Abstract
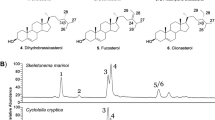
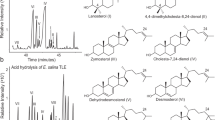
The heterotrophic alga Aurantiochytrium accumulates a high amount of squalene, which is a key precursor to the biosynthesis of sterols and steroids. However, post-squalene sterol pathways of the algae have been enigmatic. Using NMR-based chemical approaches complemented by in silico analysis of Aurantiochytrium genome, we identified 7 sterols including novel derivatives and predicted multiple sterol metabolic pathways. Based on known sterol distribution among eukaryotes, phylogeny, and our findings, we suggest that Aurantiochytrium could possess multiple sterol metabolic pathways of both photosynthetic and non-photosynthetic lineages with highly proliferated genes encoding the associated enzymes. Aurantiochytrium possibly possesses well-developed networks of sterol metabolic pathways.


Similar content being viewed by others
References
Akihisa T, Thakur S, Rosenstein FU, Matsumoto T (1986) Sterols of Cucurbitaceae: the configurations at C-24 of 24-alkyl-Δ5-, Δ7- and Δ8-sterols. Lipids 21:39–47
Akihisa T, Shimizu N, Ghosh P, Thakur S, Rosenstein FU, Tamura T (1987) Sterols of the cucurbitaceae. Phytochemistry 261:1693–1700
Akihisa T, Matsubara Y, Ghosh P, Thakur S, Shimizu N, Tamura T, Matsumoto T (1988) The 24α- and 24β-epimers of 24-ethylcholesta-5,22-dien-3β-ol in two Clerodendrum species. Phytochemistry 27:1169–1172
Albrizio S, Ciminiello P, Fattorusso E, Magno S, Pansini M (1994) Chemistry of Verongida sponges. I. Constituents of the Caribbean sponge Pseudoceratina crassa. Tetrahedron 50:783–788
Bajpai P, Bajpai PK, Ward OP (1994) Production of docosahexaenoic acid by Thraustochytrium aureum. Appl Microbiol Biotechnol 35:706–710
Baldauf SL (2003) The deep roots of eukaryotes. Science 300:1703–1706
Balliano G, Caputo O, Viola F, Delprino L, Cattel L (1983) Conversion of cyclolaudenol to 24α- and 24β-ethylsterols in the cucurbitaceae. Lipids 18:302–305
Desmond E, Gribaldo S (2009) Phylogenomics of sterol synthesis: insights into the origin, evolution, and diversity of a key eukaryotic feature. Genome Biol Evol 1:364–381
Emmons GT, Wilson KW, Schroepfer GJ Jr (1989) 1H and 13C NMR assignments for lanostan-3β-ol derivatives: revised assignments for lanosterol. Magn Reson Chem 27:1012–1024
Fabris M, Matthijs M, Carbonelle S, Moses T, Pollier J, Dasseville R, Baart GJ, Vyverman W, Goossens A (2014) Tracking the sterol biosynthesis pathway of the diatom Phaeodactylum tricornutum. New Phytol 204:521–535
Forestier E, Romero-Segura C, Pateraki I, Centeno E, Compagnon V, Preiss M, Berna A, Boronat A, Bach TJ, Darnet S, Schaller H (2019) Distinct triterpene synthesis in the laticifers of Euphorbia lathyris. Sci Rep. https://doi.org/10.1038/s41598-019-40905-y
Gershengorn MC, Smith AR, Goulston G, Goad LJ, Goodwin TW, Haines TH (1968) The sterols of Ochromonas danica and Ochromonas malhamensis. Biochemistry 7:1698–1706
Goad LJ, Lenton JR, Knapp FF, Goodwin TW (1974) Phytosterol side chain biosynthesis. Lipids 9:582–595
Gold DA, Grabenstatter J, de Mendoza A, Riesgo A, Ruiz-Trillo I, Summons R (2016) Sterol and genomic analyses validate the sponge biomarker hypothesis. PNAS 113:2684–2689
Hall J, Smith AR, Goad LJ, Goodwin TW (1969) The conversion of lanosterol, cycloartenol and 24-methylenecycloartanol into poriferasterol by Ochromonas malhamensis. Biochem J 112:129–130
Hannich JT, Umebayashi K, Riezman H (2011) Distribution and functions of sterols and sphingolipids. Cold Spring Harb Perspect Biol 3:1–13
Haubrich BA, Collins EK, Howard AL, Wang Q, Snell WJ, Miller MB, Thomas CD, Pleasant SK, Nes WD (2015) Characterization, mutagenesis and mechanistic analysis of an ancient algal sterol C24-methyltransferase: implications for understanding sterol evolution in the green lineage. Phytochemistry 113:64–72
Honda A, Yamashita K, Miyazaki H, Shirai M, Ikegami T, Xu GR, Numazawa M, Hara T, Matsuzaki Y (2008) Highly sensitive analysis of sterol profiles in human serum by LC-ESI-MS/MS. J Lipid Res 49:2063–2073
Iida T, Jeong M, Tamura t, Matsumoto T (1980) Identification of chondrillasterol in two Cucurbitaceae seed oils by proton nuclear magnetic resonance spectroscopy. Lipids 15: 66–68
Itoh T, Kikuchi Y, Tamura T, Matsumoto T (1981) Co-occurrence of chondrillasterol and spinasterol in two cucurbitaceae seeds as shown by 13C NMR. Phytochemistry 20:761–764
Kaya K, Nakazawa A, Matsuura H, Honda D, Inouye I, Watanabe MM (2011) Thraustochytrid Aurantiochytrium sp. 18W-13a accumulates high amounts of squalene. Biosci Biotechnol Biochem 75:2246–2248
Knapp FF, Goad LJ, Goodwin TW (1977) The conversion of 24-ethylidene sterols into poriferasterol by the alga Ochromonas malhamensis. Phytochemistry 16:1683–1688
Koklesnikova MD, Xiong Q, Lodeiro S, Hua L, Matsuda SP (2006) Lanosterol biosynthesis in plants. Arch Biochem Biophys 447:87–95
Kumar A, Fogelman E, Weissberg M, Tanami Z, Veilleux R, Ginzberg I (2017) Lanosterol synthase-like is involved with differential accumulation of steroidal glycoalkaloids in potato. Planta 246:1189–1202
Lenton JR, Hall J, Smith AR, Ghisalberti EL, Rees HH, Goad LJ, Goodwin TW (1971) The utilization of potential phytosterol precursors by Ochromonas malhamensis. Arch Biochem Biophys 143:664–674
Lewis TE, Nichols PD, McMeekin TA (2001) Sterol and squalene content of a docosahexaenoic-acid-producing thraustochytrid: influence of culture age, temperature, and dissolved oxygen. Mar Biotechnol 3:439–447
Li ZY, Ward OP (1994) Production of docosahexaenoic acid by Thraustochytrium roseum. J Ind Microbiol 13:238–241
Lu Y, Zhou W, Wei L, Li J, Jia J, Li F, Smith SM, Xu J (2014) Regulation of the cholesterol biosynthetic pathway and its integration with fatty acid biosynthesis in the oleaginous microalgae Nannochloropsis oceanica. Biotechnol Biofuels 7:1–15
Miller BM, Haubrich BA, Wang Q, Snell WJ, Nes WD (2012) Evolutionary conserved ∆25(27)-olefin ergosterol biosynthesis pathway in the alga Chlamydomonas reinhardtii. J Lipid Res 53:1636–1645
Nagano N, Taoka Y, Honda D, Hayashi M (2009) Optimization of culture conditions for growth and docosahexaenoic acid production by a marine thraustochytrid, Aurantiochytrium limacinum mh0186. J Oleo Sci 58:623–628
Nakazawa A, Matsuura H, Kose R, Kato S, Honda D, Inouye I, Kaya K, Watanabe MM (2012) Optimization of culture conditions of the thraustochytrid Aurantiochytrium sp. strain 18W-13a for squalene production. Bioresour Technol 109:287–291
Nakazawa A, Kokubun Y, Matsuura H, Yonezawa N, Kose R, Yoshida M, Tanabe Y, Kusuda E, Thang DV, Ueda M, Honda D, Mahakhant A, Kaya K, Watanabe MM (2014) TLC screening of thraustochytrid strains for squalene production. J Appl Phycol 26:29–41
Nes WD (2011) Biosynthesis of cholesterol and other sterols. Chem Rev 111:6423–6451
Nes WR, Krevitz K, Joseph J, Nes WD (1977) The phylogenetic distribution of sterols in tracheophytes. Lipids 12:511–527
Ohyama K, Suzuki M, Kukuchi J, Saito K, Muranaka T (2009) Dual biosynthetic pathway to phytosterol via cycloartenol and lanosterol in Arabidopsis. PNAS 106:725–730
Ori K, Koroda M, Mimaki Y, Sakagami H, Sashida Y (2003) Lanosterol and tetranorlanosterol glycosides from the bulbs of Muscari paradoxum. Phytochemistry 64:1351–1359. https://doi.org/10.1016/s0031-942
Patterson GW (1967) Sterols of Chlorella. II. The occurrence of an unusual sterol mixture in Chlorella vulgaris. Plant Physiol 42:1457–1459
Patterson GW (1971) Distribution of sterols in algae. Lipids 6:120–127
Patterson GW (1991) Sterols of algae. In: Patterson GW, Nes WD (eds) Physiology and biochemistry of sterols. American Oil Chemists’ Society Champaingn, Illinois, pp 118–157
Patterson GW, Doyle PJ, Dickson LG, Chan JT (1974) Effects of triparanol and AY-9944 upon sterol biosynthesis in Chlorella. Lipids 9:567–574
Řezanka T, Vyhnalek O, Podojil M (1986) Identification of sterols and alcohols produced by green algae of the genera Chlorella and Scenedesmus by means of gas chromatography-mass spectrometry. Folia Microbiol 31:44–49
Riisberg I, Orr RJS, Kluge R, Shalchian-Tabrizi K, Bowers HA, Patil V, Edvardsen B, Jakobsen KS (2009) Seven gene phylogeny of heterokonts. Protist 160:191–204
Schaller H (2004) New aspects of sterol biosynthesis in growth and development of higher plants. Plant Physiol Biochem 42:465–476
Smith TJ (2000) Squalene: potential chemopreventive agent. Expert Opin Investig Drugs 9:1841–1848
Smith AR, Goad LJ, Goodwin TW (1972) Incorporation of stereospecifically labeled mevalonic acid into poriferasterol by Ochromonas malhamensis. Phytochemistry 11:2775–2781
Suzuki M, Xiang T, Ohyama K, Seki H, Saito K, Muranaka T, Hayashi H, Katsube Y, Kushiro T, Shibuya M, Ebizuka Y (2006) Lanosterol synthase in dicotyledonous plants. Plant Cell Physiol 47:565–571
Volkman JK (2016) Sterols in microalgae. In: Borowitzka MA, Beardall J, Raven JA (eds) The physiology of microalgae. Springer, Cham, pp 485–505
Weete JD, Kim H, Gandhi SR, Wang Y, Dute R (1997) Lipids and ultrastructure of Thraustochytrium sp. ATCC 26185. Lipids 32:839–845
Wilson WK, Sumpter RM, Warren JJ, Rogers PS, Ruan B, Schroepfer GJ Jr (1996) Analysis of unsaturated C27 sterols by nuclear magnetic resonance spectroscopy. J Lipid Res 37:1529–1555
Yao L, Gerde JA, Lee SL, Wang T, Harrata KA (2015) Microalgae lipid characterization. J Agric Food Chem 63:1773–1787
Yokochi T, Honda D, Higashihara T, Nakahara T (1998) Optimization of docosahexaenoic acid production by Schizochytrium limacinum SR21. Appl Microbiol Biotechnol 49:72–76
Acknowledgments
The authors thank Ms. Yasuko Yoshikawa and Ms. Junko Hayashi (National Institute for Environmental Studies, Japan) for measurement of EIMS spectrum. The authors also thank Dr. Daiske Honda and Ms. Mayumi Ueda (Konan University, Japan) for their taxonomic advices about A. limacinum and A. mangrovei.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yoshida, M., Ioki, M., Matsuura, H. et al. Diverse steroidogenic pathways in the marine alga Aurantiochytrium. J Appl Phycol 32, 1631–1642 (2020). https://doi.org/10.1007/s10811-020-02078-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10811-020-02078-4