (Yang and Wei 2020). Previously Kim et al. (2012) reported a constant extraction yield of FX from P. tricornutum (15.42–16.51 mg g−1) and suggested P. tricornutum as a promising sustainable source for large-scale FX production.
In P. tricornutum the primary carotenoid FX plays a crucial role in harvesting light for photosynthesis, which is an essential physiobiological process for maintaining microalgal growth and development. Therefore, FX has been used as a biomarker to determine the effects of environmental changes on P. tricornutum. For example, FX concentration decreases with decreasing iron concentration (Kosakowska et al. 2004) and during silicon starvation (Zhao et al. 2014), whereas heat-exposed cells increasingly accumulate FX (Feijão et al. 2018).
Several studies have been conducted to enhance the yield of high-value products, including FX in P. tricornutum, based on the known effects of environmental and culture conditions (Gómez-Loredo et al. 2015; Gao et al. 2017; McClure et al. 2018; Conceição et al. 2020). Briefly, these studies modified culture medium components such as nitrogen and silicate; and light quality to optimize the production of FX in P. tricornutum (Gao et al. 2017; McClure et al. 2018; Nur et al. 2018; Wang et al. 2018; Yi et al. 2019). Recently, an approach combining light emitting diodes (LED), nitrogen supplements, and mixotrophic mode to improve biomass and FX production simultaneously, resulted in higher FX productivity (Yang and Wei 2020). Notably, the investigation of appropriate light intensity seems to be a prerequisite for optimizing FX production in P. tricornutum (Nur et al. 2018; Wang et al. 2018; Yi et al. 2019; Yang and Wei 2020).
Light, the critical energy source for photosynthesis, has a profound influence on FX biosynthesis. The effects of light quality and intensity on FX productivity in P. tricornutum have been investigated. Previous findings suggest that the absorption of different light qualities can alter FX production in P. tricornutum (Valle et al. 2014; Yi et al. 2019; Yang and Wei 2020). For example, monochromatic blue light alone and the combination of red and blue light or red, blue, and white light can improve FX production in P. tricornutum (Yi et al. 2019; Yang and Wei 2020). More studies have investigated the effect of white light on FX production, as white light or sunlight is the fundamental photon source for all phototrophic organisms. Recently, the effect of light intensity and other essential factors on P. tricornutum growth and FX productivity were examined (Gómez-Loredo et al. 2015; McClure et al. 2018). Previous findings indicate a similar phenomenon in which FX accumulation is favored by low light (LL) but greatly decreased by high light (HL), consistent with the present results (Gómez-Loredo et al. 2015; McClure et al. 2018; Conceição et al. 2020).
The FX biosynthetic pathway in P. tricornutum was first described by Dambek et al. (2012). Geranylgeranyl pyrophosphate (GGPP) from the 2-C-methyl-D-erythritol 4-phosphate pathway initiates the carotenogenic pathway for FX biosynthesis (Dambek et al. 2012; Mikami and Hosokawa 2013; Zarekarizi et al. 2019). Phytoene synthase (PSY) catalyzes the conversion of GGPP to phytoene, and phytoene desaturase (PDS) desaturates phytoene to ζ-carotene. Moreover, carotenoid isomerase (CRITISO), lycopene β-synthase (LCYB), zeaxanthin synthase (ZEP), and violaxanthin de-epoxidase-like protein 1 (VDL1) catalyze the sequential conversion of ζ-carotene to intermediates including prolycopene, lycopene, β-carotene, zeaxanthin, antheraxanthin, violaxanthin, and neoxanthin. Diadinoxanthin (DD) and FX are the two final co-products of carotenogenesis. In particular, the regulation of genes involving the FX biosynthetic pathway was validated with various approaches. First, overexpression of genes such as PSY and DXS and triple transformant overexpressing genes violaxanthin de-epoxidase (VDE), Vde-related (VDR), and ZEP enhance the FX accumulation in P. tricornutum (Kadono et al. 2015; Eilers et al. 2016; Manfellotto et al. 2020). The second approach is to investigate the effect of environmental factors on FX biosynthesis and regulatory genes. For instance, LED light shift and nitrogen supplement improved FX yield in P. tricornutum along with the upregulation of genes such as PSY, ZEP, VDE, and fucoxanthin-chlorophyll-protein b (FCP-b). Notably, a recent study attempting to create simultaneous knockout (KO) of multiple LHCF genes reveals the phenotypic differences in term of FX-brown color accumulation between mutants and wild type (Sharma et al. 2021).
In addition to the major chlorophyll (Chl) a pigment, P. tricornutum has Chl c and FX pigments, which replace Chl b and lutein found in plants, respectively, and are bound to antenna proteins, forming the fucoxanthin-chlorophyll a/c binding protein (FCP) complex, a functional light-harvesting complex (LHC). Antenna proteins are so called apoproteins which are encoded by Lhc-family genes including LHCF, encoding the major fucoxanthin Chl a/c protein; the red algal-like LHCR; and LI818-like LHCX genes (Nymark et al. 2009). Compared with plant LHC, the diatom FCP complex has a different pigment composition and flexible structures to sustain life under fluctuating light conditions (Büchel 2019). The FCP complex is responsible for light-harvesting and electron transfer reactions during photosynthesis. In P. tricornutum the FCP complex is associated with photosystems (PS) I and II in the thylakoid membranes (TM) (Wang et al. 2019). Apart from light harvesting capacity, FCPs have been also investigated for the photoprotective capacity (Giovagnetti and Ruban 2017; Nagao et al. 2021). FX dominance in the FCP pigment pool implies a high correlation between FX biosynthesis and FCP complex formation in the TM. Upon light change, TM structure is supposed to change including membrane lipid and FCP antennae compositions, FCP-bound pigments, number of Cyt b6/f, and ATP synthase complex (Lepetit et al. 2012). For example, HL resulted in a decrease of monogalactosyl-diacylglycerol (MGDG) but the accumulation of digalactosyl-diacylglycerol (DGDG) and sulfoquinovosyl-diacylglycerol (SQDG) and phosphtidylglycerol (PG) in TM of P. tricornutum compared to LL culture (Lepetit et al. 2012). Higher amount of total secondary pigments (DD and DT) is obtained from FCP fraction of TM isolated from HL-grown Cyclotella meneghiniana (Lepetit et al. 2010).
Numerous studies have reported the effect of light intensity on FX production in P. tricornutum; however, the regulatory mechanism of FX biosynthesis under different light conditions has not been fully elucidated. This study aimed to explore the FX production regulated by different light intensities, especially via the interdependence of FX biosynthesis and FCP apoprotein composing of FCP complex.
Materials and methods
Chemicals
HPLC-grade solvents such as acetonitrile, ethanol, methanol, and water were purchased from Daejung Chemicals and Metals (South Korea). Standard FX was purchased from Sigma Aldrich (USA).
P. tricornutum culture and microalgal growth
Phaeodactylum tricornutum Bohlin strain 4 (Pt4, UTEX 646) was obtained from the University of Texas (https://utex.org/). F/2 medium components were prepared as described by Guillard and Ryther (1962). The P. tricornutum cultivation method described by Vuong et al. (2019) was adopted to utilize self-assembled light systems. Phaeodactyum tricornutum was first continuously acclimated at 20 ± 2 °C, 100 μmol photons m−2 s−1 fluorescent light with air supplement for several passages. An approximate 1 × 105 cells mL−1 of the stock was inoculated into 2 L of 3% artificial seawater supplemented F/2 medium components. The culture was grown under a constant LL (approximately 20 μmol photons m−2 s−1) or HL (approximately 200 μmol photons m−2 s−1) for 10 days. For the light transition experiment, light intensity was switched from high-to-low light (HL/LL) and low-to-high light (LL/HL) on day 5 of the 10-day culture period. Cell density was recorded daily at the same time to determine the microalgal growth. A volume of 1 mL sample was withdrawn for cell counting using a Neubauer hemocytometer. Three biological replicates were performed for each light intensity.
Determination of FX concentration and production
FX concentrations obtained from fresh biomass were determined using high performance liquid chromatography (HPLC) with diode-array detection (DAD). A 10 mL sample was harvested daily from day 2 to day 10. The cell pellet was collected using centrifugation (3,000 × g, 15 min). The fresh pellet was re-suspended with 1 mL distilled water and transferred into new micro-centrifuge tube. The sample was washed once and recollected using centrifugation (8,000 × g, 5 min, 4 °C). Supernatant was removed carefully by pipetting and quick spinning to remove all remained water. Sample tubes are dried at room temperature for approximately 5 min.
Weights of the empty tube and the sample tube were measured using a calibrated analytical balance (Mettler Toledo, Switzerland). Collected samples were frozen in liquid nitrogen and stored at -80 °C until further analysis.
The FX extraction and HPLC quantification method were similar to a previously published method, with minor modifications (Koo et al. 2019). FX was extracted from each harvested biomass by adding 1 mL of absolute ethanol. The sample was vortexed vigorously and sonicated in an ultrasonic bath (UCP-10, USA) for 15 min at room temperature. The extract was filtered via a 0.45 μm membrane filter and used for HPLC quantification. HPLC analysis was performed using an Agilent HPLC–DAD 1260 series (Agilent Technologies, USA). Briefly, FX separation was performed with a CAPCELL PAK C18 MG II column (5 μm particle size, 250 × 4.6 mm I.D.) by using binary mobile phases including water and acetonitrile at a flow rate of 1 mL min−1. For the gradient condition, 90% acetonitrile was increased to 100% over 8 min and maintained for 3 min, and then decreased to 90% over 5 min. Calibration curve of FX standard was constructed with the concentration range of 3.125 ~ 50 μg mL−1 and used for FX content quantification. Three biological replicates were performed for each group. FX productivity was determined as follows:
where: CFX (mg g−1) is FX content calculated with the standard curve; m (mg) is the fresh weight harvested from 10 mL sample; V is the volume of sample (mL).
Thylakoid membrane fraction isolation
Fresh biomass was collected from a 100 mL sample on day 5 of culture according to the above protocol, with minor modifications, including washing with phosphate-buffered saline and freezing with liquid nitrogen before storage. The TM isolation method described previously (Lepetit et al. 2007) was adopted with some modifications. Briefly, fresh samples were thawed quickly on ice and suspended in 2 mL of isolation buffer[10 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES) pH 7.4, 5 mM ethylenediamine tetraacetic acid (EDTA), 2 mM potassium chloride (KCl), and 1 M sorbitol]. The cells were lysed using a sonicator at 30% amplification (5 cycles of 15 s on and 15 s off). The lysates were centrifuged (3,000 × g, 12 min, 4 °C) to remove the unbroken cells and cell debris. The TM fraction was collected from the supernatant using centrifugation (20,000 × g, 20 min, 4 °C). The membrane fraction was washed twice with washing buffer (10 mM HEPES pH 7.4, 5 mM EDTA, and 2 mM KCl) via centrifugation (20,000 × g, 20 min, 4 °C). The TM fraction was re-suspended in small volume of washing buffer, flash-frozen in liquid nitrogen, and stored at -80 °C until FCP semi-quantification. For pigment analysis, relative TM fraction amount was measured in the same way of fresh weight determination and stored at -80 °C until pigment extraction. Three biological replicates were performed for each group.
Determination of photosynthetic pigment concentrations in thylakoid membrane fraction
The photosynthetic pigments Chls a, c and FX were extracted from the TM fraction by adding 1 mL of absolute methanol. The extraction was performed according to a previous FX extraction protocol provided in the upper part. Photosynthetic pigment-containing extracts were collected by centrifugation (8,000 × g, 5 min, 4 °C), and Chl a and c concentrations were determined using a Cary 60 UV–vis spectrophotometer at ambient temperature. According to the extinction coefficients reported by Jeffrey and Humphrey (1975) and adjusted by Ritchie (2006), absorbance values were recorded at wavelengths of 665 nm and 632 nm.
FX concentration was measured using HPLC–DAD as mentioned in the previous experiment.
Semi-quantification of FCP in isolated TM fraction
The Pierce Rapid Gold BCA protein assay kit (Thermo Fisher Scientific, USA) was used for protein quantification of TM fractions. Proteins (20 μg) were precipitated overnight in a five-fold volume of cold acetone at -20 °C. Protein precipitates were pelleted by centrifugation (> 20,000 × g, 10 min, 4 °C) and briefly dried. The pellets were resuspended in loading buffer (LDS sample buffer, 50 mM dithiothreitol) and denatured by heating at 100 °C for 3 min. Protein samples were resolved on 4–12% NuPAGE Bis–Tris gels (Life Technologies, USA). The gels were stained with the Coomassie Brilliant Blue reagent. Each target band was cut into 1 mm3 cubes and in-gel digested according to Shevchenko et al. (2006). After tryptic digestion, the peptide extracts were concentrated using SpeedVac (SPD1030, Thermo Fisher Scientific). Dried peptides were reconstituted in 10 μL of solvent A (0.1% formic acid in water) and centrifuged. Peptides (2 μL) were loaded for LC–MS/MS analysis using an Easy-nLC 1000 system (Thermo Fisher Scientific) and Q Exactive Orbitrap high-resolution mass spectrometer (Thermo Fisher Scientific). The samples were trapped on an Acclaim PepMap 100 C18 column (75 μm × 2 cm, Thermo Fisher Scientific) and separated on an EASY-Spray column (75 μm × 15 cm; particle size, 3 μm; pore size, 100 Å; 35 °C; PepMapM RSLC C18, Thermo Fisher Scientific) using a 90 min gradient of solvents A and B (0.1% formic acid in acetonitrile). The gradient was 5–40% solvent B at a flow rate of 300 nL min−1. Eluting peptides were ionized with a voltage of 2.2 kV. We performed a full MS scan ranging from 400 to 2,000 m/z with a resolution of 70,000, followed by a data-dependent higher-energy collision dissociation MS/MS scan of the top 10 ions. Parameters used for MS/MS scan were as follows: resolution, 17,500; loop count, 10 (Top 10); isolation window, 2.0 m/z; normalized collision energy, 27; peptide match, preferred; exclude isotope, on; dynamic exclusion, 20.0 s. Raw tandem spectra files were searched against P. tricornutum database (txid: 556,484; downloaded on 6 June 2021 from UniProtKB: http://www.uniprot.org/proteomes) using Proteome Discoverer (v2.4.1.15; Thermo Fisher Scientific). The search settings were as follows: Sequest HT search engine, trypsin (full), maximum 2 missed cleavage sites, MS tolerance 10 ppm, MS/MS tolerance 0.02 Da, carbamidomethylation (C) for fixed modification, and oxidation (M) for variable modification. Peptide/protein identification and quantitation were performed using a Scaffold (v4.11.1, Proteome Software Inc.) with a peptide/protein false discovery rate threshold of 0.01/0.01. For protein identification, a minimum of two peptides were required.
RNA sequencing of P. tricornutum on day 5 of culture
RNA was isolated from HL and LL culture samples on day 5. A Qiagen Plant RNeasy Mini kit (Qiagen, Germany) was used to isolate total RNA from P. tricornutum, according to the manufacturer’s instructions. RNA concentration and purity were determined using a NanoDrop ND-1000 UV/Vis spectrometer (Thermo Fisher Scientific) and Tris–borate-EDTA agarose gel electrophoresis, respectively. The qualified samples were subjected to RNA sequencing (SEEDERS Inc., Korea). RNA degradation was examined by measuring RNA integrity number (RIN > 7). The integrity of RNA samples were computed by using Agilent 2100 bioanalyzer system coupled with Agilent 2100 Expert software (Agilent Technologies, USA) following the manual instruction. The pre-cDNA library was prepared using the Illumina RNA-Seq protocol and sequenced using Illumina HiSeq 2000. Raw sequence reads have been deposited in GenBank, National Center for Biotechnology Information (NCBI) under accession BioProject PRJNA779221. Raw sequence reads were pre-processed using the DynamicTrim and LengthSort programs from the SolexaQA package. Further transcriptome data analyses were described by Kwon et al. (2021). In this study, differentially expressed genes (DEGs) related to carotenoid, Chl biosynthesis, and photosynthesis were identified using the TBLASTN tool with an e-value of ≤ 1e−10. HL samples were selected as a control group for the normalzing analyses.
Quantitative real-time polymerase chain reaction (qRT-PCR)
Primers were obtained from previous studies or self-designed using GenScript, and their specificity was confirmed using the National Center for Biotechnology Information Blast tool. Primer information is provided in Table S1. Total RNA was isolated from the HL and LL cultures harvested at two-day intervals. RNA isolation was performed using RNAiso Plus, according to the protocol. Reverse transcription was conducted using the TOYOBO ReverTra Ace–α- high efficiency reverse transcription kit, according to the manufacturer’s instructions for cDNA synthesis. The histone H4 (H4) gene was selected as the reference gene. Roche SYBR Master Mix and Applied Biosystems 7500 were used for qRT-PCR. qRT-PCR was run for 35 cycles. The threshold and baseline were set automatically but manually double-checked for proper analysis. Cycle threshold (CT) values were extracted from the results to calculate the relative expression of target gene transcripts normalized against the reference gene H4, according to the 2−∆∆CT method (Livak and Schmittgen 2001). Three biological replicates were performed for each light intensity. In this experiment, HL sample at day 2 was selected as a control group for the normalization to determine fold change in the transcription levels of investigated genes.
Validation of RNA sequencing data
Raw data of all biological replicates was first analyzed with correlation matrix-online software found in the resources of Statistical tools for high-throughput data analysis (STHDA, USA). Then the correlation graph was constructed using Google Sheets (https://docs.google.com/spreadsheet/). A Pearson correlation test between gene expressing levels based on RNA sequencing at day 5 and qRT-PCR analysis was performed using online tool Google add-ons, XLMiner Analysis ToolPak to determine correlation coefficient (r).
Statistical analysis
All data are expressed as mean ± standard deviation of biological triplicates. Statistical analyses, including unpaired t-test, two-tailed analysis, and two-way analysis of variance (ANOVA), followed by Tukey’s test, were performed using the statistical software PRISM 7 (p < 0.05 or p < 0.01, 95% or 99% confidence interval).
Results
Effect of light intensity on cell growth and FX production
Phaeodactylum tricornutum was grown under continuous LL or HL conditions for 10 days. Compared with LL, HL enhanced the growth rate of P. tricornutum (Fig. 1a). Consistent with previous studies (McClure et al. 2018; Conceição et al. 2020), FX content in the LL-grown culture was significantly higher than that in the HL-grown culture (Fig. 1b). A positive correlation between cell density and biomass was observed because the increase of cell number was paralleled by the gradual increase of biomass (Fig. S1). In the LL culture, FX continuously accumulated until day 6 and reached a maximum value of approximately 1.7 mg g−1. In contrast, FX content in the HL culture was lower, and reached a maximum of approximately 0.54 mg g−1 on day 3. The LL culture produced less biomass (Fig. S1); however, excessive FX accumulation indicated that FX production in LL-grown P. tricornutum culture was approximately 2–3 times higher than that in the HL culture. The highest FX productivity of approximately 2.52 mg L−1 day−1 was observed in the LL-grown culture (Fig. 1c). The samples at different growth stages and with different FX content were collected on day 5 for pigment and FCP proteomic analyses of the TM fraction and RNA transcriptomic analysis.
Effect of different light intensities on the growth of P. tricornutum and fucoxanthin (FX) production. (a) Growth rate of P. tricornutum microalgae was determined using cell density (× 104 cells mL−1) measured daily for 10 days of culture; (b) FX content (mg g−1, fresh weight) in P. tricornutum was measured using HPLC; (c) FX productivity (mg L−1 day−1) by P. tricornutum. Cell growth and FX measurement were carried out in biological triplicates. Different letters denote significant differences analyzed using two-way ANOVA, followed by Tukey’s test, p < 0.05
Relative quantification of photosynthetic pigments in the TM fraction
The photosynthetic machinery comprising PSII-FCP and PSI-FCP super complexes has been characterized in the P. tricornutum TM (Büchel 2019). Therefore, determining the composition of photosynthetic pigments extracted from the TM fraction could reveal the effect of light intensity on the density of pigment-containing complexes, essentially FCP complex. Quantification of photosynthetic pigments demonstrated that all pigments, including Chl a, Chl c, and FX, were significantly higher in the LL culture TM than in the HL culture TM (Fig. 2a-b). Briefly, the Chl a content extracted from the LL culture TM fraction was 0.541 mg gTM−1 which was nearly 5-times higher than Chl a content extracted from the HL culture TM fraction. Chl c was detected in trace amounts; however, a significantly higher Chl c content was obtained from the LL culture TM fraction. Notably, HPLC analysis revealed that the FX content obtained from the LL culture TM fraction (0.446 mg gTM−1) was nearly four times greater than that obtained from the HL culture TM fraction (0.108 mg gTM−1).
Thylakoid membrane (TM) fraction analysis. (a) Chlorophylls a, c quantified using spectrophotometer in TM fractions isolated from high light (HL)- or low light (LL)-grown P. tricornutum on day 5; (b) Fucoxanthin content (mg g−1, thylakoid membrane) in P. tricornutum was measured using HPLC; (c) Separation of the FCP antennae-enriched band from the total protein pool of TM fractions using SDS-PAGE before LC–MS/MS proteome analysis. In SDS-PAGE analysis, 20 μg of total protein was loaded each lane. The experiment was performed in biological triplicate. Asterisk denotes the significant difference between HL and LL culture analyzed using unpaired two-tailed t-test, p < 0.01. FX, fucoxanthin; TM, thylakoid membrane. Lanes 1, 2, 3: three replications under HL condition; Lanes 4, 5, 6: three replications under LL condition
Identification and quantification of FCPs
To identify the protein composition of FCP complexes, we performed sodium dodecyl-sulfate polyacrylamide gel electrophoresis (SDS-PAGE) of the TM fraction proteins obtained from P. tricornutum. As shown in Fig. 2c, the intensity of FCP-enriched bands with a molecular weight of approximately 18–19 kDa was higher in the LL culture TM fraction than in the HL culture TM fraction. These protein bands were extracted and used for liquid chromatography with tandem mass spectrometry (LC–MS/MS); as shown in Table 1, normalized weighted spectrum count (NWSC) revealed that FCP antenna were enriched in the protein pools of the LL culture TM fraction (56.4%) and the HL culture TM fraction (45.5%). The Lhcf3/4 antenna protein was the predominant protein in the TM fractions of both HL and LL cultures. The NWSC of Lhcf3/4 antenna protein was 161 ± 7 in the HL culture TM fraction and 292 ± 9 in the LL culture TM fraction. Remarkably, the NWSC of Lhcf5 antenna protein was nearly 20 times higher in the LL culture TM (216 ± 18) than in the HL culture TM (12 ± 2). The NWSC of Lhcf8 antenna protein, the second most abundant protein in the HL culture TM, doubled from 39 ± 9 under HL to 93 ± 23 under LL. Similarly, a drastic increase in Lhcf10 antenna protein levels of approximately four times was observed in the LL culture TM fraction compared to the HL culture TM fraction.
Transcriptome analysis of day 5 P. tricornutum
After sequence pre-processing, the transcriptome annotation of P. tricornutum on day 5 under LL or HL conditions was performed based on the reference functional information of genes identified in P. tricornutum and other Stramenopiles from the Non-Redundant database. In particular, 10,133 of 10,408 reads had a functional description with a mapping rate of approximately 91.04% relative to the reference genome. The transcriptome analysis revealed that different light intensities resulted in the differential expression of functional genes or DEGs. Compared with those in the HL culture, 429 DEGs were upregulated and approximately 507 DEGs were downregulated in the LL culture (Fig. S2). Further Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis revealed that most DEGs were related to metabolism (Fig. S2), and global and overview maps (Fig. S2). Gene ontology analysis classified DEGs into three main categories: biological processes, cellular components, and molecular functions. DEGs involved in cell growth- and photosynthesis-related processes are presented in Fig. 3a (Lou et al. 2020). In the biological process category, most key genes regulating cell growth and photosynthesis were downregulated, whereas four genes related to photosynthesis and photoreceptors including ATP-dependent zinc metalloprotease (PtFTSH), zinc metalloproteinase aureolysin (PtAUR), cryptochrome photolyase family 1 PtCPF, and sensor kinase protein 3 (PtSKP3) were upregulated in the LL culture. In the cellular component category, most genes involved in cellular component formation were downregulated, excluding the higher expression of genes involved in the formation of integral (34 genes) and intrinsic (34 genes) membrane components, in the LL culture. In the molecular function category, lyase activity (18 genes) and transferase activity (29 genes) were upregulated, whereas transporter activity (34 genes), ATPase activity (25 genes), and binding activity (82 genes) were strongly downregulated in the LL culture. Notably, 17 genes involved in the carotenogenic pathway of FX biosynthesis in P. tricornutum were identified (Fig. 3b). RNA sequencing revealed the upregulation of PtPSY, PtPDS2, PtCRTISO isoforms 1, 2, and 4, and PtVDL1 in LL-grown P. tricornutum. We identified 17 Lhcf, 14 Lhcr, and 4 Lhcx genes associated with FCP antennae biosynthesis. Among the 17 Lhcf genes which is responsible for the major fucoxanthin-Chl a/c binding protein, only PtLhcf5 (2.3-fold), PtLhcf8 (1.9-fold), and PtLhcf15 (1.8-fold) were upregulated in P. tricornutum LL culture on day 5 (Fig. 3c).
Gene ontology enrichment analysis. (a) Gene ontology enrichment analysis of differentially expressed genes identified in low or high light-grown P. tricornutum on day 5; expression regulation of target genes involved in (b) the carotenogenic pathway; and (c) FCP antenna biosynthesis in response to low light. n.d.: not detected. Results were normalized to data obtained from high light (HL)-grown P. tricornutum on day 5 of culture
Determination of potential key genes regulating FX biosynthesis
qRT-PCR analysis was performed to examine the expression pattern of potential key genes selected among the upregulated genes and genes encoding dominant FCP antennae. In addition, a positive correlation between qRT-PCR data and transcriptomic data at day 5 was validated with Pearson correlation test (r = 0.636, Fig. S3). The transcript levels of four genes (PtPSY, PtPDS1, PtPDS2, and PtVDL1) involved in carotenogenesis and three genes (PtLhcf3/4, PtLhcf5, and PtLhcf8) were measured at two-day intervals (Fig. 4). Consistent with the transcriptomic analysis results, the qRT-PCR results demonstrated the upregulation of the selected genes throughout the LL culture period. Notably, PtPDS2 exhibited the greatest change in transcript levels, which were upregulated by more than sixfold. Carotenogenesis genes generally maintained high transcript levels throughout the culture period. In contrast, Lhcf genes reached the maximum transcript level on day 2, which then decreased drastically until day 10. Among the three Lhcf genes, PtLhcf5 in the LL culture on day 2 exhibited over fourfold higher transcript levels than those in the HL culture.
Time course qRT-PCR analysis of the transcript levels of genes involved in carotenogenic pathway for fucoxanthin biosynthesis and of Lhcf genes encoding FCP antennae in P. tricornutum. Different letters denote significant differences analyzed using two-way ANOVA, followed by Tukey’s test, p < 0.05. PSY, phytoene synthase; PDS, phytoene desaturase; VDL, violaxanthin de-epoxidase-like gene; ZEP, zeaxanthin epoxidase; Lhcf, light harvesting chlorophyll a/c-fucoxanthin protein. Results were normalized to data obtained from high light (HL)-grown P. tricornutum on day 2 of culture
Effect of light transition on the cell growth of and FX production in P. tricornutum
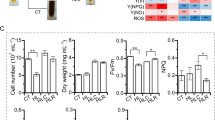
The results shown in Fig. 1 indicate that higher biomass was obtained under HL, whereas higher FX content was obtained under LL. Therefore, a light transition approach was expected to improve FX productivity of P. tricornutum. Although cell growth was likely saturated by HL/LL transition, LL/HL transition significantly increased cell density, which was 1.6 × 107 cells mL−1 at day 10 (Fig. 5a). Compared with continuous illumination, both light transitions resulted in similar cell densities as the HL culture (1.4 × 107 cells mL−1, Fig. 1a). However, LL/HL and HL/LL transitions did not positively impact FX biosynthesis compared with continuous LL. LL/HL transition decreased FX content from 1.85 mg g−1 on day 5 to 0.56 mg g−1 on day 10 (Fig. 5b). Interestingly, HL/LL transition slightly decreased FX content from 0.82 mg g−1 on day 5 to 0.68 mg g−1 on day 10 (Fig. 5b). At the harvesting point, the overall FX production following HL/LL and LL/HL transitions (1.25 mg L−1 day−1 and 1.26 mg L−1 day−1, respectively) was significantly lower than that in the LL culture (2.52 mg L−1 day−1; Fig. 5c). To examine photosynthetic pigment alterations and protein levels in the FCP-enriched region of the TM fraction in response to light transitions, we performed pigment quantification and SDS-PAGE analyses of the TM fractions isolated from P. tricornutum on day 5 (before light transition) and day 8 (3 days after light transition; Fig. S4). Only slight differences in FX content and FCP-enriched band intensity were observed following HL/LL transition. In contrast, a significant reduction in Chl a and FX content, coupled with the fading of the FCP-enriched band, was observed following LL/HL transition.
Effect of light transition on the growth of P. tricornutum and fucoxanthin (FX) production. (a) Growth rate of P. tricornutum microalgae was determined by measuring cell density (× 104 cells mL−1) daily for 10 days; (b) FX content (mg g−1, fresh weight) in P. tricornutum measured using HPLC analysis; (c) FX produced by P. tricornutum (mg L−1 day−1). Light transition was the switch from high-to-low light (HL/LL) and from low-to-high light (LL/HL). Cell growth and FX measurement were carried out in biological triplicates. Different letters denote significant differences analyzed using two-way ANOVA, followed by Tukey’s test, p < 0.05. Black arrow marks the time point of light transition
Discussion
Notably, different light intensities exerted different effects on FX production and cell growth (Fig. 1). Consistent with previous results (Gómez-Loredo et al. 2015; McClure et al. 2018; Wang et al. 2018; Conceição et al. 2020), we observed that FX production in P. tricornutum was consistently favored by lower light intensity. Particularly, P. tricornutum strain UTEX 646 cultured under LL intensity (20 μmol photons m−2 s−1) contained over 1.7 mg g−1 FX content of biomass, whereas this value was three times lower under HL intensity (200 μmol photons m−2 s−1) (Wang et al. 2018). In contrast, HL accelerated the growth rate, but negatively affected FX biosynthesis. Irradiance, a critical growth factor for photosynthesis, is known for its diverse effect on the microalgal growth (Singh and Singh, 2015). HL facilitates the conversion of inorganic carbon into biomass production. Our transcriptome analysis revealed that most genes involving cellular metabolism were down-regulated in response to LL (Fig. S1). In addition, GO enrichment analysis revealed the higher number of genes regulating cell division was down-regulated in LL culture compared to HL culture (Fig. 3). Another common observation was that FX accumulation was strongly related to the microalgal growth (McClure et al. 2018; Wang et al. 2018). Similar results of light-dependent FX production and growth were also observed in other diatoms, such as Nitzchia levis, O. aurita, and Cylindrotheca fusiformis (Xia et al. 2013; Lu et al. 2018; Wang et al. 2018).
These repeated observations prompted the question of the fundamental mechanism regulating FX biosynthesis at different light intensities. In this study, we hypothesized that FX biosynthesis was highly correlated with FCP complex formation at different light intensities. Under LL conditions, P. tricornutum requires more FCP complexes to compensate for the reduction in photon flux. Thus, photosynthetic pigments including FX and FCP antennae should be provided to form more FCP complexes (Fig. 6). Higher FX content and FCP-enriched protein bands obtained from LL culture TM shown in Fig. 2 strongly supported our hypothesis. The increase in FCP antenna levels in the LL culture TM fraction suggested that P. tricornutum might require more FCP complexes or larger complexes to overcome the lack of photon flux under LL. In contrast to LL condition, excess photon flux must switch on the photoprotective mechanism to prevent photoinhibition and photodamage, which can threaten microalgal viability (Domingues et al. 2012). Phaeodactylum tricornutum can induce non-photochemical Chl fluorescence quenching (NPQ) to dissipate excess energy into heat to overcome HL stress or fluctuating light (Domingues et al. 2012). NPQ is mediated by the interconversion of DD into diatoxanthin (DT), which is known as the xanthophyll cycle (Lavaud et al. 2012). Photoprotection thus took priority over photosynthesis in the HL culture, which raised the demand for the secondary pigment DD to compete with FX for the same precursor (Dambek et al. 2012). HL stress may cause alterations in the photosynthetic apparatus, such as chloroplast fragmentation and TM degradation, leading to an increase in secondary carotenoids (DD and DT) rather than in the primary carotenoid FX (Xia et al. 2013).
Hypothetical interdependence of FX biosynthesis and apoproteins composing of FCP complex integrated into thylakoid membrane. PSY, phytoene synthase; PDS, phytoene desaturase; CRTISO, carotenoid isomerase; VDL, violaxanthin de-epoxidase-like gene; LHCF, light harvesting chlorophyll a/c-fucoxanthin binding proteins; FCP, fucoxanthin-chlorophyll a/c binding protein; PSI, photosystem I; PSII: photosystem II
FCP antennae are transmembrane protein complexes, whose fundamental subunits are mainly encoded by Lhcf genes (Wang et al. 2019). The location of the FCP complex in the P. tricornutum TM was first discovered in 1992 (Pyszniak and Gibbs 1992). Recently, an increasing number of studies have focused on FCP, including its formation, structure, pigments, and protein compositions (Büchel 2003; Grouneva et al. 2011; Nymark et al. 2019; Wang et al. 2019). Nevertheless, the relationship between FX biosynthesis and FCP formation has likely been neglected, which is evident from the lack of studies focusing on it, particularly under varied illumination conditions, which is the most essential factor for photosynthesis. Our hypothesis of the correlation between FX biosynthesis and FCP formation was substantially reinforced by certain statements merely suggested elsewhere in others’ study. Briefly, genes encoding FCP antennae might be post-transcriptionally controlled under silicon starvation; therefore, FX could not efficiently bind to antenna proteins, leading to a reduced FX production (Zhao et al. 2014). Moreover, FCPs were more abundant in aluminum-treated cells, which was accompanied by enhanced photosynthetic pigments in these cells (Xie et al. 2015). In a study on improving FX production using an approach combining LED shift and nitrogen supplementation, Yang and Wei (2020) reported a significantly upregulated FCPb, which corresponds with an increase in FX content.
In the present study SDS-PAGE coupled with LC–MS/MS and RNA transcriptome analyses revealed the correlation between encoding genes and antenna proteins regulating FCP formation (Fig. 3c, Table. 1). In consistent with the enrichment of photosynthetic pigments, the levels of most identified FCP antennae increased in the LL culture. In particular, Lhcf5, Lhcf8, and Lhcf10 levels significantly increased in response to LL. Similarly, the alteration of the FCP antenna distribution caused by LL and HL was also observed by Gundermann et al. (2013) and Nagao et al. (2021). Lhcf5 was identified as a fundamental protein of the stable trimer FCPA unit isolated from P. tricornutum wild type grown under LL (Joshi-Deo et al. 2010). Moreover, a study on TM proteome identified ten Lhcf proteins in the FCP trimer units of P. tricornutum, with Lhcf3/4 being the most abundant antenna protein (Grouneva et al. 2011). The current study also identified Lhcf3/4 as a major light-harvesting protein in the FCP antennae pool of the HL and LL culture TM.
Lhcx1 was the only stress-related LI818/LhcSR-like protein identified in FCP-enriched band. Previous findings reveal that Lhcx1 gene is highly expressed under LL condition and is not induced by further HL acclimation (Nymark et al. 2009; Bailleul et al. 2010). Particularly, when Lhcx1 gene upregulated in cells exposed to HL after 30 min, its mRNA level returned to baseline or even downregulated during longer HL exposure (Nymark et al. 2009). The constitutive presence of Lhcx1 protein in cells acclimated to non-stressful condition like LL enables the organism to respond rapidly to sudden changes (Bailleul et al. 2010). Along with findings related to Lhcx1 gene expression under different light intensities, we observed that the level of Lhcx1 protein in FCP fraction of HL culture was fourfold less than that in LL culture. However, some previous findings such as the recent one from Taddei et al. (2018) indicated that 2-day HL-acclimated cells enrich more Lhcx1 protein in FCP fraction than that in LL-adapted cells. In Taddei’s study, cells were grown under HL and LL with photoperiod of 12 h: 12 h light/dark cycle while microalgal cells were continuously illuminated with constant HL and LL in our study. Another study of Taddei et al. (2016) indicated that various environmental signals can control the expression of Lhcx protein family including Lhcx1. Moreover, Lhcx1 which is fivefold accumulated more in cells grown under intermittent light than under continuous light was considered as the essential regulator of dynamic photoprotection (Giovagnetti et al. 2022). Those findings and our result suggested that P. tricornutum might possess highly flexible mechanism regulating NPQ capacity in order to adapt to different light conditions. A further study should be carried out to fully understand this complicated phenomenon.
Transcriptome analysis revealed the upregulation of PtLhcf5, PtLhcf8, and PtLhcf15 in LL culture on day 5. Previously, Lhcf15 gene was particularly noticed for its abnormal expression. The upregulation of this gene is observed in prolonged darkness and red light exposure (Nymark et al. 2013; Herbstová et al. 2017). Lhcf15 mRNA level is mainly accumulated in short time and likely stabilized (Islam et al. 2020). In this study, PtLhcf15 was upregulated in LL-grown P. tricornutum at day 5. However, Lhcf15 protein was not detected in FCP-enriched band of TM which is in consistent with the previous findings on the composition of FCP (Grouneva et al. 2011; Gundermann et al. 2013). Lhcf15 protein was only detected in red light-acclimated cells and absent in white light acclimated cells (Costa et al. 2013; Herbstová et al. 2015, 2017).
Based on proteomic result and transcriptome analysis, we performed qRT-PCR to examine the expression patterns of the selected genes encoding Lhcf antennae throughout the culture period. The results demonstrated that three genes (PtLhcf3/4, PtLhcf5, and PtLhcf8) were highly expressed in the LL culture than in the HL culture. Unlike carotenogenic genes, FCP-encoding genes had a different expression pattern: the expressing level reached a maximum on day 2 and then decreased gradually over time. Therefore, the production of pigment-binding antenna proteins likely preceded pigment production and was maximized in the early growth stage to induce FCP formation in response to the reduced photon flux caused by LL. Consequently, FX biosynthesis was possibly prolonged until the end of the exponential phase, whereas FCP antennae were synthesized in the early exponential phase. After the growth phase, inducing de novo FCP complex formation is likely difficult because that requires FX and Lhc antennae supplies.
RNA transcriptome analysis indicated that PtPSY, PtPDS2, PtCRTISO isoforms 1, 2, and 4, and PtVDL1 were highly expressed in LL cultures. In previous studies, PSY overexpression led to an increase in FX and total carotenoids (Kadono et al. 2015; Eilers et al. 2016; Manfellotto et al. 2020). Along with the enhancement of FX biosynthesis in P. tricornutum, PSY and PDS1 were upregulated in response to blue light and tryptone addition (Yang and Wei 2020). In a recent study on the correlation between specific growth rate and FX biosynthesis, PSY and PDS were upregulated in the exponential phase, which was accompanied by higher FX production than that in the stationary phase (Kwon et al. 2021). Dautermann et al. (2020) reported the essential role of VDL1 enzyme in the production of marine carotenoids. VDL1 catalyzes the conversion of violaxanthin to neoxanthin, which serves as a pool of precursors for FX synthesis. Moreover, VDL1 was previously shown to be induced by HL stress, suggesting its crucial role in the xanthophyll cycle in P. tricornutum (Kuczynska et al. 2017). In this study, PtCRTISO isoforms 1, 2, and 4 were upregulated in the LL culture on day 5; however, no previous findings demonstrate a positive correlation between CRTISO expression and FX biosynthesis. Particularly, CRTISO expression was unchanged or increased in response to HL acclimation, which had a consistent and severe impact on FX biosynthesis (Nymark et al. 2009). Furthermore, CRTISO was downregulated in the exponential phase, which is favorable for FX production (Kwon et al. 2021). Previous findings based on RNA transcriptome analysis have demonstrated that PtPSY, PtPDS1, PtPDS2, and PtVDL1 likely played critical roles in FX biosynthesis under LL conditions. Therefore, we used qRT-PCR to investigate the expression patterns of these genes at two-day intervals for 10 days. The results revealed upregulated transcript levels of PtPSY (four-fold), PtPDS1 (< two-fold), PtPDS2 (six-fold), and PtVDL1 (four-fold) in LL cultures compared to those in HL cultures (Fig. 4), which was consistent with FX accumulation under different light intensities (Fig. 1b). PtPSY, PtPDS2, and PtVDL1 might play a major role in carotenogenesis to increase FX biosynthesis. The maximum transcript levels of these genes in the earlier growth stage induced FX pigment biosynthesis following the increasing demand for FCP formation.
As mentioned above, FX has numerous human health benefits that have increased the demand for this compound as a health supplement. P. tricornutum might be a more sustainable source of FX than brown seaweeds. Compared to brown seaweeds, P. tricornutum had numerous advantages in terms of FX production; however, FX productivity in diatoms must be optimized to offset the production costs in the upstream and downstream processing (Butler et al. 2020). Therefore, more studies should be conducted to optimize FX production in P. tricornutum.
In this study, FX accumulation was favored by LL owing to the increased demand for FCP complex formation, whereas HL boosted biomass production (Fig. S1). We hypothesized that a light transition experiment might result in increased FX accumulation and biomass production and eventually enhance FX productivity. We validated the feasibility of this approach by culturing P. tricornutum under continuous LL or HL conditions until day 5, after which the LL and HL conditions were switched. Cell density, FX content, and FX productivity were measured daily (Fig. 5). Unfortunately, light transition did not have a positive effect on FX production in P. tricornutum. LL/HL transition increased biomass production to a level similar to that in the HL culture. However, after transition to HL, FX content drastically decreased at the end of cultivation. We also observed lower photosynthetic pigment concentrations and FCP enrichment in the TM on day 8 than on day 5, which might be a sign of FCP degradation caused by HL (Fig. S4). At the harvesting point (day 10), FX productivity in the HL culture increased; however, it was much lower than that in the LL culture (Figs. 1c, 5c). Despite its positive impact on biomass production, LL/HL transition severely affected FX accumulation, likely due to FCP degradation. In addition, FCP formation occurred during the early growth stage, and FCP degradation likely occurred upon exposure to HL when the prevention of photoinhibition and photodamage were prioritized. Therefore, continuous exposure to LL during the growth phase is critical to boost FX accumulation in P. tricornutum, whereas the absence of HL is crucial to prevent FCP degradation accompanied by severe FX reduction.
On the other hand, HL/LL transition resulted in equal FX productivity to LL/HL cultures on day 10 (Fig. 5c). HL/LL transition reduced the cell growth rate and did not increase cellular FX content, but rather maintained FX productivity (Fig. 5). Coincidently, no significant differences were observed in the pigmentation or FCP amount in TM fractions between days 5 and 8 (Fig. S4). However, the unexpected result of HL/LL transition could be related to the correlation between FX biosynthesis and growth phase. At day 5 under HL condition, P. tricornutum might start entering the stationary phase and essential nutrients began reducing. In this stage, cells might not grow more and FCP formation for the growth is not needed anymore.
The impact of light on cellular FX production might also be owning to the different photon flux per cell received at different growth stage. In LL/HL transition, the sudden increase of photon absorbed per cell resulted in the drastic reduction of photosynthetic pigments to prevent photodamage. In HL/LL transition, the decrease of photon absorbed per cell supposed to enhance FX accumulation but nutrient deficiency together with non de novo synthesis of LHC prevented that.
In summary, light transition experiments did not yield the expected impact on FX production which was favored by LL likely due to FCP formation only at an early growth stage. In addition, photoprotection priority in response to HL induced FCP complex degradation regardless of the growth stage.
Conclusion
Light intensities influenced FX production via the interdependence of FX biosynthesis and apoproteins demanded for FCP complex formation. We performed RNA sequencing and LC–MS/MS and identified several key genes involving carotenogenesis (PtPSY, PtPDS, and PtVDL1) and FCP complex formation (PtLhcf5 and PtLhcf8). To cope with the shortage of photon flux density in LL condition, P. tricornutum required more FCP units to enhance photon absorption that raised the demand for both Lhc antenna and FX pigment supplies. In this study, FCP complex formation occurred only in the early growth stages while FCP complex degradation could occur at any time during the culture period upon HL exposure. Therefore, HL and LL transitions in the middle of the culture period could not increase FX production compared to that observed under LL conditions.
Data availability
Raw sequencing reads obtained from RNA sequencing in this study have been deposited at NCBI SRA and are available under accession BioProject PRJNA779221 under the following link: https://www.ncbi.nlm.nih.gov/bioproject/PRJNA779221.
References
Alipanah L, Rohloff J, Winge P, Bones AM, Brembu T (2015) Whole cell response to nitrogen deprivation in the diatom Phaeodactylum tricornutum. J Exp Bot 66:6281–6296
Bae M, Kim MB, Park YK, Lee JY (2020) Health benefits of fucoxanthin in the prevention of chronic diseases. Biochim Biophys Acta-Mol Cell Biol Lipids 1865:158618–158638
Bailleul B, Rogato A, de Martino A, Coesel S, Cardol P, Bowler C, Falciatore A, Finazzi G (2010) An atypical member of the light-harvesting complex stress-related protein family modulates diatom responses to light. Proc Natl Acad Sci 107:18214–18219
Bohlin K (1897) Zur Morphologie und Biologie einzelliger Algen. (Vorläufige Mitteilung.) Öfvers Kongl Svenska Vetensk-Akad Förh 54:507–529
Bowler C, Allen AE, Badger JH, Grimwood J, Jabbari K, Kuo A, Maheswari U, Martens C, Maumus F, Otillar RP, Rayko E, Salamov A, Vandepoele K, Beszteri B, Gruber A, Heijde M, Katinka M, Mock T, Valentin K, Verret F, Berges JA, Brownlee C, Cadoret JP, Chiovitti A, Choi CJ, Coesel S, De Martino A, Detter JC, Durkin C, Falciatore A, Fournet J, Haruta M, Huysman MJJ, Jenkins BD, Jiroutova K, Jorgensen RE, Joubert Y, Kaplan A, Kroger N, Kroth PG, La Roche J, Lindquist E, Lommer M, Martin-Jezequel V, Lopez PJ, Lucas S, Mangogna M, McGinnis K, Medlin LK, Montsant A, Oudot-Le Secq MP, Napoli C, Obornik M, Parker MS, Petit JL, Porcel BM, Poulsen N, Robison M, Rychlewski L, Rynearson TA, Schmutz J, Shapiro H, Siaut M, Stanley M, Sussman MR, Taylor AR, Vardi A, von Dassow P, Vyverman W, Willis A, Wyrwicz LS, Rokhsar DS, Weissenbach J, Armbrust EV, Green BR, van de Peer Y, Grigoriev IV (2008) The Phaeodactylum genome reveals the evolutionary history of diatom genomes. Nature 456:239–244
Büchel C (2003) Fucoxanthin-chlorophyll proteins in diatoms: 18 and 19 kDa subunits assemble into different oligomeric states. Biochemistry 42:13027–13034
Büchel C (2019) How diatoms harvest light. Science 365:447–448
Butler T, Kapoore RV, Vaidyanathan S (2020) Phaeodactylum tricornutum: a diatom cell factory. Trends Biotechnol 38:606–622
Conceição D, Lopes RG, Derner RB, Cella H, do Carmo APB, D’Oca MGM, Marcelo G. Petersen R, Passos MF, Vargas JVC, Galli-Terasawa LV, Kava V (2020) The effect of light intensity on the production and accumulation of pigments and fatty acids in Phaeodactylum tricornutum. J Appl Phycol 32:1017-1025
Costa BS, Jungandreas A, Jako T, Weisheit W, Mittag M, Wilhelm C (2013) Blue light is essential for high light acclimation and photoprotection in the diatom Phaeodactylum tricornutum. J Exp Bot 64:483–493
Dambek M, Eilers U, Breitenbach J, Steiger S, Buchel C, Sandmann G (2012) Biosynthesis of fucoxanthin and diadinoxanthin and function of initial pathway genes in Phaeodactylum tricornutum. J Exp Bot 63:5607–5612
Dautermann O, Lyska D, Andersen-Ranberg J, Becker M, Fröhlich-Nowoisky J, Gartmann H, Krӓmer LC, Mayr K, Pieper D, Rij LM, Wipf HML, Niyogi KK, Lohr M (2020) An algal enzyme required for biosynthesis of the most abundant marine carotenoids. Sci Adv 6:eaaw9183
Domingues N, Matos AR, da Silva JM, Cartaxana P (2012) Response of the diatom Phaeodactylum tricornutum to photooxidative stress resulting from high light exposure. PLoS One 7:e38162
Eilers U, Bikoulis A, Breitenbach J, Büchel C, Sandmann G (2016) Limitations in the biosynthesis of fucoxanthin as targets for genetic engineering in Phaeodactylum tricornutum. J Appl Phycol 28:123–129
Feijão E, Gameiro C, Franzitta M, Duarte B, Caçador I, Cabrita MT, Matos AR (2018) Heat wave impacts on the model diatom Phaeodactylum tricornutum: searching for photochemical and fatty acid biomarkers of thermal stress. Ecol Indic 95:1026–1037
Field CB, Behrenfeld MJ, Randerson JT, Falkowski P (1998) Primary production of the biosphere: integrating terrestrial and oceanic components. Science 281:237–240
Gao B, Chen A, Zhang W, Li A, Zhang C (2017) Co-production of lipids, eicosapentaenoic acid, fucoxanthin, and chrysolaminarin by Phaeodactylum tricornutum cultured in a flat-plate photobioreactor under varying nitrogen conditions. J Ocean Univ China 16:916–924
Gille A, Stojnic B, Derwenskus F, Trautmann A, Schmid-Staiger U, Posten C, Briviba K, Palou A, Bonet ML, Ribot J (2019) A lipophilic fucoxanthin-rich Phaeodactylum tricornutum extract ameliorates effects of diet-induced obesity in C57BL/6J mice. Nutrients 11:796–815
Giovagnetti V, Ruban AV (2017) Detachment of the fucoxanthin chlorophyll a/c binding protein (FCP) antenna is not involved in the acclimative regulation of photoprotection in the pennate diatom Phaeodactylum tricornutum. Biochim Biophys Acta-Bioenerg 1858:218–230
Giovagnetti V, Jaubert M, Shukla MK, Ungerer P, Bouly JP, Falciatore A, Ruban AV (2022) Biochemical and molecular properties of LHCX1, the essential regulator of dynamic photoprotection in diatoms. Plant Physiol 188:509–525
Gómez-Loredo A, Benavides J, Rito-Palomares M (2015) Growth kinetics and fucoxanthin production of Phaeodactylum tricornutum and Isochrysis galbana cultures at different light and agitation conditions. J Appl Phycol 28:849–860
Grouneva I, Rokka A, Aro EM (2011) The thylakoid membrane proteome of two marine diatoms outlines both diatom-specific and species-specific features of the photosynthetic machinery. J Proteome Res 10:5338–5353
Guillard RR, Ryther JH (1962) Studies of marine planktonic diatoms. I. Cyclotella nana Hustedt and Detonula confervacea (Cleve) Gran. Can J Microbiol 8:229–239
Gundermann K, Schmidt M, Weisheit W, Mittag M, Büchel C (2013) Identification of several sub-populations in the pool of light harvesting proteins in the pennate diatom Phaeodactylum tricornutum. Biochim Biophys Acta-Bioenerg 1827:303–310
Herbstová M, Bína D, Koník P, Gardian Z, Vácha F, Litvín R (2015) Molecular basis of chromatic adaptation in pennate diatom Phaeodactylum tricornutum. Biochim Biophys Acta-Bioenerg 1847:534–543
Herbstová M, Bína D, Kaňa R, Vácha F, Litvín R (2017) Red-light phenotype in a marine diatom involves a specialized oligomeric red-shifted antenna and altered cell morphology. Sci Rep 7:11976
Heydarizadeh P, Boureba W, Zahedi M, Huang B, Moreau B, Lukomska E, Couzinet-Mossion A, Wielgosz-Collin G, Martin-Jézéquel V, Bougaran G, Marchand J, Schoefs B (2017) Response of CO2-starved diatom Phaeodactylum tricornutum to light intensity transition. Philos Trans R Soc B 372:20160396
Huang B, Marchand J, Blanckaert V, Lukomska E, Ulmann L, Wielgosz-Collin G, Rabesaotra V, Moreau B, Bougaran G, Mimouni V, Morant-Manceau A (2019) Nitrogen and phosphorus limitations induce carbon partitioning and membrane lipid remodelling in the marine diatom Phaeodactylum tricornutum. Eur J Phycol 54:342–358
Islam S, Sabharwal T, Wu S, Bullock TJ, Mehdy MC (2020) Early dynamics of photosynthetic Lhcf2 and Lhcf15 transcription and mRNA stabilities in response to herbivory-related decadienal in Phaeodactylum tricornutum. Sci Rep 10:2029
Jeffrey SW, Humphrey GF (1975) New spectrophotometric equations for determining chlorophylls a, b, c1 and c2 in higher plants, algae and natural phytoplankton. Biochem Physiol Pflanzen 167:191–194
Joshi-Deo J, Schmidt M, Gruber A, Weisheit W, Mittag M, Kroth PG, Büchel C (2010) Characterization of a trimeric light-harvesting complex in the diatom Phaeodactylum tricornutum built of FcpA and FcpE proteins. J Exp Bot 61:3079–3087
Kadono T, Kira N, Suzuki K, Iwata O, Ohama T, Okada S, Nishimura T, Akakabe M, Tsuda M, Adachi M (2015) Effect of an introduced phytoene synthase gene expression on carotenoid biosynthesis in the marine diatom Phaeodactylum tricornutum. Mar Drugs 13:5334–5357
Kim SM, Jung YJ, Kwon ON, Cha KH, Um BH, Chung D, Pan CH (2012) A potential commercial source of fucoxanthin extracted from the microalga Phaeodactylum tricornutum. Appl Biochem Biotechnol 166:1843–1855
Koo SY, Hwang JH, Yang SH, Um JI, Hong KW, Kang K, Pan CH, Hwang KT, Kim SM (2019) Anti-obesity effect of standardized extract of microalga Phaeodactylum tricornutum containing fucoxanthin. Mar Drugs 17:311–325
Kosakowska A, Lewandowska J, Stoń J, Burkiewicz K (2004) Qualitative and quantitative composition of pigments in Phaeodactylum tricornutum (Bacillariophyceae) stressed by iron. Biometals 17:45–52
Kuczynska P, Jemiola-Rzeminska M, Strzalka K (2017) Characterisation of carotenoids involved in the xanthophyll cycle. In: Cvetkovic DJ, Nikolic GS (eds) Carotenoids. InTechOpen, Riejeka pp 1–16
Kwon DY, Vuong TT, Choi J, Lee TS, Um JI, Koo SY, Hwang KT, Kim SM (2021) Fucoxanthin biosynthesis has a positive correlation with the specific growth rate in the culture of microalga Phaeodactylum tricornutum. J Appl Phycol 33:1473–1485
Lavaud J, Materna AC, Sturm S, Vugrinec S, Kroth PG (2012) Silencing of the violaxanthin de-epoxidase gene in the diatom Phaeodactylum tricornutum reduces diatoxanthin synthesis and non-photochemical quenching. PLoS One 7:e36806
Lepetit B, Volke D, Szabó M, Hoffmann R, Garab G, Wilhelm C, Goss R (2007) Spectroscopic and molecular characterization of the oligomeric antenna of the diatom Phaeodactylum tricornutum. Biochemistry 46:9813–9822
Lepetit B, Volke D, Gilbert M, Wilhelm C, Goss R (2010) Evidence for the existence of one antenna-associated, lipid-dissolved and two protein-bound pools of diadinoxanthin cycle pigments in diatoms. Plant Physiol 154:1905–1920
Lepetit B, Goss R, Jakob T, Wilhelm C (2012) Molecular dynamics of the diatom thylakoid membrane under different light conditions. Photosynth Res 111:245–257
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25:402–408
Lou S, Zhu X, Zeng Z, Wang H, Jia B, Li H, Hu Z (2020) Identification of microRNAs response to high light and salinity that involved in beta-carotene accumulation in microalga Dunaliella salina. Algal Res 48:101925–101935
Lu X, Sun H, Zhao W, Cheng KW, Chen F, Liu B (2018) A hetero-photoautotrophic two-stage cultivation process for production of fucoxanthin by the marine diatom Nitzschia laevis. Mar Drugs 16:219–231
Manfellotto F, Stella GR, Falciatore A, Brunet C, Ferrante MI (2020) Engineering the unicellular alga Phaeodactylum tricornutum for enhancing carotenoid production. Antioxidants 9:757–769
McClure DD, Luiz A, Gerber B, Barton GW, Kavanagh JM (2018) An investigation into the effect of culture conditions on fucoxanthin production using the marine microalgae Phaeodactylum tricornutum. Algal Res 29:41–48
Mikami K, Hosokawa M (2013) Biosynthetic pathway and health benefits of fucoxanthin, an algae-specific xanthophyll in brown seaweeds. Int J Mol Sci 14:13763–13781
Nagao R, Yokono M, Ueno Y, Suzuki T, Kumazawa M, Kato KH, Tsuboshita N, Dohmae N, Ifuku K, Shen JR, Akimoto S (2021) Enhancement of excitation-energy quenching in fucoxanthin chlorophyll a/c-binding proteins isolated from a diatom Phaeodactylum tricornutum upon excess-light illumination. Biochim Biophys Acta-Bioenerg 1862:148350–148358
Nur MMA, Muizelaar W, Boelen P, Buma AGJ (2018) Environmental and nutrient conditions influence fucoxanthin productivity of the marine diatom Phaeodactylum tricornutum grown on palm oil mill effluent. J Appl Phycol 31:111–122
Nymark M, Valle KC, Brembu T, Hancke K, Winge P, Andresen K, Johnsen G, Bones AM (2009) An integrated analysis of molecular acclimation to high light in the marine diatom Phaeodactylum tricornutum. PLoS One 4:e7743
Nymark M, Volpe C, Hafskjold MCG, Kirst H, Serif M, Vadstein O, Bone AM, Melis A, Winge P (2019) Loss of ALBINO3b insertase results in truncated light-harvesting antenna in diatoms. Plant Physiol 181:1257–1276
Petrushkina M, Gusev E, Sorokin B, Zotko N, Mamaeva A, Filimonova A, Kulikovskiy M, Maltsev Y, Yampolsky I, Vinokurov V, Namasaraev Z, Kuzmin D (2017) Fucoxanthin production by heterokont microalgae. Algal Res 24:387–393
Pyszniak AM, Gibbs SP (1992) Immunocytochemical localization of photosystem I and the fucoxanthin-chlorophyll a/c light-harvesting complex in the diatom Phaeodactylum tricornutum. Protoplasma 166:208–217
Remmers IM, Martens DE, Wijffels RH, Lamers PP (2017) Dynamics of triacylglycerol and EPA production in Phaeodactylum tricornutum under nitrogen starvation at different light intensities. PLoS One 12:e0175630
Ritchie RJ (2006) Consistent sets of spectrophotometric chlorophyll equations for acetone, methanol and ethanol solvents. Photosynth Res 89:27–41
Sharma AK, Nymark M, Flo S, Sparstad T, Bones AM, Winge P (2021) Simultaneous knockout of multiple LHCF genes using single sgRNAs and engineering of a high-fidelity Cas9 for precise genome editing in marine algae. Plant Biotechnol J 19:1658–1669
Shevchenko A, Tomas H, Havli J, Olsen JV, Mann M (2006) In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat Protoc 1:2856–2860
Singh SP, Singh P (2015) Effect of temperature and light on the growth of algae species: a review. Renew Sust Energ Rev 50:431–444
Taddei L, Stella GR, Rogato A, Bailleul B, Fortunato AE, Annunziata R, Sanges R, Thaler M, Lepetit B, Lavaud J, Jaubert M, Finazzi G, Bouly JP, Falciatore A (2016) Multisignal control of expression of the LHCX protein family in the marine diatom Phaeodactylum tricornutum. J Exp Bot 67:3939–3951
Taddei L, Chukhutsina VU, Lepetit B, Stella GR, Bassi R, van Amerongen H, Bouly JP, Jaubert M, Finazzi G, Falciatore A (2018) Dynamic changes between two LHCX-related energy quenching sites control diatom photoacclimation. Plant Physiol 177:953–965
Terasaki M, Hamoya T, Kubota A, Kojima H, Tanaka T, Maeda H, Miyashita K, Mutoh M (2021) Fucoxanthin prevents colorectal cancer development in dextran sodium sulfate-treated ApcMin/+ mice. Anticancer Res 41:1299–1305
Terry KL, Hirata J, Laws EA (1983) Light-limited growth of two strains of the marine diatom Phaeodactylum tricornutum Bohlin: chemical composition, carbon partitioning and the diel periodicity of physiological processes. J Exp Mar Biol Ecol 68:209–227
Valle KC, Nymark M, Aamot I, Hancke K, Winge P, Andresen K, Johnsen G, Brembu T, Bones AM (2014) System responses to equal doses of photosynthetically usable radiation of blue, green, and red light in the marine diatom Phaeodactylum tricornutum. PLoS One 9:e114211
Vuong TT, Kwon BR, Eom JI, Shin BK, Kim SM (2019) Interaction between marine bacterium Stappia sp. K01 and diatom Phaeodactylum tricornutum through extracellular fatty acids. J Appl Phycol 32:71–82
Wang H, Zhang Y, Chen L, Cheng W, Liu T (2018) Combined production of fucoxanthin and EPA from two diatom strains Phaeodactylum tricornutum and Cylindrotheca fusiformis cultures. Bioprocess Biosyst Eng 41:1061–1071
Wang W, Yu LJ, Xu C, Tomizaki T, Zhao S, Umena Y, Chen X, Qin X, Xin Y, Suga M, Han G, Kuang T, Shen JR (2019) Structural basis for blue-green light harvesting and energy dissipation in diatoms. Science 363:eaav0365
Xia S, Wang K, Wan L, Li A, Hu Q, Zhang C (2013) Production, characterization, and antioxidant activity of fucoxanthin from the marine diatom Odontella aurita. Mar Drugs 11:2667–2681
Xie J, Bai X, Lavoie M, Lu H, Fan X, Pan X, Fu Z, Qian H (2015) Analysis of the proteome of the marine diatom Phaeodactylum tricornutum exposed to aluminum providing insights into aluminum toxicity mechanisms. Environ Sci Technol 49:11182–11190
Yang R, Wei D (2020) Improving fucoxanthin production in mixotrophic culture of marine diatom Phaeodactylum tricornutum by LED light shift and nitrogen supplementation. Front Bioeng Biotechnol 8:820–843
Ye G, Lu Q, Zhao W, Du D, Jin L, Liu Y (2014) Fucoxanthin induces apoptosis in human cervical cancer cell line HeLa via PI3K/Akt pathway. Tumor Biol 35:11261–11267
Yi Z, Su Y, Cherek P, Nelson DR, Lin J, Rolfsson O, Wu H, Salehi-Ashtiani K, Brynjolfsson S, Fu W (2019) Combined artificial high-silicate medium and LED illumination promote carotenoid accumulation in the marine diatom Phaeodactylum tricornutum. Microb Cell Fact 18:209
Zarekarizi A, Hoffmann L, Burritt D (2019) Approaches for the sustainable production of fucoxanthin, a xanthophyll with potential health benefits. J Appl Phycol 31:281–299
Zhao P, Gu W, Wu S, Huang A, He L, Xie X, Gao S, Zhang B, Niu J, Lin AP, Wang G (2014) Silicon enhances the growth of Phaeodactylum tricornutum Bohlin under green light and low temperature. Sci Rep 4:3958–3967
Funding
This work was supported by the intramural grant (2Z06670) from the Korea Institute of Science and Technology (KIST), Republic of Korea.
Author information
Authors and Affiliations
Contributions
T.Q.T. and S.M.K. conceived and designed the study. T.Q.T. conducted experiments, collected data, and wrote manuscripts. Y.J.P., A.E., J.-H.C. and D.-G.S. assisted and partly participated in experimental design and laboratory experiments. T.T.Q. performed bioinformatics and statistical analysis with the assistance of S.Y.K. T.T.Q and S.M.K. created figures and tables with the revision of other authors. S.M.K. supervised the study, revised manuscripts, and was in charge of the final version. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Truong, T.Q., Park, Y.J., Koo, S.Y. et al. Interdependence of fucoxanthin biosynthesis and fucoxanthin-chlorophyll a/c binding proteins in Phaeodactylum tricornutum under different light intensities. J Appl Phycol 35, 25–42 (2023). https://doi.org/10.1007/s10811-022-02856-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10811-022-02856-2